External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G15860 348 / 6e-123
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
AT3G12760 114 / 8e-31
unknown protein
AT3G28970 44 / 5e-05
AAR3
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G047900
418 / 2e-150
AT1G15860 328 / 3e-115
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Potri.018G112200
314 / 2e-109
AT1G15860 267 / 4e-91
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Potri.008G083800
110 / 4e-29
AT3G12760 415 / 1e-148
unknown protein
Potri.004G122100
47 / 4e-06
AT3G28970 296 / 3e-100
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Potri.017G084300
45 / 2e-05
AT3G28970 269 / 1e-90
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039198
372 / 2e-132
AT1G15860 369 / 2e-131
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10013742
372 / 9e-131
AT1G15860 367 / 5e-129
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10019598
280 / 3e-96
AT1G15860 285 / 4e-98
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10040019
275 / 6e-94
AT1G15860 278 / 2e-95
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10026216
264 / 1e-90
AT1G15860 252 / 3e-86
Domain of unknown function (DUF298) (.1), Domain of unknown function (DUF298) (.2), Domain of unknown function (DUF298) (.3)
Lus10009599
110 / 1e-29
AT3G12760 380 / 3e-135
unknown protein
Lus10000875
110 / 1e-29
AT3G12760 383 / 3e-136
unknown protein
Lus10030685
51 / 2e-07
AT3G28970 255 / 4e-84
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
Lus10005235
45 / 2e-05
AT3G28970 258 / 1e-84
antiauxin-resistant 3, Domain of unknown function (DUF298) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0220
EF_hand
PF03556
Cullin_binding
Cullin binding
Representative CDS sequence
>Potri.003G180300.1 pacid=42785644 polypeptide=Potri.003G180300.1.p locus=Potri.003G180300 ID=Potri.003G180300.1.v4.1 annot-version=v4.1
ATGCGTCGGTCTGCATCGAAGAAGACGGGTCAATCCAATTCTACCACTGCTTCAATTACTTCATCGGCTACGGATCTTTTTCGCTCCTCCTCAAGTAAAG
CATCTTCAAAAGAAATGGAGCGAATCGATAACCTATTTTACTCATATGCAAACAGGTCTTCTGGTATGATTGACCCAGAAGGAATTGAAACTCTATGTTC
AGATATGGAGGTGGATCATACTGATGTTAGAATCTTGATGCTAGCTTGGAAAATGAGAGCTGAAAAACAGGGATACTTTACCCTGGAGGAGTGGCGACAG
GGCCTCAAATCACTGAGGGCTGATACTCTAAATAAATTAAAAAAGGCGCTCCCGGATCTAGAGAAAGAGGTTAAGAGGCCATCAAACTTTGTGGATTTCT
ATAACTATGCATTCCGGTATTGCCTAACAGAGGAGAAACAAAAGAGTATAGACATAGAGAGCATCTGCCAATTGTTGGATCTTGTTTTAGGATCCCATTT
CCAAGCCCAGGTTGATTACTTCATTGAGTATCTAAAGATTCAGAGTGATTATAAGGTCATAAACATGGATCAGTGGATGGGATTTTACCGTTTTTGCAAC
GAGATAAGTTTTCCAGACTTCAGTAATTATGATCCGGAGCTTGCATGGCCCCTGATCCTGGACAATTTTGTAGAATGGATGCGAGCCAAAAGGACCTAA
AA sequence
>Potri.003G180300.1 pacid=42785644 polypeptide=Potri.003G180300.1.p locus=Potri.003G180300 ID=Potri.003G180300.1.v4.1 annot-version=v4.1
MRRSASKKTGQSNSTTASITSSATDLFRSSSSKASSKEMERIDNLFYSYANRSSGMIDPEGIETLCSDMEVDHTDVRILMLAWKMRAEKQGYFTLEEWRQ
GLKSLRADTLNKLKKALPDLEKEVKRPSNFVDFYNYAFRYCLTEEKQKSIDIESICQLLDLVLGSHFQAQVDYFIEYLKIQSDYKVINMDQWMGFYRFCN
EISFPDFSNYDPELAWPLILDNFVEWMRAKRT
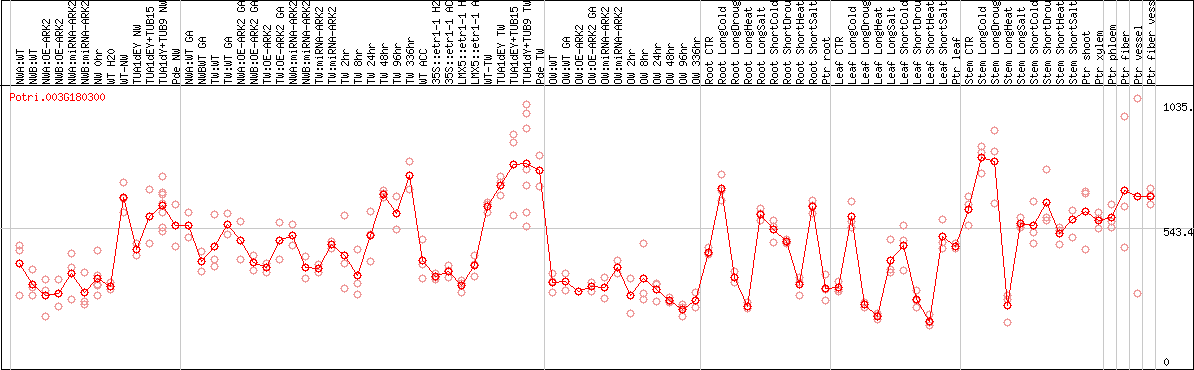
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G180300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.