Potri.003G151500 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G151500.3 pacid=42786317 polypeptide=Potri.003G151500.3.p locus=Potri.003G151500 ID=Potri.003G151500.3.v4.1 annot-version=v4.1
ATGACGACAAGTGTAGCTTCTCAATTACAAGCAATCCGGTCAGTGATTCAAACTGGTTTGGAGTCCAAAAAACGTCCCATCACCCGACCCTCCATTTTAT
TCGATCCTAAAGAAGCTGCTGATCTTGATATCGATACTATACTTGACATTGCACTTTCAGGTTTGGAAGTTCTTGTAAGCGCGGATGAAAGGTTTAAGAA
CTATAAAAATGATCTGTTTAGTCATAAAAGTAAAGAGTTGGATAGAGAATTGATGACTGGCGAGGAAAATAAGCATATTAACTCGACCATCAGTTCATAT
TTGAGGTTACTATCTGGACATCTTCAATTGGCTGCATCACTTAGGACACTTGAGTACTTGATACGAAGATACAAGATACATGTATACAATTTCGAGGATT
TGATTTTATGCTCATTGCCTTACCATGATACTCATGCATTTGTTCGAATAGTTCAGTTGATTGATACAAGGAATGGTAAATGGAAGTTTCTAGATGGTGT
AAAAGCATCAGGTGCACCGCCACCTAGAAATGTCATGGTGCAGCAGTGTGTTCGAGATATGGGAGTTTTGGAGGCGTTGTGCAACTATGCATCTCCTGCA
AAGAAGTTCCAGCCATCAAGGTCTGTTGTCAGCTTTTGCACAGCAGTGGTCATAGAGGTATTGGGTTCTATAACTACAGTCAATACAGATGTTGTGCAGA
GAATCCTTCCATTTGTTATTTCTGGACTTCAGCCTGGCAGCAAAGGAGGTTCAGATCATAAGGCTGCTGCTTTGATGATTGTTTGTCTATTAGCAAATAA
GGTTTCCTTGTCTCCAAAACTTGTCAAAAGTTTGATGCGATCAATTGCTGAGATAGCTCAGAAAGATGCAAGCAAGTCAACTGATCTGCAATGGTTTCGT
TTGTCAATTATGGCTCTAATAAATCTCGTTCAGTTGCAATCAATCGATGTGTTCCCAAAGAAGGTACTGGAGATTTTGAAGGAAACCAGGGAAATAGCTG
GGGTTCTTATGGGACTTTCTAAGGAGTTCAATATCGATAGATTTCTTGCTGTGCTTTTGGAAGCTCTGGTTGACAACAGTTCTTCTGATGATACATATCA
TCATGTTTTGGTATCCATATTGGAGACAGTTCCCATAAAGAATTTTGTTGATCGTGTTGTTTCAAAGGTCCTCTTGTCTTGCATGAAAATGTCTCAGAAA
AATAGCAACCCATCATCTCAATCTGGAAGCTGGGCAAAGGACATTCTGATGGTTATTAATAAAATTTATCCATTTGAACTTCATCAAGCTGTTCAGAAGT
TTTTGGAGGACACCAAAGTGCAATCCAAAAATGATGATGCAGTTTTTGAGATTTGCAAAATGTTGGATGGGAACTTGGATATGTCAGCTTCCATCTCTGA
TTCTAAAATTTGGCTTGCTTTGCACCATCCAAAGGCTGAGGTCCGGCGTGCTACATTGTCTGGTTTAAACAGACATGTTGATCTGAAAAACATGGCTGTT
GATTCAAAGAGGCTTGTAACCATTCAAGATGCTGTATTGTGCCAACTTCGTGATGATGACCTAACAGTTGTTCAAGCAGCTTTATCTCTTAAAGGATTGT
