External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G33720 109 / 4e-28
AP2/B3-like transcriptional factor family protein (.1)
AT1G78640 92 / 6e-21
unknown protein
AT4G02870 74 / 5e-15
unknown protein
AT1G13260 53 / 5e-08
AP2_ERF
EDF4, RAV1
ETHYLENE RESPONSE DNA BINDING FACTOR 4, related to ABI3/VP1 1 (.1)
AT1G25560 53 / 6e-08
AP2_ERF
EDF1, TEM1
TEMPRANILLO 1, ETHYLENE RESPONSE DNA BINDING FACTOR 1, AP2/B3 transcription factor family protein (.1)
AT1G68840 53 / 8e-08
AP2_ERF
EDF2, RAV2, RAP2.8, TEM2, AtRAV2
TEMPRANILLO 2, RELATED TO AP2 8, ETHYLENE RESPONSE DNA BINDING FACTOR 2, related to ABI3/VP1 2 (.1.2)
AT3G25730 51 / 3e-07
AP2_ERF
EDF3
ethylene response DNA binding factor 3 (.1)
AT4G01500 49 / 2e-06
B3
NGA4
NGATHA4, AP2/B3-like transcriptional factor family protein (.1)
AT5G26805 47 / 2e-06
unknown protein
AT4G03170 45 / 2e-05
B3
AP2/B3-like transcriptional factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G191901
448 / 1e-161
AT2G33720 119 / 7e-32
AP2/B3-like transcriptional factor family protein (.1)
Potri.004G099600
242 / 3e-81
AT2G33720 112 / 7e-30
AP2/B3-like transcriptional factor family protein (.1)
Potri.004G045800
50 / 4e-07
AT1G28300 201 / 2e-60
LEAFY COTYLEDON 2, AP2/B3-like transcriptional factor family protein (.1)
Potri.010G129200
50 / 7e-07
AT1G25560 441 / 1e-154
TEMPRANILLO 1, ETHYLENE RESPONSE DNA BINDING FACTOR 1, AP2/B3 transcription factor family protein (.1)
Potri.008G117100
46 / 1e-05
AT1G25560 422 / 6e-148
TEMPRANILLO 1, ETHYLENE RESPONSE DNA BINDING FACTOR 1, AP2/B3 transcription factor family protein (.1)
Potri.011G054000
46 / 1e-05
AT1G28300 154 / 5e-43
LEAFY COTYLEDON 2, AP2/B3-like transcriptional factor family protein (.1)
Potri.016G074500
44 / 4e-05
AT2G36080 219 / 9e-70
AP2/B3-like transcriptional factor family protein (.1.2)
Potri.006G186301
44 / 6e-05
AT1G51120 287 / 3e-95
AP2/B3 transcription factor family protein (.1)
Potri.001G452200
44 / 6e-05
AT2G46870 198 / 4e-60
NGATHA1, AP2/B3-like transcriptional factor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027591
94 / 2e-22
AT4G02870 75 / 2e-15
