Potri.003G197100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G197100.2 pacid=42786683 polypeptide=Potri.003G197001.1.p locus=Potri.003G197100 ID=Potri.003G197100.2.v4.1 annot-version=v4.1
ATGCATGAACCAGGTTCGAATACTGATTTTCTTCTTGAATCCATTTTCATTAGTGGGGTCTGCGGATCGTTGCACTTGGTTTTGTTACTTGCTTTATGTG
TCTCGTTTTTGTGTAAGAAACTCAGCAGGTGGGGTGATGGTGAGGGTTCTTCAGAGATGTTAATGATGAAGAGGAGGTTTTTGTGGTATAAACAGACCTT
GGTCTGTTGTTTGGGTGTCTCGGTTTTTAATTTCATCTTGTGTTTGTTGAGCTACTTTTATTTGTATGGAAATGTTTTGTCTGATGGTGAAATAATGACC
CTTTTGGATTTAGGGCTGAAAACACTCTCGTGGGGTGCACTTGTTGTTTATTTGCATACCCAATTCTTTAACTCAGGTGAAAAAATGTTCCCTTTGTCAT
TGAGAGTTTGGTGGGGTTTCTATTTGGCTATTTCTTGTTATTGCTTTGTTGTAGATGTTTTCCTTCATCGCAAACACGGGTCTTTGGAGATAGAGTGGTG
TTTAGTGTCTGATGTGGTCTCTGTGTTTTCTGGATTGTTTCTTTGTTATGTTGGGTTTTTGAGAAGTGATATTCAAGATGTACTTGGAGAACCCCTTTTG
AATGGTGATTCTAGTTCGATTAATAATTTGGAGACGAGTAATTCTAGGGGGGGAGATACGGTGACTCCTTTTGGAAATGCTGGTCTTTTCAGTATTCTCA
CCTTCTCTTGGATGAATTCTTTGATTGCTGCTGGCAATAAGAAAACTTTAGACCTTGAAGATGTTCCTCAACTTCACGGTGTTGACAGTGTGGTTGGAGC
TTTTCCAGTTTTTAAAAATAAGCTTGAGTCAGATTGTGGTAGAGTTACCGGGTTCAAGCTCGCGAAAGCACTGTTCCTGTTAGTCTGGAAAGAAATTTTA
AAGACAGCTTTATTGGCGTTGATATGCACGTTATGTTCTTTTGTTGGTCCCTACCTTATCGACGCTTTTGTTCAATGCCTTGAAGGGCGAGGAGAATTCA
AGAATCAAGGTTATATTTTAGCTTCCACCTTTGTGGCTGCAAAGCTTGCTGAGTGCCTCGCACACAGGCATTCATCTTTTAGGTTGCAACAAATTGGAAC
AAGGCTCCGCGCAGTAACAGCCACGATGATTTACAACAAGAGCTTGACCATTTCCTGTCAGTCAAAGCAGGGGCATTCAAGTGGAGAAATGATCAATATT
ATGACAATTGATGCTGATAGACTTGGCACCTTTAGTCAGTTCATACACGATCCATGGTTGGTCATCTTGCAAGTTTGTTTGGCCCTGCTGATACTGTACA
GAAATTTGGGACTGGGGTCGGTTGCTGGCTTTGTTGCGACAGGAATTGTCATGTCGTTAAACTATCCTTTCGGAAGATTAGAGGAGAAGTTTCAAGACAA
