Potri.003G198401 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.003G198401.1 pacid=42785055 polypeptide=Potri.003G198401.1.p locus=Potri.003G198401 ID=Potri.003G198401.1.v4.1 annot-version=v4.1
ATGCATGCGCCAGGTTCTAACATTGATTTTCTTCTTAAATCCATTTTCATACGTGGGTTCTGTGGATCATTGCACTTGGTTTTGTTACTCGCTTTATCTG
TCTCATATGTGTGCAAGAAACTGAGCAGGAGGGGTGATGGGGAGGGCTCTAAAGAGACGTTAAAGATAAAGAGGAGGTTTATGTGGTATAAACAGACTAT
GGTCTGTTGTTTGGGTGTCTCAGTGTTTAATTTCATCTTGTGTTTGTTGAGCTACTTTTATCTGTATGGAAATGTCTGGTCTGATGGTGAAGCAATGACC
CTTTTGGATTTTGGGCTCAGAACACTCTCTTGGGGTGCACTTGTTGTTTACTTGCATACCCAGTTTTTTAACTCAGGTGAAAAGATGTTCCCTTTGTTAT
TGAGAGTTTGGTGGGGTTTCTATTTGGCTATTTCTTGTTATTGCTTTTTTGTAGATGTTTTCCTTCATCACAAACATGTGTCTTTGGAGATAGAGTGGTA
TTTAGTGTCTGATGTGGTCTCTGTGTTTACTGGATTGTTTCTTTGTTATGTTGGGTCTTTGAGAAGTGATATTCAAGATGTACTTGAAGAACCCCTTTTG
AATGGTGATTCTAGCTCGATTGATAATTTAGAGAACAGGGGGGCAGATACGGTGACTCCTTTTGGAAACGCTGGTCTTTTCAGTATTCTAACCTTTTCTT
GGATGAATTCTTTAATTGCTGCTGGCAACAAGAAAACCTTAGACCTTGAAGATGTTCCTCAACTTCACGGTGTCGATAGTGTGGTTGGAGCTTTTCCAGT
TTTTAAAAATAAGCTTGAGTCGGATTGTGGTAGAGTTACAAGGTTCAAGTTCGCAAAAGCTTTGTTCCTGTTAGTCTGGAAAGAAATTTTATGGACAGCT
TTATTGGCGTTGATTCACACGTTAAGTTCTTATGTTGGTCCCTACCTTATCGATGTTTTTGTTCAATGCCTTGACGGGCGAGGAGAATTCAAGAATCAAG
GTTACATTTTGGCTTCCGCCTTTGTGGTTGCAAAGCTCGCTGAGTGCCTCGCACAGAGGCATTTGCGCTTTAGGTTGCAGCAAATTGGAACAAGGCTCCG
CGCAGTAACAGCCACGATGATATACAACAAAAGCTTGACCATTTCCAGTCATTCGAAGCAAGGGCATTCGAGTGGACAAATAATCAATATTATGACTATT
GATGCTAATAGACTTGGCATCTTTAGTTGGTACATGCACGATCCATGGTTGGTCATCTTGCAAGTTTGTTTGGCCTTGCTGATACTGTACAGAAATTTGG
GACTCGGGTCAGTTGCTGGCTTTGTTGCGACAGTAATTGTCATGTCGTTAAACTATCCTTTCGGGAGACTGGAGGAGAAGTTTCAAGACAAGTTAATGGA
ATCAAAAGATAAACGAATGAAGGCAACCACGGAGATTTTGAGAAATATGAGAATTCTCAAGCTTCAGGGATGGGAAATGAAGTTTTTGTCTAAGATTCTT
GAACTCAGGAAAGTAGAGACACGGTGGTTGAAAAAATACTTATATACTTCAGAAGTGATCACTGTTGTCGCCTGGGTTACTCCGACTGTTGTGGCTGTGG
CCACCTTTGGTACTTGCATGCTTATGGGAGTCCCGCTTGACTCAGGGAAGGTCTTATCTGCACTTGCAACATTCGAGATTCTTCAATCCCCAATCTATAA
CCTTCCCAATACAGTTTCTATGCTAATTCAAACAAAGGTTTCTCTCGACAGAATTGCATCTTTTCTTTGTCTCGATGACTTGCAGCCTGATGCTATAGAG
AAGCTTCCAGTAGGCAGTTCTGATACAGCAATTGAGATTGTTGATGGAAATTTCTCTTGGGATTTCTCTTCACCCTGTGCAACATTGAAAGACATAAACT
