Potri.003G169000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G169000.1 pacid=42785828 polypeptide=Potri.003G169000.1.p locus=Potri.003G169000 ID=Potri.003G169000.1.v4.1 annot-version=v4.1
ATGAGTAGAGAGAAGTTTAATAAATTAAGACACGACCATTGTTGTTTTCTCTCTGTATCCATCTTTGTTGTTGCATTTGCTTTATCATTGAGCCCTGCTT
ATTGTCAAGATATCGGTGGCTATGATGACAATCCAGCCACACAGGAGCTATTTTCAGAGCTGGTTTACAAGAGTTTCTCAAATTTTACCTCCGTGTTCAA
GCAAGATATTGTCAAATATTTTGGTTTCTGCATAACAGACGTGGATGAAGATTGGAATATGGCATTCAATTTCTCCAAAGGTACACAATTCATATCCAAT
TGCGCCAAGAAGACAAAAGGAGACATGCTAAGGCGTACATGCACAGCAGCAGAAATAAAGTTCTATTTCAACAATCTCTTTGAAAAAGGAGCAAAGAAAA
GCAATTATTTGAAACCTAACAAGAACTGTAATTTGAGTTCATGGGTTTCTGGGTGTGAGCCAGGATGGGCTTGTGGAGTTGGTAAAGGTGAAAAGGTTGA
CCTCAGAAATTCAAAGGACATGCCTTTTAGAACGACAAATTGTGCAGCTTGTTGTGAGGGTTTCTTCTGCCCTCATGGTATTACTTGCATGATACCTTGC
CCACTGGGTGCTCACTGTCCCCTTGCAAAACTGAACAAAACCACTGGCATATGCGACCCATATCATTACCAATTGCCTCCAGGAAAGCCAAACCATACTT
GTGGTGGAGCGGATATTTGGGCTGATATTTTGAGTAGTAGCGAAATATTCTGTTCAGCAGGATCATACTGTCCGTCCACAATTCAAGAGATTCCATGCAG
TCGTGGACATTACTGCAGGACTGGTTCAACGTCTCAAACAGGGTGTTTTAATCTGGCAACTTGTGAAACACAATCAGCAAATCAGAATATCACTGCATAT
GGCATTTTATTTTTCGCTGGATTAAGCTTTCTGCTTATCATTATTTATAACTGTTCTGATCAAGTTCTTGCAACTCGAGAGAGAAGACAAGCAAAGACAA
GGGAGAAGGCAGTTCAAAGTGTAAGAGAAACTGCTCAAGCTCGTGAAAAATGGAAATCTGCAAGAGACATTGCCAAGAAGGGTGCGATTGGATTACAAAC
CCAGCTGTCACGCACATTTTCTCGTACAAAGTCTAAAAGGCCAGTAGAACAGCTAAAAGGTTTTGGTCAAGCTAAACCCGGATCGGATGCTGCTTTACCA
CCCATGCCGGTAAGCAGTTCATCTCAGCAATCATCTGGAAAAGGGAAGAAAAAGGGAAAGAGCAACCTCTCACAGATGCTGGATGATATTGAGAACAACC
CCGAGGGTCACGAAGGCTTTAATTTGGAAATTGGAGATAAAAATATCAGAAAAAATGCACCAAGGGGTAAGCAGTTGCATACTCAGAGTCAAATGTTCAG
GTATGCATATGGACAAATCGAGAGGGAGAAAGCTATGCAGGAGCAGAACAAGAATTTGACCTTCTCTGGGGTAATTTCAATGGCTAACGATACCGAAATC
