Potri.003G197200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G197200.2 pacid=42787408 polypeptide=Potri.003G197200.2.p locus=Potri.003G197200 ID=Potri.003G197200.2.v4.1 annot-version=v4.1
ATGTTAATGATGAAGAGGAGGTTTTTGTGGTATAAACAGACCTTGGTCTGTTGTTTGGGTGTCACGGTTTTTAATTTCATCTTGTGTTTGTTGAGCTACT
TTTATTTGTATGGAAATGTTTTGTCTGATGGTGAAATAATGACCCTTTTGGATTTAGGGATTAGAACACTCTCTTGGGGTGCACTTGTTGTTTATTTGCA
TACCCAATTCTTTAACTCAGGTGAAAACATGTTCCCTTTGTTATTGAGAGTTTGGTGGGGTTTCTATTTGGCTATTTCTTGTTATTGCTTTCTTGTAGAT
GTTTTCATTCATCACAAACACGGGTCTTTGGAGATAGAGTGGTATTTGGTGTCTGATGCGGTCTCTGTGCTTACTGGATTGTTTCTTTGTTATGTTGGGT
TTTTGAGAAGCGATATTCAAGATGTACTTGGAGAACCCCTTTTGAATGGTGATTCTAGTTCGATTAATAATTTGGAGACAAGTAATTCTAGGGGGGGAGA
TACGGTGACTCCATTTGGAAATGCTGGTCTTTTCAGTATTCTCACCTTCTCTTGGATGAATTCTTTAATTGCTGCTGGCAATAGGAAAATTTTAGACCTT
GAAGATGTTCCTCAACTTCACGGTGTTGACAGTGTGGTTGGAGCTTTTCCAGTTTTTAAAAATAAGCTTGAGTCAGATTGTGGTAGAGCTACCAGGTTCA
AGCTCGCGAAAGCACTGTTCCTGTTAGTCTGGAAAGAAATTTTAAAGACAGCTTTATTGGCGTTGACACACACGTTATGTTCTTATGCTGGTCCCTACCT
TATCGACGCTTTTGTTCAATGCCTTGAAGGGCGAGGAGAATTCAAGAATCAAGGTTATATTTTGGCTTCCACCTTTGTGGCTGCAAAGCTTGCTGAGTGC
CTCGCACACAGGCATTTATCTTTTAGGTTGCAACAAATTGGAACAAGGCTCCGTGCAGCAACAGCCACGATGATTTACAACAAGAGCTTGACCATTTCCT
GTCAGTCAAAGCAGGGGCATTCAAGTGGAGAAATGATCAATATTATGACAATTGATGCTGATAGACTTGGCATCTTTAGTTGGTACATACACGATCCATG
GTTGGTCATCTTGCAAGTTTGTTTGGCCTTGCTGATACTGTACAGAAATTTGGGACTCGGGTCAGTTGCTGGCTTTGTTGCGACAGTAATTGTCATGTCG
TTAAACTATCCTTTCGGAAGATTAGAGGAGAAGTTTCAAGACAAGTTAATGGAATCAAAAGATAAACGAATGAAGGCAACCGTGGAGATTTTGAGAAATA
TGAGAATTCTCAAGCTTCAGGGATGGGAAATGAAGTTTTTGTCTAAGATTCTTGAACTCAGGGAAGTAGAGACACGGTGGCTGAAGAAATACTTCTATAA
TTCAGTAGTGATCACTGTTGTCTCTTGGGCTACTCCCACTGTTGTGGCTGTGGCCACCTTCGGTACTTGCATGCTAATGGGAATCCCGCTTGAATCAGGG
AAGGTCTTATCTGCACTCGCAACATTTGGGATTCTTCAATCCCCAATCTATAACCTTCCCGATACAGTTTCTATGCTAATTCAGACAAAGGTTTCTCTCG
ACAGAATTGCATCTTTTCTTTGTCTTGATGACTTGCAGCCTGATGCTATAGAGAAGCTTCCAGGAGGCAGTTCTGATACAGCAATTGAGATTGTTGATGG
AAATTTCTCTTGGGATTTGTCTTCACCCAGCGCAACATTGAAAGATATAAACTTCAAAGTGCTGAATGGTATGAAGGTAGCTGTTTGTGGTACTGTTGGT
TCAGGCAAGTCGAGCTTGCTTTCCTCCATTTTGGGAGAGTTACCCAAGATCTCAGGGACACTCAAGTTGTGTGGGACAAAGGCGTATGTTGCTCAGTCAC