TCAAAGTGTTCAATGGTATGAAGGTAGCTGTATGTGGTACTGTTGGATCAGGCAAGTCGAGCTTGCTTTCCTCCATTTTAGGAGAATTACCCAAGATCTC
AGGGACACTTAAGCTGTGCGGGACAAAGGCCTACGTTGCTCAGTCACCTTGGATACAGAGTGGTAAGATAGAAGAGAACATATTGTTTGGTAAGGAGATG
GACAGAGAAAGGTACGAGAAGGTACTTGAAGCATGTTCCCTGAAGAAGGACCTGGAAATCCTCTCATTCGGTGATCAAACAGTAATTGGTGAGAGGGGTA
TCAACTTGAGTGGTGGACAGAAGCAAAGAATACAGATAGCACGTGCTCTCTACCAAGATGCCCAAATCTATCTATTTGATGACCCTTTCAGTGCCGTGGA
TGCTCATACTGGATCTCATTTGTTTAAAGAAGTTTTGCTTGGTCTTTTAAGTTCAAAAACTGTTATTTATGTTACTCATCAGGTTGAGTTCTTATCCACT
GCTGATCTAATCTTGGTCATGAAAGATGGAAGGATCGCGCAGGCTGGAAAGTATGATGACATTCTCAATTCAGGATCTGATTTTACGGTTCTTGTTGGTG
CACATAAGGCAGCTTTATCAGTTCTTGATTCCAGACAGGCAGGACCAGTTTCCGAAAATGAAAGTGTCAGAGACAACAATGGAGGCGAGAACAGTACTGA
TGGGATTGTACATAACGAAGGAAATAAAGATTCACAAATTGGTAAAGCAGATGAGGTAGCTGAGCCACAAGCACAACTCATTCAAGAAGAAGAGAGAGAG
AAAGGCAGAGTTGGGTTTCAAATCTATTGGAAATATATCACAACAGCATACGGAGGAGCTCTGGTGCCCTTTATATTACTGGCACAACTTCTCTTTCAGA
TCCTTCAAATTGGTAGCACTTATTGGATGGCATGGGCAACTCCTGTGTCAAAGGATGTGAAACCTGTTGTTAGTGGATCTAGACTGTTAATCGTCTATGT
ATCTCTGGTTATTGGAAGTTCTTTTGGCATCCTGGCTCGAGTCATGCTTCTTGTAACAGCAGGGTACAAAACTGCAACTCTACTATTCAACAAATTGCAT
CTGTGCATTTTTCGTGCTCCCATGTCCTTTTTTGACGCGACCCCTAGTGGACGAATCATTAACAGAGCTTCTACAGATCAAAGTGCATTAGAGATGGAAA
TTCCATATATAATTGGGGAACTTGCCATCCAAGCAATCACGCTTCTGGGAATTATTGCGGTGATGTCTCAGGTGGCATGGCAAGTTTTTATGGTTTCCAT
CCCTGTGATTGCTGCCTGTATCTGGTATCAGCAATATTACATTCCCTCAGCAAGAGAGCTTTCACGGTTGATTGGAGTATGCAACGCTCCAGTTATCCAA
AATTTTGCAGAAACAATTTCAGGAGCAACAACGATCAGGAGCTTTGATCAAGAATCAAGATTCGAAGAAATAAATATGAAACTAACAGATGCATATTCTC
GGCCCAAATTTCATAATTCTGCTGCAATGCAATGGCTTTGCTTCCGAATGGATATGTTCAGTTCTATTACATTTGCTTTCTGTCTGTTTCTTTTAGTCTC
TTTTCCAGAAAAAATTAATCCAGCCATTGCAGGCTTAGCTGTTACGTATGCGCTTGGCCTACACATGGCACAATATGTGTTGATATGGTGTTTCTGCAAT
TGCGAGAACAAACTCATATCAGTAGAGAGAATTCTTCAGTACATTTCTATACCTGCCGAGCCTCCTCTTGTAATTGAAGCGAACAGGCCTGGCCATTCTT
GGCCATCACATGGCGAAATTGACATCGATAATCTGCAGGTCCGATATGCCCCACACATGCCACTTGTGCTGCGTGGTCTCTCATGCACTTTCCCAGGAGG
AAAGAAAACTGGTATTGTTGGGAGAACAGGCAGTGGTAAATCTACTCTCATACAGGCTCTTTTCCGAACCGTAGAACCTGCAGCTGGTCAGATTATGATT
GACAGCATTGATATCTCACTGATTGGCCTGCATGATTTAAGGTCAAGACTGAGCATTATTCCCCAGGATCCAACCATGTTCGAAGGGACTGTTAGAAGTA