unknown protein
Lus10008543
52 / 1e-08
AT4G02870 53 / 3e-09
unknown protein
Lus10041487
49 / 2e-06
AT1G13260 416 / 8e-146
ETHYLENE RESPONSE DNA BINDING FACTOR 4, related to ABI3/VP1 1 (.1)
Lus10034276
45 / 1e-05
AT1G68840 233 / 4e-76
TEMPRANILLO 2, RELATED TO AP2 8, ETHYLENE RESPONSE DNA BINDING FACTOR 2, related to ABI3/VP1 2 (.1.2)
Lus10021707
45 / 3e-05
AT1G13260 369 / 1e-127
ETHYLENE RESPONSE DNA BINDING FACTOR 4, related to ABI3/VP1 1 (.1)
Lus10021326
42 / 0.0003
AT5G06250 202 / 2e-63
AP2/B3-like transcriptional factor family protein (.1.2)
PFAM info
Representative CDS sequence
>Potri.003G191700.1 pacid=42785261 polypeptide=Potri.003G191700.1.p locus=Potri.003G191700 ID=Potri.003G191700.1.v4.1 annot-version=v4.1
ATGGAGAAACAACAACAATTTTGTGTTTTGAGTCCTTGTTTTCCTTCCAGGAACACCAATAATCCTAAGAAAAGAAAAGAATGTGAATGTCAACAAGCCG
AGAGAGAAGTGTTTGCTGGGTCACAGCCAATTTTCAATGAGCATCCTTTTTGGTTCAAGAAGCCAAAAACAAGCAGCGGAATTGGTTCCATTGTTGTTGC
TTCAAGAAATCCAGGGAGGGTTTTAAAGGAATCAAAGCTTTTTGATGAAAACTGGGTTTTCAAGAACTCTGTTGATCCTGATTCAAGAAACAATGAATCT
GGAGAAGAGTTGGAGAGAAGGGTGGCTTTTAATCTTAGGACGATGAGATCGTCTTTTAGCCTTGGGGATTTGCTAGAGGAGAGGTCTCGTGGGGTATCAA
CTCAACTAAGTCTGTATCATCATGACCCATTCGAGATCAAGAAGAAGATGAAGCCAAGTGATCTGGGTAATCTATGCAGGCTGCTTGTGTCTGCAGATTT
GGTTGAGAAACATATCCTACCTTTCTTGAACGAGGATCAAACTAAACAGGTCTGGGTTTGGGATATCGATACTGGAAGTATGCATCAGTTGGTCTTCAAA
AGATGGAGCACTTCTAAAAGCTATATTTTCAATGATGGCTGGACAAAGCACTTCGTTAAAAGGAGGAATTTGAGGGAAAGCGATGAAATTGGACTCTACT
GGGACAATGATCAGTCAAGATTTCACTTCAGTGTTCTTAGTAGGGCTGCTGCTGTGGCCAAGATTAGAAATTAG
AA sequence
>Potri.003G191700.1 pacid=42785261 polypeptide=Potri.003G191700.1.p locus=Potri.003G191700 ID=Potri.003G191700.1.v4.1 annot-version=v4.1
MEKQQQFCVLSPCFPSRNTNNPKKRKECECQQAEREVFAGSQPIFNEHPFWFKKPKTSSGIGSIVVASRNPGRVLKESKLFDENWVFKNSVDPDSRNNES
GEELERRVAFNLRTMRSSFSLGDLLEERSRGVSTQLSLYHHDPFEIKKKMKPSDLGNLCRLLVSADLVEKHILPFLNEDQTKQVWVWDIDTGSMHQLVFK
RWSTSKSYIFNDGWTKHFVKRRNLRESDEIGLYWDNDQSRFHFSVLSRAAAVAKIRN
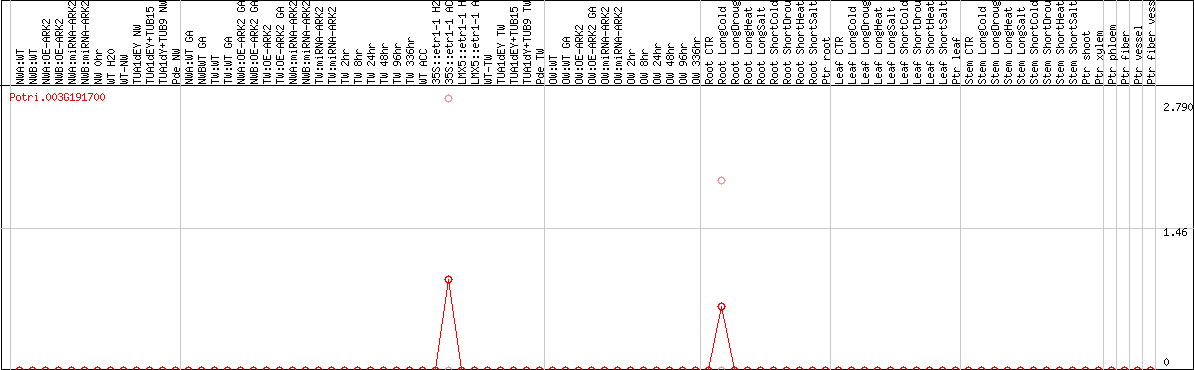
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G191700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.