CTGAAATCATCAGTCCTTCTGATCTTCTCAAAGCATTAGATGGCGTGCTTAAAAAATGTGTTAGCACCCTAAGGTCAGGTGCATCAGATAAAGCTGCTTT
AGCTAATGACGTTGCAATTGCATTTTTGAAGACTGCTGTCTCAACATTTCATGATCAGATTGATTATTCAAAGAAACTTGCTGCCATGATGTTTCCTCTT
CTCCTAATTTTTCAAAAGACACAGAGCCTAAATTTGGAGGTCTTGGAATTAGTGAAAGAAGTGAAGTGGCCATTTTATAATGATCTCACCGCTGTTTCCT
CTGAAGTGGTGAAATTACAACAGGAAGTTATATCTTCGATCAACATGAAGATTGTTAATGGCTTGGCAGAAACATTTTCAATGCACCCAGGTAAGTATAT
GACCTGGCTTGTCGACAGCTCCAGTGACTGTACTGTCTCGAAGACGCTTCTGCTCTTGGTTCTGATGCAATCATTTATTAGGCCAAAGAATAAAAGTGAG
CAATTCTCAGCACTTTTTGAAGCTTTCTTTTCTTTTCTGAAGACCGAGTGGGAGCTGCAGTCTGCAGTTGTTTCTGGGAATGAGTTCAATAATGAAATGC
TTCAATGGGATTGTGGAAGGTTCCTAGATCAGCTGTTTGACACCGATCTTAAGGCATTAAATATTAATATCCTAATCTGCACCTTTTGGAGGTTACTGGA
GGCTTTTACTTCAATGGAGGACAATCAGCAGTTGATTAGCAGCAGACATACAGATTTATTTGTCTTTTTCTCGAACTCTCAGTCCAAGCATTTTTTCAAG
GAGCATCTTCATTACCTTGTTACAAAATGCAAGATCTCTCCTATTGATTTTCTGTCTGGTTTTTACACTAATGAAGACATTTCCATTACTGTCCAAGTTG
AGAGCCTTCATTGTCTTGCATTCCTTTGCTCAGAACCCGACGATAGATTGCTGCTTCAGCTTCTATTCAGTTTCCCCTCACTTCTTGTTCCTCTTGCCTC
TGATAGCCAGGATCTCAGGATTGCTTCCATGGGCTGCATTGAAGGGTTATCTGCTCTGTCACATCGCGCTGACTATTTAAGCAAGAAAAATGGTATTTCT
GGGAACAATGCAAATTGGAGTCATTTTCTTGATGAGTTATTGGGCCTGATTGTGCAACAAAAAAGGCTTATATTATCAGACAGTAATTTTCTACCCTCGT
TTTTGTGCTGTTTGCTTGGCTCATCCAGGAATAGCCTTCTGGTACCACAAAATGTTGAACAAAGATTTGATCAATCTACAAAAGAAAAGATTCTTGCCTT
TGTCTTGGGGTCTGGACTTCAACTTTCCTCGTTTGCAAAGATGATGATTATATCTTTGCTTAAAGGAATGGGCAGTGCGTTACTGCATGTTAAAGAAGCT
GAGTCATTGCTGTCTCAACTTCTGAAAAGACGCCGTCAATATTATTTCGAGGTGGACCGGTCATCCCAGAAATTGTCCAAAACCGAAGTCAAGATCTTGT
GCCTTCTACTGGAGGTTTGTGCCATGCCTCCATCACTGGAGGGGCATGCTTGTGAAGATTATCTATTAAAGGCCCTGCAACTGGATGGGCTCTCTTCTGA
AGAATTTGCCATTATAGAACCTTGCATTACTGTTTTGCAGAAGCTAAGTGCCCCATTATACAGTGGATTGACAACTGAAAAACAGGAGCTTCTATTTCGT
GAGCTTGTGATTTTGTTTCGCAATGCTAATGGTGATATACAAAATGCAACAAGAGAGGCCTTGATGCGCTTAAATATTACCTGCTCTACGGTAGTCCATA
