Potri.003G178900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G178900.1 pacid=42786150 polypeptide=Potri.003G178900.1.p locus=Potri.003G178900 ID=Potri.003G178900.1.v4.1 annot-version=v4.1
ATGGAGAGTGCTGTTATTTCTAGAGGTAGTGACAGTTTTAGAGGTAGTTCTAGAGGAGTTTCGTCAGTATGGAGAAACAGCACTGTGGAAGTGTTTTCAA
GATCTTCAAGAGAAGAAGATGATGAAGAGGCTCTAAAATGGGCTGCGCTTGAGAAACTACCTACCTATGATCGTTTAAGGAAAGGTATATTAACGAGTGC
GTCAAGAGGTATAATTAGTGAGGTAGATATTGAGAACCTTGGAGTCCAAGAGAGAAAACAGTTGCTTGAGAGGTTGGTCAAAGTTGCAGACGAGGATAAT
GAGAAGTTCTTGTGGAAACTCAAGAACCGTGTTGAAAGGGTTGGAATTGAGTTTCCAACAATTGAAGTTCGGTACGAGAATCTAAATATTGAAGCAGAAG
CTTACGTGGGAAGTAGTGCTTTGCCTTCATTCGCCAAGTTCATTTTTCATATAATAGAGGGTTTTTTTATTGCTCTTCATGTCCTTCCAAGTAGAAAGAA
GCCACTCACCATCCTTAAGGATGTTAGTGGAATCATCAAGCCATCAAGATTGACTTTGCTTTTGGGTCCTCCGAATTCTGGGAAGACCACACTGTTGTTG
GCAATGGCTGGAAAGCTTGATCCTAGTCTGAAGTTTTCTGGTCATGTGACTTACAATGGTCATGAGATGAACGAATTTGTACCACAAAGAACGGCTGCCT
ATGTCAGTCAACATGATCTCCATATAGGAGAAATGACTGTGAGGGAGACTTTGGAATTCTCTGCCAGATGCCAAGGGGTTGGCCACCTACATGAGATGCT
AGCAGAGTTGTCGAGAAGAGAGAAAGAAGCCAACATCAAGCCTGATCAAGATGTTGATGTCTTCATGAAGGCAGTAGCAACACAAGGTCAAGAGGCCAGT
GTCATCACCGATTACGTCTTGAAGATATTAGGATTGGAAGTCTGCGCAGATACTTTGGTAGGGGATGAAATGATAAGGGGTATATCTGGAGGACAAAGGA
AGCGTGTTACAACCGGGGAAATGCTGGTCGGACCATCAAGGGCATTGCTTATGGATGAGATTTCTACAGGCCTGGATAGCTCAACAACTTACCAAATTGT
GAACTCATTAAAGCAAACTATTCACGTTCTCAATTGCACTGCTGTCATCTCACTTCTCCAGCCAGCACCAGAGACTTATGACCTCTTTGATGACATCATT
CTCCTGTCAGATGGCCAAATTGTGTACCAGGGTCCTCGTGAAAATGTGCTTGGATTTTTTGAACATATGGGCTTCAAGTGTCCCGACAGAAAAGGCGTGG
CAGACTTTCTACAAGAAGTGACATCAAAAAAAGATCAAGAGCAGTATTGGGCAATTAAAGATCAGCCTTACAGGTTTGTCAGGGTCAACGAATTTTCTGA
GGCATTTCAGTCATTCAACGTGGGACGAAAGATCGCAGATGAGCTTTCAATCCCATTTGACAAGACTAAGAACCATCCAGCTGCATTGGTAAACAAAAAA
TATGGAGCTGGAAAGATGGATCTGCTTAAAGCTAACTTCTCAAGAGAATACTTGCTCATGAAGAGGAACTCATTTGTCTACATCTTCAAGATATGCCAAC
TGACAGTAGTGGCATTGATTTCCATGTCACTCTTTTTTCGGACCAAGATGCATCATGATACGGTTGCCGATGGAGGAATTTATACCGGTGCTTTGTTCTT
CACTGTGATCATTATAATGTTTAATGGAATGTCAGAGCTATCTATGACCATCGTAAAGCTTCCTGTGTTTTACAAGCAAAGGGAACTCCTATTCTTTCCT
CCATGGGCATATTCAATTCCTCCATGGATTCTGAAAATTCCCGTCACGTTTGTTGAAGTTGCTGCATGGGTGCTCCTAACCTATTATGTAATTGGATTTG