CTTGGATACAGAGTGGTACGATAGAAGAGAACATATTGTTTGGTAAGGAGATGGACAAAGAAAGGTACGATAAGGTACTTGAAGCATGTTCCCTGAAGAA
GGACCTGGAAATCCTCTCATTCGGGGATCAAACAGTTATTGGTGAGAGGGGTATCAACTTGAGTGGTGGACAGAAGCAAAGAATACAGATAGCACGTGCT
CTCTACCAGGATGCCCAAATCTATCTATTTGATGACCCTTTCAGTGCCGTGGATGCTCATACTGGATCTCATTTGTTTAAAGAAGTTTTGCTTGGTCTTT
TAAGTTCAAAAACTGTTATTTATGTTACTCATCAGGTTGAGTTCTTATCCGCTGCTGATCTAATCTTGGTCATGAAAGATGGAAGGATCGCGCAGGCTGG
GAAGTATGACGAAATTCTCAATTCAGGATCTGATTTTAAGGTACTTGTAGGTGCACATAAGGCAGCTTTATCAGTTCTGGATTCCAGACAGGCAGGAGCA
GTTTCCGAAAATGAAAGTGTCAGAGACAACAATGGAGGGGAGAACAGTACTGATAGGATTGTACATGACGAAGGAAATAAAGATTCACAAATTGGTAAAG
CAGATGACGTAGCCGAGCCACAAGCACAACTCATTCAAGAAGAAGAGAGAGAGAAAGGCAGTGTTGGGTTTCAAATCTATTGGAAATATATCACAACAGC
ATACGGAGGAGCTCTGGTGCCCTTTATATTACTGGCACAACTTCTCTTTCAGATCCTTCAAATTGGTAGCACTTATTGGATGGCATGGGCAACTCCTGCG
ACAAAGGATGTGAAACCTGGTGTTAGTGGATCTAGACTGTTAATCGTCTATGTATCTCTGGTTATTGGAAGTTCTTTTTGCATCCTGGCTCGAGCCATGC
TTCTTGTAACAGCAGGGTACAAAACTGCAACTCTACTATTCAACAAATTGCATCAGTGCATTTTTCGTGCTCCCATGTCCTTTTTTGACGCAACCCCTAG
TGGACGAATCATTAACAGAGCTTCTAAAGATCAAAGTGCACTAGAAATGCAAATTCCAGATCTAGTTGGGGGTCTTGCCTTCGAAGCAATCATGCTTCTG
GGAATCATAGCAGTGATGTCTCAAGTGGCATGGCAAGTTTTTATAGTTTCCATCCCTGTAATTGCTGCCTGTATCTGGTATCAGCAATATTACATACCTG
CAGCAAGAGAGCTTTCACGGTTGATTGGAGTATGCAATGCTCCAGTTATCCAAAATTTTGCAGAAACAATTTCAGGAGCAACAACGATCAGGAGCTTTGA
TCAAGAATCAAGATTCCAAGAAATAAATATGAAACTAACAGATGCATATTCTCGGCCCAAATTTCATAATTCTGCTGCAATGCAATGGCTTTGCTTCCGC
ATGGATATGTTCAGTTCTGTTACGTTTGCTTTCTGTCTGTTTCTTTTAGTCTCTTTTCCAGAAAGAACTAATCCAGCCATTGCAGGCTTAGCTGTTACAT
ATGCGCTTGAGCTACACATGGCACAATTTGGGTTGATATGGAATTTCTGCATTTGTGAGAACAAACTCATATCAGTAGAGAGAATTCTTCAGTACATGTC
TATTCCTGCTGAGCCTCCTCTTGTAATTGAATCAAAGAGGCCTGACCATTCTTGGCCATCACATGGTAAAATTGATATCGATAATCTACAGGTCCGATAT
GCCCCACACATGCCACTGGTGCTGCGTGGTCTCTCATGCACTTTCCCAGGAGGGAAGAAAACTGGTATTGTTGGGAGAACAGGCAGCGGTAAATCTACTC
TCATACAGGCTCTTTTCCGAACCGTTGAACCTGCAGCTGGTCAGATTATGATTGACAGCATTGATATCTCATTGATTGGCCTGCATGATTTAAGGTCAAG
ACTGAGCATTATTCCCCAGGATCCAACCATGTTCGAAGGGACTGTGAGAAGTAATCTGGACCCACTTGAAGAGTACACAGATGAACAGATCTGGGAGGTT
CTGGACAAGTGCCAACTTGGAGATGAAGTTAGGAAAAAGGAAAGGAAGCTAGATTCCACAGTTATCGAGAATGGGGAGAACTGGAGTATGGGCCAGAGGC