GTTAATGGAATCAAAAGATAAACGAATGAAGGCAACCGTGGAGATTTTGAGAAATATGAGAGTTCTCAAGCTTCAGGGATGGGAAATGAAGTTTTTGTCT
AAGATTCTTGACCTCAGGGAAGTAGAGACACGGTGGCTGAAGAAATACTTCTATAATTCAGTAGTGATCACTGTTGTCTGCTGGGCTACTCCCACTGTTG
TGGCTGTGGCCACCTTCGGTACTTGCATGCTAATGGGAATCCCTCTTGAATCAGGGAAGGTCTTATCTGCACTCGCAACATTCGAGATTCTTCAATCCCC
AATCTATAACCTTCCCGATACAGTTTCTATGCTAATTCAGACAAAGGTTTCTCTCGACAGAATTGCATCTTTTCTTTGTCTTGATGACTTGCAGCCTGAT
GCTATAGAGAAGCTTCCAGGAGGCAGTTCTGATACAGCAATTGAGATTGTTGATGGAAATTTCTCTTGGGATTTGTCTTCACCCAGCGCAACATTGAAAG
ATATAAACTTCAAAGTGCTTAATGGTATGAAGGTAGCTGTTTGTGGTACTGTTGGTTCAGGCAAGTCGAGCTTGCTTTCCTCCATTTTGGGAGAGTTACC
CAAGATCTCAGGGACACTCAAGTTGTGTGGGACAAAGGCTTATGTTGCTCAGTCACCTTGGATACAGAGTGGTACGATAGAAGAGAACATATTGTTTGGT
AAGGTGATGGACAGAGAAAGGTACGATAAGGTACTTGAAGCATGTTCCCTGAAGAAGGACCTGGAAATCCTCTCATTCGGTGATCAAACGGGTATTGGTG
AGAGGGGTATCAATTTGAGTGGTGGACAGAAGCAAAGAATACAGATAGCACGTGCTCTCTACCAAGATGCTCAAATCTATCTATTTGATGACCCTTTCAG
TGCCGTGGATGCTCATACTGGATCTCATTTGTTTAAAGAAGTTTTGCTTGGTCTTTTAAGTTCAAAAACTGTTATTTATGTTACTCATCAGGTTGAGTTC
TTATCCGCTGCTGATCTAATCGTGGTCATGAAAGATGGAAGGATCGCGCAGGCAGGGAAGTATGACGACATTCTCAATGCAGGATCTGATTTTAAGGTAC
TTGTAGGTGCACTCAAGACAGCTTTATCAGTTCTTGATTCCAGACATGCAGGACCAGTGTCCGAAAATGAAAGTGTCAGAGACAACAATGGAGGCGAGAA
CAGTACTGATAGGATTGTACATAACGAAGGAAATAAAGATTCACAAATTGGTAAAGCAGATGAGGTAGCCGAGCCACAAGCACAACTCATTCAAGAAGAA
GAGAGAGAGAAAGGCAGTGTTGGGTTTCAAATCTATTGGAAATATATCACAATAGCATACGGAGGCGCTCTGGTCCCCTTTATATTACTGGCACAACTTC
TCTTTCAGATCCTTCAAATTGGTAGCACTTATTGGATGGCATGGGCAACTCCTGTGTCAAAGGATGTGAAACCTGGTGTTAGTGGATCTAGACTGTTAAT
CGTCTATGTATCTCTGGTTATTGGAAGTTCTTTTTGCATGCTCGCTCAAGCCATGCTTTTTGTAACAGCAGGGTACAAAACTGCAACTCTACTATTCAAC
AAATTGCATCTGTGCATTTTTCGTGCTCCCATGTCCTTTTTTGAGGCAACCCCTAGCGGGCGAATCATTAACAGAGCTTCTACAGATCAAAGTGCATTAG