AGGAAAAGGCCTTCTATTGAGGTTGCTTTTAAAGATTTAACCCTCACTTTGACGACTAAACACAAACATCTGCTAAGATGTGTGACTGGGAAATTATCAC
CTGGCCGAGTTTCTGCTGTCATGGGTCCGTCTGGGGCTGGAAAAACAACATTTCTTTCTGCTTTGACTGGAAAAGCAACTGGCTGCACCATGTCTGGTAT
GGTTCTTGTAAATGGTAAGATGGAACCAATCCAAGCATACAGGAAAATCATTGGTTTTGTGCCTCAAGATGATATTGTGCATGGAAACCTAACAGTGGAG
GAGAATCTCTGGTTCAGTGCAAGATGCAGACTCTCTGCTGATCTACCAAAACCAGAAAAGGTTCTAGTTGTTGAAAGAGTTATTGAGTCCTTGGGACTGC
AGGCAGTGCGAGATTCTCTGGTTGGGACGGTGGAAAAGCGAGGAATTTCTGGAGGTCAAAGAAAACGTGTCAACGTTGGTCTTGAAATGGTTATGGAACC
TTCACTTCTAATATTAGATGAACCCACATCAGGCTTGGACAGTTCATCGTCCCTGTTGCTACTTAGAGCTCTTCGACGCGAAGCTCTTGAAGGAGTAAAC
ATATGCATGGTTGTACACCAACCAAGCTATACATTATTCAGGATGTTTGATGATTTGATACTTCTAGCTAAAGGTGGCTTGACAGCATATCACGGATCGG
CCAAGAAAGTCGAGGAGTATTTTGCTGGCCTTGGAATTACTGTACCTGAGCGCGTGAATCCTCCAGATTACTTCATTGATATTTTGGAGGGCATAGCGAA
ACCAAAATCAGGGGTGAACTATAAGCAACTTCCAGTTAGATGGATGCTTCACAATGGCTACCCTGTTCCAATGGATATGCTGCAGAATACAGATGGATTG
GGAGCATCCTCCAGTGAAAATTCTGCTCATGGAGCAAGTGAAGTTGGATCTGAAACAGGGTCTTTAGCTGGAGATTTTTGGCATGATTTGAAATCTAATG
TTGAGTCAGAGAAAGACAACTTGAAACCCAACGTTTTGAAATCAGGTGACTTATCCGAACGGAGAAGCCCTGGAGTATATCAGCAGTATAGATACTTCCT
GGGGAGGGTTGGAAAGCAGCGGCTACGAGAAGCAAGGGCACAAGCTGTAGATTATTTGATTTTATTGCTAGCTGGAATCTGCTTAGGAACACTAGCTAAA
GTGAGCGATGAAACTTTTGGGGTGGTTGGTTACACATACACCGTCATTGCAGTTTCACTACTGTGTAAGATCGCGGCTTTGAGGTCATTTTCTCTGGATA
AGCTACATTACTGGAGAGAGAGATCTTCTGGGATGAGCAGCTTGGCATACTTTCTTGCAAAGGATACAATAGACCATTTCAGCACAATTGTCAAGCCACT
TGTCTATCTGTCAATGTTCTACTTCTTCAACAATCCACGATCAACAGTCTTCGATAACTACATAGTCTTAATCTGTCTGGTGTATTGTGTGACTGGAATA
GCCTATGCATTGGCCATTTTCTTTGAACCAGGTCCAGCCCAACTGTGGTCAGTGCTGCTTCCGGTTGTTCTGACCCTGATAGCAACCCGTACTGAAAATG
