External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G07010 394 / 1e-137
ATST2A
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
AT5G07000 375 / 3e-130
ATST2B
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2B, sulfotransferase 2B (.1)
AT3G45070 304 / 1e-102
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G43690 298 / 5e-100
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT3G45080 290 / 4e-97
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G13420 286 / 1e-95
SST1, ATST4B
ARABIDOPSIS THALIANA SULFOTRANSFERASE 4B, sulfotransferase 4B (.1)
AT2G03770 281 / 2e-93
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT2G03760 279 / 7e-93
AtSOT12, AtSOT1, ATST1, RAR047, ST
ARABIDOPSIS THALIANA SULFOTRANSFERASE 1, sulphotransferase 12 (.1)
AT2G14920 274 / 1e-90
ATST4A
ARABIDOPSIS THALIANA SULFOTRANSFERASE 4A, sulfotransferase 4A (.1)
AT2G03750 273 / 3e-90
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G193200
630 / 0
AT5G07010 384 / 1e-133
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Potri.003G193400
512 / 0
AT5G07010 393 / 6e-137
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Potri.003G193300
510 / 0
AT5G07010 378 / 4e-131
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Potri.003G193500
401 / 2e-140
AT5G07010 446 / 1e-157
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Potri.003G189100
362 / 2e-125
AT5G07010 295 / 1e-98
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Potri.003G188800
344 / 2e-118
AT5G07010 325 / 2e-110
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Potri.010G138501
335 / 9e-115
AT3G45070 351 / 3e-121
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.010G138450
331 / 5e-113
AT3G45070 350 / 1e-120
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.003G189066
328 / 4e-112
AT5G07010 324 / 5e-110
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008673
387 / 5e-135
AT5G07010 429 / 3e-151
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Lus10026147
380 / 3e-132
AT5G07010 425 / 2e-149
ARABIDOPSIS THALIANA SULFOTRANSFERASE 2A, sulfotransferase 2A (.1)
Lus10017878
273 / 7e-90
AT1G18590 289 / 3e-96
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
Lus10041611
240 / 7e-77
AT3G45070 239 / 9e-77
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10003070
227 / 3e-72
AT3G45070 219 / 2e-69
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10033720
214 / 7e-67