ATATGAAAATTCCACATACAGTTGAGGGCCTTGCCTTCGAAGCAATCATGCTTCTGGGAATCATAGCAGTGATGTCTCAAGTGGCATGGCAAGTTTTTAT
AGTTTCCATCCCCGTAATTGCTGCCTGTATCTGGTATCAGCAATACTACATTCCCTCAGCAAGAGAGCTTTCACGGTTGATTGGAGTATGCAACGCTCCA
GTTATCCAAAATTTTGCAGAAACAATTTCAGGAGCAACAACGATCAGGAGCTTTGATCAAGAATCAAGATTCGAAGAAATAAATATGAAACTAACAGATG
CATATTCTCGGCCCAAATTTCATAATTCTGCTGCAATGCAATGGCTTTGCTTCCGAATGGATATGTTCAGTTCTATTACGTTTGCTTTCTGTCTGTTTCT
TTTAGTCTCTTTTCCAGAAAGAACTAATCCAGCCATTGCAGGCTTAGCTGTTATGTATGCGCTTGAGCTACACAGGGCACAATTTGGGTTGATATGGTGT
TTCTGCGATTGCGAGAACGAACTCATATCAGTAGAGAGAATTCTTCAGTACATTTCTATTCCTGCCGAGCCTCCTCTTGTAATTGAAGCGAACAGGCCTG
ACCATTCTTGGCCATCACATGGTGAAGTTGACATCGATAATCTGCAGGTCCGATATGCCCCACACATGCCACTTGTGCTGCGTGGTCTCTCATGCACTTT
CCCAGGAGGGAAGAAAACTGGTATTGTTGGGAGAACAGGCAGTGGTAAATCTACTCTCATACAGGCTCTTTTCCGAACTGTTGAACCTGCAGCTGGTCAG
ATTATGATTGACAGCATTGATATCTCATTGATTGGCCTGCATGATTTAAGGTCAAGACTGAGCATTATTCCCCAGGATCCAACCATGTTCGAAGGGACTG
TTAGAAGTAATCTGGACCCACTTGAAGAGTACACAGATGAACAGATCTGGGAGGTTCTGGACAAGTGCCAACTTGGAGATGAAGTTAGGAAAAAGGAAAG
GAAGCTAGATTCCACAGTTATCGAGAATGGGGAGAACTGGAGTATGGGCCAGAGGCAGCTAGTCTGTCTTGGGCGTGTGCTACTCAAGAAAAGTAAGGTG
TTGGTCCTTGATGAAGCTACTGCCTCAGTTGACACAGCCACTGATAATCTTATTCAGCAAACGCTCAGGAAGCACTTCTCTGACTGTACAGTGATAACCA
TTGCACATAGAATAACCTCTGTTCTTGACAGTGACATGGTTTTGCTTCTAAGTCAAGGTCTCATTGAGGAGTATGATTCTCCAACAAGGTTGTTAGAAAA
CAAGTCATCGTCATTCTCCCAACTTGTAGCAGAGTACACAGTGAGGTCAAATACCAGTTTCGAGAAATCAACTGGATTAAATTTATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G197100.2 pacid=42786683 polypeptide=Potri.003G197001.1.p locus=Potri.003G197100 ID=Potri.003G197100.2.v4.1 annot-version=v4.1