ATCCAAATGTTGAAAGGCTTTTAAGGCAGTACTTTCTGCTTTTGCTAATTAACCAGATGGCATCTGCACTATTTCGATTTATTGCTGCAGCTGGAAGGAA
CATGATTGTTGCTAACACTTTTGGGTCTTTTGCGCTACTTACACTTTTTGCATTGGGTGGCTTTATCTTGTCTCGAGAGCAAATAAAAAAGTGGTGGATA
TGGGGTTACTGGCTTTCACCATTGATGTATGGGCAGAACGCAATAGTAGTGAATGAGTTCCTTGGACATAGCTGGAGTCATATTCCTGGAACCTCCACAG
AACCACTAGGAATCCAAGTTTTGAAGAGTCGCGAATTTTTCACTGAAGCAAATTGGTATTGGATAGGAGTAGGGGCAACAGTAGGATTCATGCTACTGTT
CAATATCTGCTTCGCTTTGGCCCTCACTTTCCTCAATGCATTTGAGAAGCCTCAAGCTTTTATATTTGAAGAATCCGAAAGGGAAGGTTCTGTTGGCAAA
ACAGGAGGAGCTGTTCAGTTATCAAACCATGGAAGTAGCCACAAAAACAAAACAGAGAATGGAGATGAAATTAATAGAAATGGTTTTGCGTCCATTGGTG
AAGCCAGCGATAACAGGAAGAGAGGGATGGTTCTTCCATTTGAACCACATTCCATCACATTTGATGATGTTATTTACTCCGTTGACATGCCACAGGAAAT
GAAAATTCAAGGAGTAGTTGAGGATAGATTGGTGCTTTTGAAGGGTGTGAATGGTGCTTTCAGGCCTGGTGTCCTTACAACTTTGATGGGTGTTAGTGGT
GCAGGAAAAACGACTTTGATGGATGTGCTGGCAGGTAGGAAAACTGGTGGATATATTGAGGGAGACATCAAAATTTCTGGGTACCCAAAGAAGCAAGAAA
CTTTTGCTAGAATTGCAGGATACTGTGAACAAAATGACATTCACTCTCCACATGTTACTGTCTACGAATCCTTACTCTACTCAGCCTGGCTTCGTCTACC
CCCTGAAGTTGATTCTGAAACCAGAAAGATGTTCATTGACGAAGTCATGGAGCTTGTGGAGTTGGATTCGTTGCGCAATGCATTAGTTGGGTTGCCTGGT
GTGAATGGTTTGTCCACTGAGCAGCGCAAGCGTCTAACCATTGCAGTAGAGCTCGTTGCAAACCCCTCTATCATATTCATGGATGAGCCAACTTCAGGGT
TAGATGCTAGAGCTGCTGCAATTGTGATGAGAACAGTAAGAAACACTGTGGACACAGGAAGAACAGTTGTATGCACCATCCACCAGCCCAGTATAGATAT
ATTTGATGCTTTTGATGAGCTATTCCTGATGAAGCGAGGAGGAGAAGAGATATATGTGGGGCCATTGGGTCACCATTCTACCCATCTAATTAAATACTTT
GAGGCAATTGAAGGAGTGAGTAAAATTAAAGATGGTTATAATCCAGCAACATGGATGTTGGAAGTTACTGCTTCATCACAAGAGATGGCTTTGGAGGTTG
ATTTTGCTAATATTTACAAGAATTCAGATCTGTTCAGGAGAAACAAAGCTCTCATTGCGGAATTAAGCACACCTGCTCCCGGCTCCAAGGACGTTCATTT
CCCTACACGGTATTCAACATCATTTTTCACACAATGTATGGCTTGCTTGTGGAAGCAACATTGGTCATACTGGAGGAACCCACCGTACACGGCCGTGAGA
TTTCTTTTCACAACCTTTATAGCCTTGATGTTTGGAACAATGTTCTGGGACCTTGGCTCCAAAGTGAAAACTACCCAAGATCTTTCCAACGCAATGGGTT
CGATGTATGCAGCAGTTCTCTTCCTTGGTTTTCAAAATGGAACAGCTGTGCAGCCAGTGGTGGCTGTTGAAAGAACTGTCTTCTACAGAGAAAGGGCTGC
TGGAATGTATTCAGCTTTGCCCTATGCCTTTGCACAGGCCCTGATTGAGCTTCCATATGTCTTTGTTCAAGCTGCAGTCTACGGCGTTATAGTTTATGCA
ATGATTGGATTTGAATGGACTGCAGCTAAGTTCTTTTGGTATCTATTCTTTATGTACTTCACACTATTATACTTCACCTTTTATGGAATGATGGCTGTAG