AT1G74100 229 / 6e-73
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
Lus10031624
210 / 8e-66
AT3G45070 209 / 2e-65
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10018047
207 / 2e-64
AT1G74100 208 / 7e-65
CORONATINE INDUCED-7, ARABIDOPSIS SULFOTRANSFERASE 5A, sulfotransferase 16 (.1)
Lus10034079
205 / 4e-64
AT3G45070 206 / 1e-64
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10033721
214 / 5e-64
AT1G18590 223 / 6e-67
ARABIDOPSIS SULFOTRANSFERASE 5C, sulfotransferase 17 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00685
Sulfotransfer_1
Sulfotransferase domain
Representative CDS sequence
>Potri.003G193250.1 pacid=42784973 polypeptide=Potri.003G193250.1.p locus=Potri.003G193250 ID=Potri.003G193250.1.v4.1 annot-version=v4.1
ATGGAAGACACCCAAATGATCACCGGAAACAACAATAGTCTTGCAGAAACCGATGAGATTGCTGACCTCCTGTCATCCCTTCCCAAAGAAAAGTCATGGC
GATTAGGTTATCTGTATCGATACCAAGGCTTTTGGTGTCCTGAAAAGCAGATTCCTGCTGTCATTGCATTTCAAAAGCACTTCATAGCACAGAAAACAGA
TACTATACTAGTCACCATGCCTAAATCAGGCACAACATGGTTGAAAGCCTTGGCATTTTCCATTATGAATCGTGCAAAATATACACCCTCGTGTAGCCCC
TTGAACTCTGTCAACCCTCATGATCTTGTACCTTTCTTTGAGTTTAGGCTTTACGCAAATAACCAACTTCCTGACCTGTCTACCTTTCCATCCCCTAGAA
TGTTTGCTACTCATGTGCCATATCCATCACTACCGGATTCCATCAAGAACTCCGGCTGTCGAATTGTTTGTCTTTGCAGGAATCCATTTGACAACTTTAT
CTCCTTGTGGCATTTCGCCTCCAAAGCAAGACATGAAAGTCTTGGGCCATTATCTATGGAGGATTGTTTCGATAGTTTTTGCAATGGACTTGGAGGATTC
GGTCCCTTTTTTGACCACGTATTAGGGTATTGGAGAGAAAGCTTAGAGAGACCAGAGGAGGTTCTGTTTCTCACGTATGAGGACATGAAAGAAGACATTA
ATTCTCAGATGAAAAGGTTAGCTGAGTTCCTGGGCTGTCCTTTTTCCTTGGAAGAAGAGGCAGATGGGGTTGTGGAAGGAATATCAAAGTTGTGTAGCTT
CAGCAATATGAAGGACAAAGAGATCAACAAGACTGGTAAGTCTATCCCACACTTTGAGAACAAGACTCTCTTTAGAAGAGGTGAAGTAGGGGATTGGGTC
AATTACCTTTCTCCTGAGATGGTGGATCGTTTGAACAAAATCATGGAACAAAAGCTGGCTGGTTCTGGTTTGAAGTTCAAGACTGGTTTGTAG
AA sequence
>Potri.003G193250.1 pacid=42784973 polypeptide=Potri.003G193250.1.p locus=Potri.003G193250 ID=Potri.003G193250.1.v4.1 annot-version=v4.1
MEDTQMITGNNNSLAETDEIADLLSSLPKEKSWRLGYLYRYQGFWCPEKQIPAVIAFQKHFIAQKTDTILVTMPKSGTTWLKALAFSIMNRAKYTPSCSP
LNSVNPHDLVPFFEFRLYANNQLPDLSTFPSPRMFATHVPYPSLPDSIKNSGCRIVCLCRNPFDNFISLWHFASKARHESLGPLSMEDCFDSFCNGLGGF
GPFFDHVLGYWRESLERPEEVLFLTYEDMKEDINSQMKRLAEFLGCPFSLEEEADGVVEGISKLCSFSNMKDKEINKTGKSIPHFENKTLFRRGEVGDWV
NYLSPEMVDRLNKIMEQKLAGSGLKFKTGL
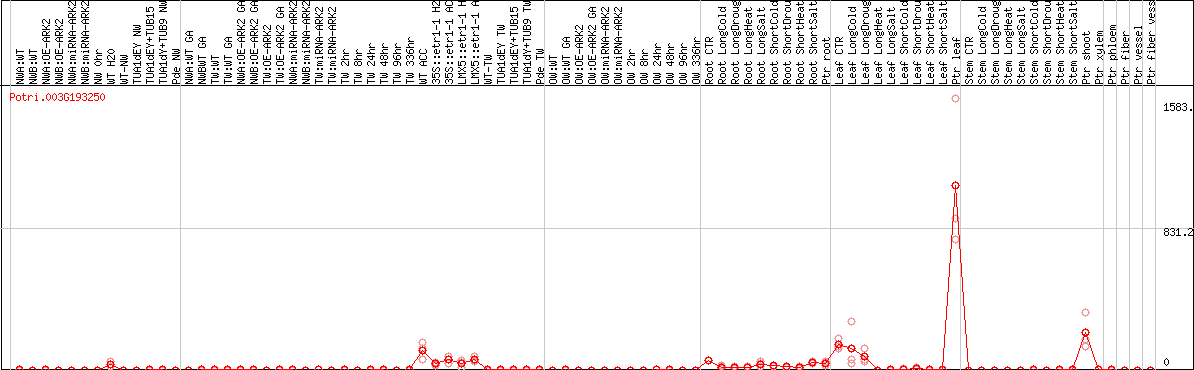
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G193250 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.