External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G64660 288 / 1e-96
ATMGL
methionine gamma-lyase (.1)
AT3G01120 99 / 6e-24
AtCGS1, ATCYS1, CGS1, MTO1
METHIONINE OVERACCUMULATION 1, CYSTATHIONINE GAMMA-SYNTHASE 1, CYSTATHIONINE GAMMA-SYNTHASE, A. thaliana cystathionine gamma-synthetase 1, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
AT1G33320 77 / 2e-16
Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
AT3G57050 62 / 3e-11
CBL
cystathionine beta-lyase (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G084000
306 / 2e-103
AT1G64660 689 / 0.0
methionine gamma-lyase (.1)
Potri.017G086500
96 / 9e-23
AT3G01120 720 / 0.0
METHIONINE OVERACCUMULATION 1, CYSTATHIONINE GAMMA-SYNTHASE 1, CYSTATHIONINE GAMMA-SYNTHASE, A. thaliana cystathionine gamma-synthetase 1, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
Potri.003G146600
57 / 7e-10
AT1G64660 239 / 1e-76
methionine gamma-lyase (.1)
Potri.016G038200
57 / 2e-09
AT3G57050 714 / 0.0
cystathionine beta-lyase (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000880
306 / 6e-104
AT1G64660 639 / 0.0
methionine gamma-lyase (.1)
Lus10001672
296 / 2e-100
AT1G64660 546 / 0.0
methionine gamma-lyase (.1)
Lus10017333
297 / 5e-100
AT1G64660 676 / 0.0
methionine gamma-lyase (.1)
Lus10000881
278 / 4e-93
AT1G64660 588 / 0.0
methionine gamma-lyase (.1)
Lus10000074
213 / 1e-70
AT1G64660 244 / 2e-80
methionine gamma-lyase (.1)
Lus10033233
195 / 1e-61
AT1G64660 377 / 8e-130
methionine gamma-lyase (.1)
Lus10030694
92 / 3e-21
AT3G01120 702 / 0.0
METHIONINE OVERACCUMULATION 1, CYSTATHIONINE GAMMA-SYNTHASE 1, CYSTATHIONINE GAMMA-SYNTHASE, A. thaliana cystathionine gamma-synthetase 1, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
Lus10005226
90 / 1e-20
AT3G01120 741 / 0.0
METHIONINE OVERACCUMULATION 1, CYSTATHIONINE GAMMA-SYNTHASE 1, CYSTATHIONINE GAMMA-SYNTHASE, A. thaliana cystathionine gamma-synthetase 1, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
Lus10027313
59 / 6e-10
AT3G57050 683 / 0.0
cystathionine beta-lyase (.1.2.3)
Lus10039016
56 / 6e-09
AT3G57050 688 / 0.0
cystathionine beta-lyase (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0061
PLP_aminotran
PF01053
Cys_Met_Meta_PP
Cys/Met metabolism PLP-dependent enzyme
Representative CDS sequence
>Potri.003G187032.2 pacid=42786039 polypeptide=Potri.003G187032.2.p locus=Potri.003G187032 ID=Potri.003G187032.2.v4.1 annot-version=v4.1
ATGATTCTCTCCCCGGCAAAGCTTGGGGCAGATGTGGTTGTGCATAGCATCTCCAAGTATATTAGTGGTGAAGCGGATATTATCGCAGGTGCGGTTTGTG
GGCCTGCTGATATAGTGGACGAAATGAGAGACCTTCACGGAGGGCCTTTGATGTTTCTTGGCCCAACAATGAATCCCAAGCTAGCCTTCGAGATATCAGG
GAGAATCCCTCACCTGAGCATGAGGATGAAGAAACATTGCAAACGAGCAATGAAATATGTCACCAAGATGAAGGAACTGGACCCTAACCTAGATGTCTTA
TACCCCGGGCTTAAAGAGCATCCACAGCACAGTCTCTTGAAGTCCATGTCTAATAAGGATTATGGGTTCGGTGGGGTTTTCTGCATTGACATGAAAACTG
AAGGAAGGGCCTATCGCTTGATGGACAAATTGCAAAACAGCACCAAATTTGGGTTCATGGGCGTTAGCTTGGGCTATTATGAGACCCTCATGACTTGCTC
TTCCAGGAGCACTAATAGTATGATGGACAGCAAGGAAAAGGAAAGTGCCGGCATTTCACCAGGGCTGGTAAGATTTTCGGTGGGATACGTGGGGACTTTT
GAGCAGAAATGGAGCCAATTCACCAAGGCATATTCAGAAATGTAA
AA sequence
>Potri.003G187032.2 pacid=42786039 polypeptide=Potri.003G187032.2.p locus=Potri.003G187032 ID=Potri.003G187032.2.v4.1 annot-version=v4.1
MILSPAKLGADVVVHSISKYISGEADIIAGAVCGPADIVDEMRDLHGGPLMFLGPTMNPKLAFEISGRIPHLSMRMKKHCKRAMKYVTKMKELDPNLDVL
YPGLKEHPQHSLLKSMSNKDYGFGGVFCIDMKTEGRAYRLMDKLQNSTKFGFMGVSLGYYETLMTCSSRSTNSMMDSKEKESAGISPGLVRFSVGYVGTF
EQKWSQFTKAYSEM
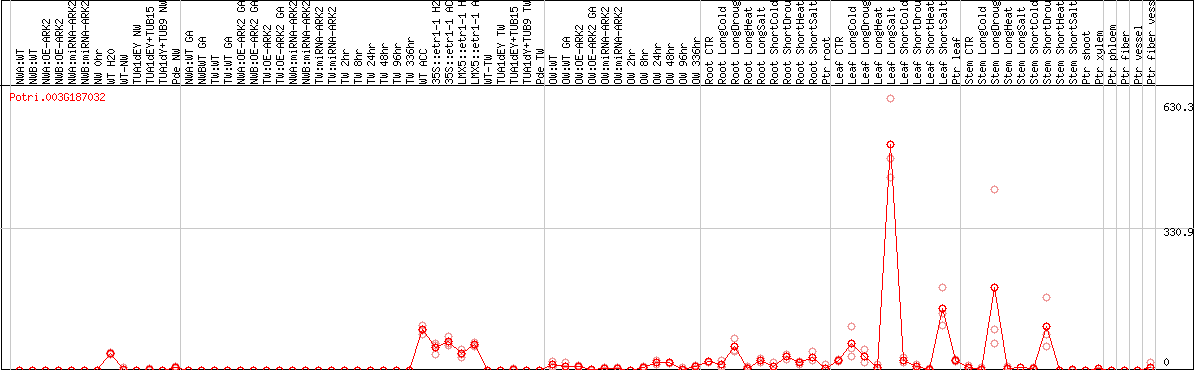
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G187032 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.