CAATCAAATTCATATTCGAACAAGAAAGTCGTATAGGTGGTTCTGCATCCGGGAAGAAGAAAAGGAAATCTATAGTACATCAAACATCCACCTTAGATGG
TGATGTGGTTTGTAAAGTAGAAACTGCATTATGCTTGCTAAGCTCCCTCCTTGACATATTGATTCTCAAGAAAGATATTGCTAGCAGGGAGCATCTGATA
GGGCCCTTATTCAAACTTCTTGAGAAAATCTTTTCGGATGATTGGATGCCTGCCCGAGATGAAAACTGGATCAAAGCTTCATATGGAGTTTCCCAGACTG
GGTCTAGCACAATTTGTTACACTCAGCAAACACTTCTACTTGTTCTTGAAGATATTATCGGTTCACTTAAAAATGTAATCCCATTAAAGGATGACATAAC
AAATAAAATCAATATCAAATTGTTGATCATGTGTGCCCGGTCTGCTAAGCATGGAGTCGTCCGCAACCATGTATTTTCACTGTTATCATCCATTGTAAAG
GTTGTTCCAGAGAATATAATGGGGTATATATTGGATATATTTACCGTTGCTGGAGAATCAACTGTTTCACAGATTGACAGCCATTCACAACATGTGTTTG
AGGATCTTATATCTGCAGTTGTCCCCTGCTGGCTGGCTGAGACACGCAACACAGACAAATTGCTCCAGGTTTTTGTGAATGTATTGCCCAAAATTGCTGA
GCATAGGAGGCTGTCAATCGTCGTGTACTTGTTGAGGACTTTGGGTGAACATAATAGCTTGGCTTCGTTGCTTGCCCTTCTTTTCCGGTCCTTAGTTTCT
AGGAAAGGACTCTCCCTCCTTGATGAAACAAATGATCTTACATCTTCTGCAGAGAGAGAGTGGGAGTATGCATTTGCAATTCGGATATGTGAGCAATATT
CATGCAGGATTTGGCTTCCCTCCCTTGTTCCTCTGCTTCAACTAATAGGAGCTGGCAATTCATGCCAAGAAATTTTTATGGAATTGCTATTTGCAACAGA
ATTTATTTTACACAAGTTAGAAGATCCAGAATTCTCTTTCAAACTTGATTCAAGCGAGGATTCAGATAAAATTCAGGAAACCCTTCAAGAACTTTTGGAG
CACGTTGTTTGCCTTTCACAACTAAGCGATTTGAGAAGAAAACAAATAAATGTGCCTGTTAGAGTCAGAAAGGAAATGAAAGAGTGTATGCATGGTGTTT
TGAGGTCTACTACAGCTGTTATGATCCCCTCCGCATACTTCAGAGGCATCATTAGTTTGCTTTGCAATTCAGATGGAAATGTGAAAAAGAAGGCTCTTGG
GCTTCTTTCTGAGACGTTGAAGAAGCGTGAGTCAATTAAAACGAAACACAAGGGAAGAAGAGATTCCATTGCAAGTTCAATTACTGATTGGTTCCATGTG
GATGGGAGTACCCTCGACTCTTTTCAGCAGATGTGTTTGGAAATTGCTCGGCTAATTGATGATACAATGGATGACTCAGATACCTCCTTGAAACTGTCAG
CAGTTTCTACACTGGAAGTTTTGGCCCATAGATTTTCTTCTAATTATTCTGTCTTCAGCATGTGCCTTCCATCTATCACAAAAGGCATTTGTTCGAACAA
CTTGGCTATTTCTTCTAGCTGCCTTCGAACAACTGGTGCTCTAGTCGATGCTCTTGGGCCAAGAGCATTTGTTCAGCTTCCTCAAATAATGGAGAATGTG
ATAAAAACTTCTAGCAAATTCTCTGCTGCACTGTCTCTTCCTGAAGAATCTCTCATGCTCTCTATTCTTCTTGCTTTAGAAGCTGTTGTAGACAAGCTGG