ATGGCGTTGTGAATTATATATCTAATTTGTGCTATACTAAGTGGGCGCTGGAAGCATTTGTGATATCAAATGCTAAAAGATATTATGGTGTTTGGCTGAT
AACAAGGTGCGGTTCACTCATGGAAAGTGGGTATGATTTAGGTCATTGGTATCGGTCTTTAATCCTCCTCGTTTTGACTGGCATAGTTAGTCGTGTTGCT
GCCTTTTTTATTTTGATTACCGTCAACAGGAAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G169000.1 pacid=42785828 polypeptide=Potri.003G169000.1.p locus=Potri.003G169000 ID=Potri.003G169000.1.v4.1 annot-version=v4.1
MSREKFNKLRHDHCCFLSVSIFVVAFALSLSPAYCQDIGGYDDNPATQELFSELVYKSFSNFTSVFKQDIVKYFGFCITDVDEDWNMAFNFSKGTQFISN
CAKKTKGDMLRRTCTAAEIKFYFNNLFEKGAKKSNYLKPNKNCNLSSWVSGCEPGWACGVGKGEKVDLRNSKDMPFRTTNCAACCEGFFCPHGITCMIPC
PLGAHCPLAKLNKTTGICDPYHYQLPPGKPNHTCGGADIWADILSSSEIFCSAGSYCPSTIQEIPCSRGHYCRTGSTSQTGCFNLATCETQSANQNITAY
GILFFAGLSFLLIIIYNCSDQVLATRERRQAKTREKAVQSVRETAQAREKWKSARDIAKKGAIGLQTQLSRTFSRTKSKRPVEQLKGFGQAKPGSDAALP
PMPVSSSSQQSSGKGKKKGKSNLSQMLDDIENNPEGHEGFNLEIGDKNIRKNAPRGKQLHTQSQMFRYAYGQIEREKAMQEQNKNLTFSGVISMANDTEI
RKRPSIEVAFKDLTLTLTTKHKHLLRCVTGKLSPGRVSAVMGPSGAGKTTFLSALTGKATGCTMSGMVLVNGKMEPIQAYRKIIGFVPQDDIVHGNLTVE
ENLWFSARCRLSADLPKPEKVLVVERVIESLGLQAVRDSLVGTVEKRGISGGQRKRVNVGLEMVMEPSLLILDEPTSGLDSSSSLLLLRALRREALEGVN
ICMVVHQPSYTLFRMFDDLILLAKGGLTAYHGSAKKVEEYFAGLGITVPERVNPPDYFIDILEGIAKPKSGVNYKQLPVRWMLHNGYPVPMDMLQNTDGL
GASSSENSAHGASEVGSETGSLAGDFWHDLKSNVESEKDNLKPNVLKSGDLSERRSPGVYQQYRYFLGRVGKQRLREARAQAVDYLILLLAGICLGTLAK
VSDETFGVVGYTYTVIAVSLLCKIAALRSFSLDKLHYWRERSSGMSSLAYFLAKDTIDHFSTIVKPLVYLSMFYFFNNPRSTVFDNYIVLICLVYCVTGI
AYALAIFFEPGPAQLWSVLLPVVLTLIATRTENDGVVNYISNLCYTKWALEAFVISNAKRYYGVWLITRCGSLMESGYDLGHWYRSLILLVLTGIVSRVA
AFFILITVNRK
|
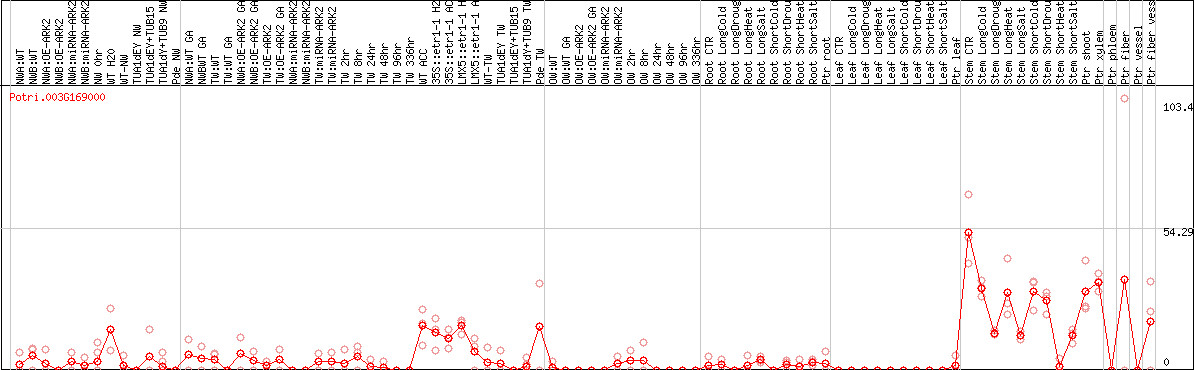
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G169000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