AGCTAGTCTGTCTTGGGCGTGTGCTCCTCAAGAAAAGTAAGGTCTTGGTCCTTGATGAAGCTACTGCCTCAGTTGACACAGCCACTGATAATCTTATTCA
GCAAACGCTCAGGCAGAACTTCTCTGACTGTACAGTGATAACCATTGCACATAGAATAACCTCTGTTCTTGACAGTGACATGGTTTTGCTTCTAAGTCAA
GGTCTCATTGAGGAATATAATTCGCCAACAAGGTTGTTAGAAAACAAGTCATCGTCATTCTCCCAACTTGTAGCAGAGTACACAGTGAGGTCAAATACCC
GTTTCGAGAAATCAACTGGATTAAATTTATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G197200.2 pacid=42787408 polypeptide=Potri.003G197200.2.p locus=Potri.003G197200 ID=Potri.003G197200.2.v4.1 annot-version=v4.1
MLMMKRRFLWYKQTLVCCLGVTVFNFILCLLSYFYLYGNVLSDGEIMTLLDLGIRTLSWGALVVYLHTQFFNSGENMFPLLLRVWWGFYLAISCYCFLVD
VFIHHKHGSLEIEWYLVSDAVSVLTGLFLCYVGFLRSDIQDVLGEPLLNGDSSSINNLETSNSRGGDTVTPFGNAGLFSILTFSWMNSLIAAGNRKILDL
EDVPQLHGVDSVVGAFPVFKNKLESDCGRATRFKLAKALFLLVWKEILKTALLALTHTLCSYAGPYLIDAFVQCLEGRGEFKNQGYILASTFVAAKLAEC
LAHRHLSFRLQQIGTRLRAATATMIYNKSLTISCQSKQGHSSGEMINIMTIDADRLGIFSWYIHDPWLVILQVCLALLILYRNLGLGSVAGFVATVIVMS
LNYPFGRLEEKFQDKLMESKDKRMKATVEILRNMRILKLQGWEMKFLSKILELREVETRWLKKYFYNSVVITVVSWATPTVVAVATFGTCMLMGIPLESG
KVLSALATFGILQSPIYNLPDTVSMLIQTKVSLDRIASFLCLDDLQPDAIEKLPGGSSDTAIEIVDGNFSWDLSSPSATLKDINFKVLNGMKVAVCGTVG
SGKSSLLSSILGELPKISGTLKLCGTKAYVAQSPWIQSGTIEENILFGKEMDKERYDKVLEACSLKKDLEILSFGDQTVIGERGINLSGGQKQRIQIARA
LYQDAQIYLFDDPFSAVDAHTGSHLFKEVLLGLLSSKTVIYVTHQVEFLSAADLILVMKDGRIAQAGKYDEILNSGSDFKVLVGAHKAALSVLDSRQAGA
VSENESVRDNNGGENSTDRIVHDEGNKDSQIGKADDVAEPQAQLIQEEEREKGSVGFQIYWKYITTAYGGALVPFILLAQLLFQILQIGSTYWMAWATPA
TKDVKPGVSGSRLLIVYVSLVIGSSFCILARAMLLVTAGYKTATLLFNKLHQCIFRAPMSFFDATPSGRIINRASKDQSALEMQIPDLVGGLAFEAIMLL
GIIAVMSQVAWQVFIVSIPVIAACIWYQQYYIPAARELSRLIGVCNAPVIQNFAETISGATTIRSFDQESRFQEINMKLTDAYSRPKFHNSAAMQWLCFR
MDMFSSVTFAFCLFLLVSFPERTNPAIAGLAVTYALELHMAQFGLIWNFCICENKLISVERILQYMSIPAEPPLVIESKRPDHSWPSHGKIDIDNLQVRY
APHMPLVLRGLSCTFPGGKKTGIVGRTGSGKSTLIQALFRTVEPAAGQIMIDSIDISLIGLHDLRSRLSIIPQDPTMFEGTVRSNLDPLEEYTDEQIWEV
LDKCQLGDEVRKKERKLDSTVIENGENWSMGQRQLVCLGRVLLKKSKVLVLDEATASVDTATDNLIQQTLRQNFSDCTVITIAHRITSVLDSDMVLLLSQ
GLIEEYNSPTRLLENKSSSFSQLVAEYTVRSNTRFEKSTGLNL
|
DESeq2's median of ratios [POPLAR]
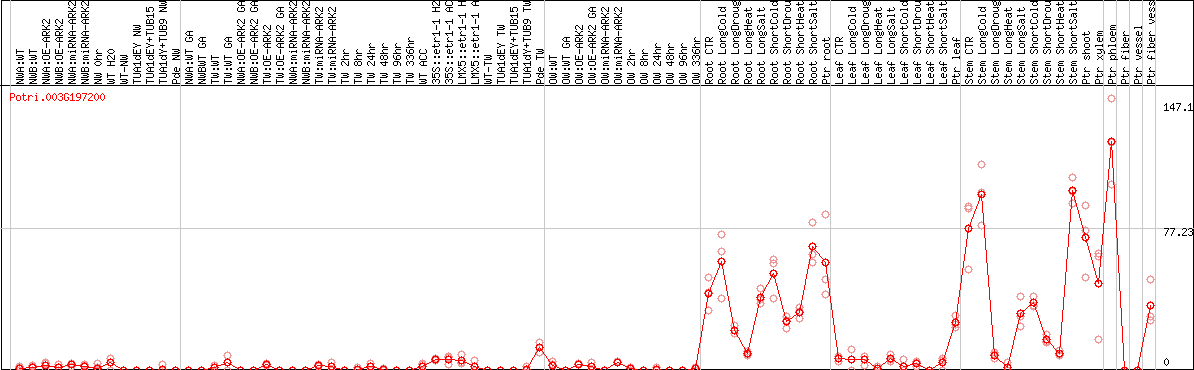
Coexpressed genes
Potri.003G197200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