GTGGTTTCCTCAATCCATATCTTGAAGATATTATACGACTCGTGGTGCATGGTCCTGAGTACACATCAGGATCAAAAATGAAACTGAGACAGAAAGCTGA
TGCAGTACGGAAGCTGCTTACTGAGAAGATCCCTGTTCGACTAGCTCTTCCACCGTTGTTGAAGATGTATCCTGATACTGTGGAGGCTGGGGATTCAAGT
TTGGCAGTTTTCTTTGAAATGCTTGGAAGCTTAGTCGGTACAATGGACAGATCATCTGTTGGTGGTTATAATGAAACCATTTTTGACCTCTGCTTGCGTG
CTCTTGATCTTCGTCGTCAACATCCTGTTTCAATTCAAAATATTGATCTAGTTGAGAAAAGCATTATCAATGCAATGATTGCTCTGACAATGAAACTTAC
GGAGACCATGTTTAAGCCGCTCTTTATTAGAAGTATTGAATGGGCTGAGTCATATGTAGAAGAAAATGACAGTAAGGACAATGTTATAGATAGGGCCATA
TCTTTCTATGGTCTGGTAAACAAACTTGCTGAGAATCATCGGTCTCTGTTCGTCAGTTACTTTGAGTACTTGCTAGAGGGTTGTGTTCGGCATCTTACCA
ATATTGTAAAGCCTAAAGGTGCTGGTTTGATTCAAAAGAAAAAGAAAGCTAAGATTCAGGAAGCTGGAAGTGATATAAAAGAGAACAGTGTCCTAACACT
TAAAAGCTGGCATCTCAGGGCTTTGGTCATTTCAGCTTTACATAAATGCTTTTTGTACGACACTGGGAGCCGGAAGTTTCTTGATTCGTCAAAATTTCAG
GTTCTGTTGAAGCCCATTGTATCACAGCTTATAGCCGAACCACCAGCATTACTGGAAGAGCATCCAAGTATCCCATCAGTTAACGAAGTTGATGAATTGT
TGGTAGTTTGTATAGGTCAAATGGCAGTTACTGCAGGAACTGATCTTCTTTGGAAGCCCCTGAACCATGAGGTATTGTTGCAAACTCGGAGTGACAAGAT
ACGATCTCGAATTCTAGGACTGAGAATTGTTAAATATCTTATGGATAACCTGAAAGATGAATATTTGGTCTTCCTGCCTGAAACCATCCCATTCCTTGGT
GAATTGCTCGAGGACTTGGAGTTACCTGTTAAATCTCTGGCCCAGGATGTTCTAAAGGAAATGGAGTCCATGAGTGGTGAAAGCCTTCAGCAATATCTCT
GA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.003G151500.3 pacid=42786317 polypeptide=Potri.003G151500.3.p locus=Potri.003G151500 ID=Potri.003G151500.3.v4.1 annot-version=v4.1
MTTSVASQLQAIRSVIQTGLESKKRPITRPSILFDPKEAADLDIDTILDIALSGLEVLVSADERFKNYKNDLFSHKSKELDRELMTGEENKHINSTISSY
LRLLSGHLQLAASLRTLEYLIRRYKIHVYNFEDLILCSLPYHDTHAFVRIVQLIDTRNGKWKFLDGVKASGAPPPRNVMVQQCVRDMGVLEALCNYASPA
KKFQPSRSVVSFCTAVVIEVLGSITTVNTDVVQRILPFVISGLQPGSKGGSDHKAAALMIVCLLANKVSLSPKLVKSLMRSIAEIAQKDASKSTDLQWFR
LSIMALINLVQLQSIDVFPKKVLEILKETREIAGVLMGLSKEFNIDRFLAVLLEALVDNSSSDDTYHHVLVSILETVPIKNFVDRVVSKVLLSCMKMSQK
NSNPSSQSGSWAKDILMVINKIYPFELHQAVQKFLEDTKVQSKNDDAVFEICKMLDGNLDMSASISDSKIWLALHHPKAEVRRATLSGLNRHVDLKNMAV