MHEPGSNTDFLLESIFISGVCGSLHLVLLLALCVSFLCKKLSRWGDGEGSSEMLMMKRRFLWYKQTLVCCLGVSVFNFILCLLSYFYLYGNVLSDGEIMT
LLDLGLKTLSWGALVVYLHTQFFNSGEKMFPLSLRVWWGFYLAISCYCFVVDVFLHRKHGSLEIEWCLVSDVVSVFSGLFLCYVGFLRSDIQDVLGEPLL
NGDSSSINNLETSNSRGGDTVTPFGNAGLFSILTFSWMNSLIAAGNKKTLDLEDVPQLHGVDSVVGAFPVFKNKLESDCGRVTGFKLAKALFLLVWKEIL
KTALLALICTLCSFVGPYLIDAFVQCLEGRGEFKNQGYILASTFVAAKLAECLAHRHSSFRLQQIGTRLRAVTATMIYNKSLTISCQSKQGHSSGEMINI
MTIDADRLGTFSQFIHDPWLVILQVCLALLILYRNLGLGSVAGFVATGIVMSLNYPFGRLEEKFQDKLMESKDKRMKATVEILRNMRVLKLQGWEMKFLS
KILDLREVETRWLKKYFYNSVVITVVCWATPTVVAVATFGTCMLMGIPLESGKVLSALATFEILQSPIYNLPDTVSMLIQTKVSLDRIASFLCLDDLQPD
AIEKLPGGSSDTAIEIVDGNFSWDLSSPSATLKDINFKVLNGMKVAVCGTVGSGKSSLLSSILGELPKISGTLKLCGTKAYVAQSPWIQSGTIEENILFG
KVMDRERYDKVLEACSLKKDLEILSFGDQTGIGERGINLSGGQKQRIQIARALYQDAQIYLFDDPFSAVDAHTGSHLFKEVLLGLLSSKTVIYVTHQVEF
LSAADLIVVMKDGRIAQAGKYDDILNAGSDFKVLVGALKTALSVLDSRHAGPVSENESVRDNNGGENSTDRIVHNEGNKDSQIGKADEVAEPQAQLIQEE
EREKGSVGFQIYWKYITIAYGGALVPFILLAQLLFQILQIGSTYWMAWATPVSKDVKPGVSGSRLLIVYVSLVIGSSFCMLAQAMLFVTAGYKTATLLFN
KLHLCIFRAPMSFFEATPSGRIINRASTDQSALDMKIPHTVEGLAFEAIMLLGIIAVMSQVAWQVFIVSIPVIAACIWYQQYYIPSARELSRLIGVCNAP
VIQNFAETISGATTIRSFDQESRFEEINMKLTDAYSRPKFHNSAAMQWLCFRMDMFSSITFAFCLFLLVSFPERTNPAIAGLAVMYALELHRAQFGLIWC
FCDCENELISVERILQYISIPAEPPLVIEANRPDHSWPSHGEVDIDNLQVRYAPHMPLVLRGLSCTFPGGKKTGIVGRTGSGKSTLIQALFRTVEPAAGQ
IMIDSIDISLIGLHDLRSRLSIIPQDPTMFEGTVRSNLDPLEEYTDEQIWEVLDKCQLGDEVRKKERKLDSTVIENGENWSMGQRQLVCLGRVLLKKSKV
LVLDEATASVDTATDNLIQQTLRKHFSDCTVITIAHRITSVLDSDMVLLLSQGLIEEYDSPTRLLENKSSSFSQLVAEYTVRSNTSFEKSTGLNL
|
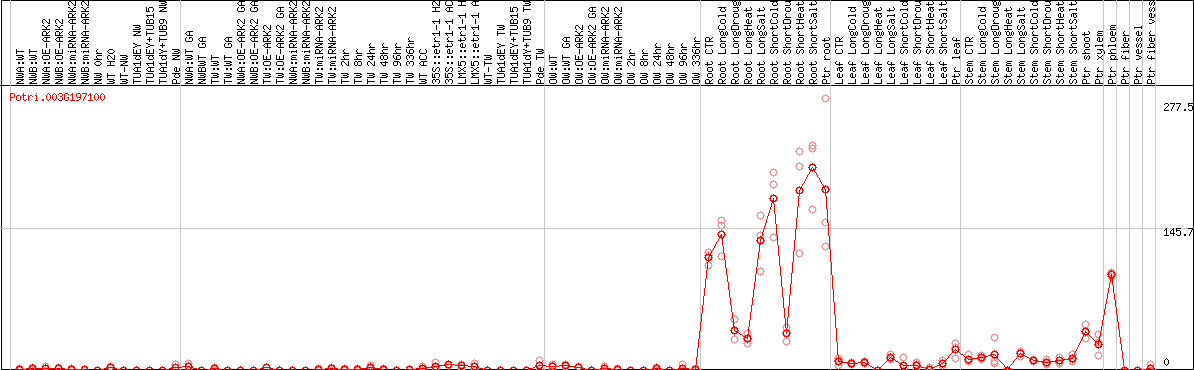
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G197100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