CTGTGACTCCAAACCACCACATAGCTGGCATAGTTTCCACAGCATTTTATGCAATCTGGAACCTCTTCTCAGGGTTTATAATTCCCCGAACTCGGATACC
TATATGGTGGAGATGGTACTACTGGGGATGCCCCGTATCATGGTCCTTGTATGGATTGGTTGTGTCACAGTATGGAGATATACAGGAACCCATTACCGCA
ACTCAAACAGTGGAAGGATATGTCAAGGATTACTTCGGTTTTGACCACGATTTTCTTGGAGTAGTTGCAGCCGTGGTTCTTGGGTGGACTGTCCTCTTTG
CCTTCATTTTCGCCTTCTCTATAAAGGCCTTCAACTTCCAGAGGCGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G178900.1 pacid=42786150 polypeptide=Potri.003G178900.1.p locus=Potri.003G178900 ID=Potri.003G178900.1.v4.1 annot-version=v4.1
MESAVISRGSDSFRGSSRGVSSVWRNSTVEVFSRSSREEDDEEALKWAALEKLPTYDRLRKGILTSASRGIISEVDIENLGVQERKQLLERLVKVADEDN
EKFLWKLKNRVERVGIEFPTIEVRYENLNIEAEAYVGSSALPSFAKFIFHIIEGFFIALHVLPSRKKPLTILKDVSGIIKPSRLTLLLGPPNSGKTTLLL
AMAGKLDPSLKFSGHVTYNGHEMNEFVPQRTAAYVSQHDLHIGEMTVRETLEFSARCQGVGHLHEMLAELSRREKEANIKPDQDVDVFMKAVATQGQEAS
VITDYVLKILGLEVCADTLVGDEMIRGISGGQRKRVTTGEMLVGPSRALLMDEISTGLDSSTTYQIVNSLKQTIHVLNCTAVISLLQPAPETYDLFDDII
LLSDGQIVYQGPRENVLGFFEHMGFKCPDRKGVADFLQEVTSKKDQEQYWAIKDQPYRFVRVNEFSEAFQSFNVGRKIADELSIPFDKTKNHPAALVNKK
YGAGKMDLLKANFSREYLLMKRNSFVYIFKICQLTVVALISMSLFFRTKMHHDTVADGGIYTGALFFTVIIIMFNGMSELSMTIVKLPVFYKQRELLFFP
PWAYSIPPWILKIPVTFVEVAAWVLLTYYVIGFDPNVERLLRQYFLLLLINQMASALFRFIAAAGRNMIVANTFGSFALLTLFALGGFILSREQIKKWWI
WGYWLSPLMYGQNAIVVNEFLGHSWSHIPGTSTEPLGIQVLKSREFFTEANWYWIGVGATVGFMLLFNICFALALTFLNAFEKPQAFIFEESEREGSVGK
TGGAVQLSNHGSSHKNKTENGDEINRNGFASIGEASDNRKRGMVLPFEPHSITFDDVIYSVDMPQEMKIQGVVEDRLVLLKGVNGAFRPGVLTTLMGVSG
AGKTTLMDVLAGRKTGGYIEGDIKISGYPKKQETFARIAGYCEQNDIHSPHVTVYESLLYSAWLRLPPEVDSETRKMFIDEVMELVELDSLRNALVGLPG
VNGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFDAFDELFLMKRGGEEIYVGPLGHHSTHLIKYF
EAIEGVSKIKDGYNPATWMLEVTASSQEMALEVDFANIYKNSDLFRRNKALIAELSTPAPGSKDVHFPTRYSTSFFTQCMACLWKQHWSYWRNPPYTAVR
FLFTTFIALMFGTMFWDLGSKVKTTQDLSNAMGSMYAAVLFLGFQNGTAVQPVVAVERTVFYRERAAGMYSALPYAFAQALIELPYVFVQAAVYGVIVYA
MIGFEWTAAKFFWYLFFMYFTLLYFTFYGMMAVAVTPNHHIAGIVSTAFYAIWNLFSGFIIPRTRIPIWWRWYYWGCPVSWSLYGLVVSQYGDIQEPITA
TQTVEGYVKDYFGFDHDFLGVVAAVVLGWTVLFAFIFAFSIKAFNFQRR
|
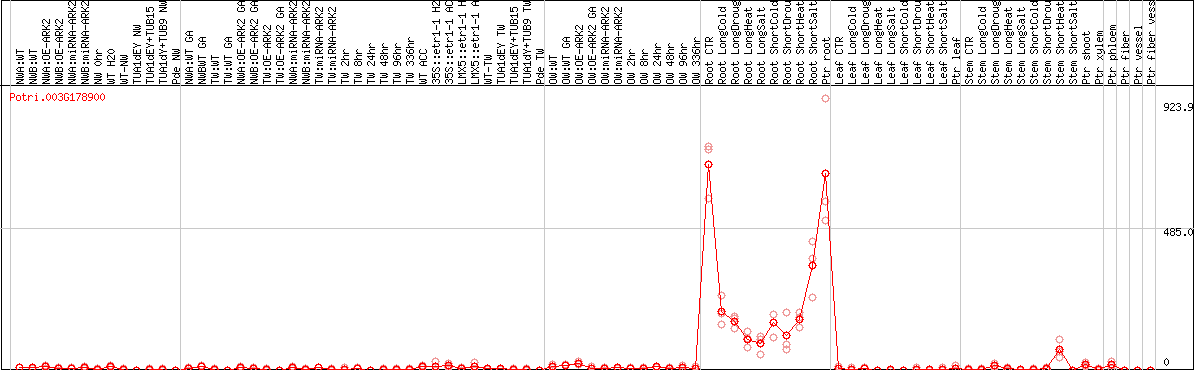
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G178900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