DSKRLVTIQDAVLCQLRDDDLTVVQAALSLKGLSEIISPSDLLKALDGVLKKCVSTLRSGASDKAALANDVAIAFLKTAVSTFHDQIDYSKKLAAMMFPL
LLIFQKTQSLNLEVLELVKEVKWPFYNDLTAVSSEVVKLQQEVISSINMKIVNGLAETFSMHPGKYMTWLVDSSSDCTVSKTLLLLVLMQSFIRPKNKSE
QFSALFEAFFSFLKTEWELQSAVVSGNEFNNEMLQWDCGRFLDQLFDTDLKALNINILICTFWRLLEAFTSMEDNQQLISSRHTDLFVFFSNSQSKHFFK
EHLHYLVTKCKISPIDFLSGFYTNEDISITVQVESLHCLAFLCSEPDDRLLLQLLFSFPSLLVPLASDSQDLRIASMGCIEGLSALSHRADYLSKKNGIS
GNNANWSHFLDELLGLIVQQKRLILSDSNFLPSFLCCLLGSSRNSLLVPQNVEQRFDQSTKEKILAFVLGSGLQLSSFAKMMIISLLKGMGSALLHVKEA
ESLLSQLLKRRRQYYFEVDRSSQKLSKTEVKILCLLLEVCAMPPSLEGHACEDYLLKALQLDGLSSEEFAIIEPCITVLQKLSAPLYSGLTTEKQELLFR
ELVILFRNANGDIQNATREALMRLNITCSTVVHTIKFIFEQESRIGGSASGKKKRKSIVHQTSTLDGDVVCKVETALCLLSSLLDILILKKDIASREHLI
GPLFKLLEKIFSDDWMPARDENWIKASYGVSQTGSSTICYTQQTLLLVLEDIIGSLKNVIPLKDDITNKINIKLLIMCARSAKHGVVRNHVFSLLSSIVK
VVPENIMGYILDIFTVAGESTVSQIDSHSQHVFEDLISAVVPCWLAETRNTDKLLQVFVNVLPKIAEHRRLSIVVYLLRTLGEHNSLASLLALLFRSLVS
RKGLSLLDETNDLTSSAEREWEYAFAIRICEQYSCRIWLPSLVPLLQLIGAGNSCQEIFMELLFATEFILHKLEDPEFSFKLDSSEDSDKIQETLQELLE
HVVCLSQLSDLRRKQINVPVRVRKEMKECMHGVLRSTTAVMIPSAYFRGIISLLCNSDGNVKKKALGLLSETLKKRESIKTKHKGRRDSIASSITDWFHV
DGSTLDSFQQMCLEIARLIDDTMDDSDTSLKLSAVSTLEVLAHRFSSNYSVFSMCLPSITKGICSNNLAISSSCLRTTGALVDALGPRAFVQLPQIMENV
IKTSSKFSAALSLPEESLMLSILLALEAVVDKLGGFLNPYLEDIIRLVVHGPEYTSGSKMKLRQKADAVRKLLTEKIPVRLALPPLLKMYPDTVEAGDSS
LAVFFEMLGSLVGTMDRSSVGGYNETIFDLCLRALDLRRQHPVSIQNIDLVEKSIINAMIALTMKLTETMFKPLFIRSIEWAESYVEENDSKDNVIDRAI
SFYGLVNKLAENHRSLFVSYFEYLLEGCVRHLTNIVKPKGAGLIQKKKKAKIQEAGSDIKENSVLTLKSWHLRALVISALHKCFLYDTGSRKFLDSSKFQ
VLLKPIVSQLIAEPPALLEEHPSIPSVNEVDELLVVCIGQMAVTAGTDLLWKPLNHEVLLQTRSDKIRSRILGLRIVKYLMDNLKDEYLVFLPETIPFLG
ELLEDLELPVKSLAQDVLKEMESMSGESLQQYL
|
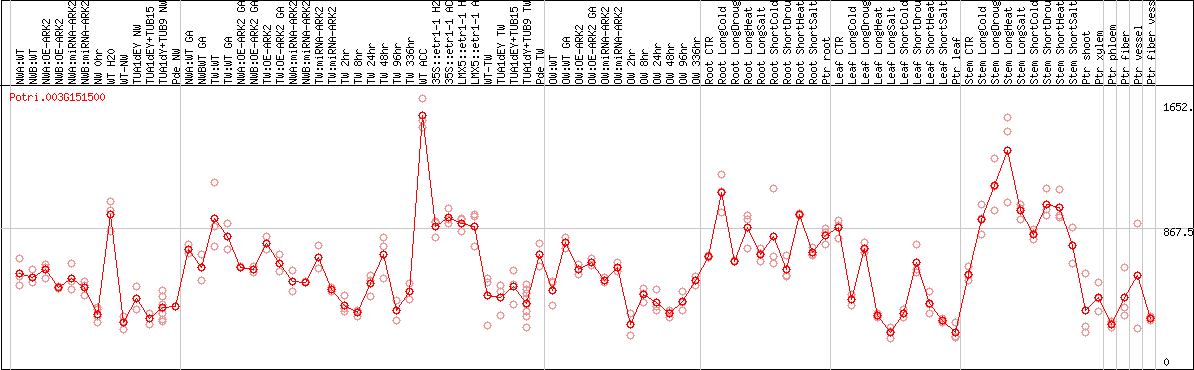
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G151500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