ATTTGGACCCACTTGAAGAGTACACAGATGAACAGATCTGGGAGGTTCTGGATAAGTGCCAACTTGGAGATGAAGTTAGGAAAAAGGAAAGGAAGCTAGA
TTCCACAGTTATCGAGAATGGGGAGAACTGGAGTATGGGCCAGAGGCAACTAGTCTGTCTAGGGCGTGTGCTTCTCAAGAAAAGTAAGGTCCTGGTCCTT
GATGAAGCTACTGCCTCAGTTGACACAGCCACTGATAATCTTATTCAGCAAACGCTCAGGCAGCATTTCTCTGACTGTACAGTGATAACCATTGCACATC
GAATAACCTCTGTTCTTGACAGTGACATGGTTTTGCTTCTAAGTCATGGTCTCATTGAGGAATATGATTCTCCAACAAGGTTGTTAGAGAACAAGTCATC
GTCATTCTCCCAACTTGTAGCAGAGTACACAGTGAGGTCAAATACCAGTTTCGAGAAATCAGCTGGAATAAATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.003G198401.1 pacid=42785055 polypeptide=Potri.003G198401.1.p locus=Potri.003G198401 ID=Potri.003G198401.1.v4.1 annot-version=v4.1
MHAPGSNIDFLLKSIFIRGFCGSLHLVLLLALSVSYVCKKLSRRGDGEGSKETLKIKRRFMWYKQTMVCCLGVSVFNFILCLLSYFYLYGNVWSDGEAMT
LLDFGLRTLSWGALVVYLHTQFFNSGEKMFPLLLRVWWGFYLAISCYCFFVDVFLHHKHVSLEIEWYLVSDVVSVFTGLFLCYVGSLRSDIQDVLEEPLL
NGDSSSIDNLENRGADTVTPFGNAGLFSILTFSWMNSLIAAGNKKTLDLEDVPQLHGVDSVVGAFPVFKNKLESDCGRVTRFKFAKALFLLVWKEILWTA
LLALIHTLSSYVGPYLIDVFVQCLDGRGEFKNQGYILASAFVVAKLAECLAQRHLRFRLQQIGTRLRAVTATMIYNKSLTISSHSKQGHSSGQIINIMTI
DANRLGIFSWYMHDPWLVILQVCLALLILYRNLGLGSVAGFVATVIVMSLNYPFGRLEEKFQDKLMESKDKRMKATTEILRNMRILKLQGWEMKFLSKIL
ELRKVETRWLKKYLYTSEVITVVAWVTPTVVAVATFGTCMLMGVPLDSGKVLSALATFEILQSPIYNLPNTVSMLIQTKVSLDRIASFLCLDDLQPDAIE
KLPVGSSDTAIEIVDGNFSWDFSSPCATLKDINFKVFNGMKVAVCGTVGSGKSSLLSSILGELPKISGTLKLCGTKAYVAQSPWIQSGKIEENILFGKEM
DRERYEKVLEACSLKKDLEILSFGDQTVIGERGINLSGGQKQRIQIARALYQDAQIYLFDDPFSAVDAHTGSHLFKEVLLGLLSSKTVIYVTHQVEFLST
ADLILVMKDGRIAQAGKYDDILNSGSDFTVLVGAHKAALSVLDSRQAGPVSENESVRDNNGGENSTDGIVHNEGNKDSQIGKADEVAEPQAQLIQEEERE
KGRVGFQIYWKYITTAYGGALVPFILLAQLLFQILQIGSTYWMAWATPVSKDVKPVVSGSRLLIVYVSLVIGSSFGILARVMLLVTAGYKTATLLFNKLH
LCIFRAPMSFFDATPSGRIINRASTDQSALEMEIPYIIGELAIQAITLLGIIAVMSQVAWQVFMVSIPVIAACIWYQQYYIPSARELSRLIGVCNAPVIQ
NFAETISGATTIRSFDQESRFEEINMKLTDAYSRPKFHNSAAMQWLCFRMDMFSSITFAFCLFLLVSFPEKINPAIAGLAVTYALGLHMAQYVLIWCFCN
CENKLISVERILQYISIPAEPPLVIEANRPGHSWPSHGEIDIDNLQVRYAPHMPLVLRGLSCTFPGGKKTGIVGRTGSGKSTLIQALFRTVEPAAGQIMI
DSIDISLIGLHDLRSRLSIIPQDPTMFEGTVRSNLDPLEEYTDEQIWEVLDKCQLGDEVRKKERKLDSTVIENGENWSMGQRQLVCLGRVLLKKSKVLVL
DEATASVDTATDNLIQQTLRQHFSDCTVITIAHRITSVLDSDMVLLLSHGLIEEYDSPTRLLENKSSSFSQLVAEYTVRSNTSFEKSAGIN
|
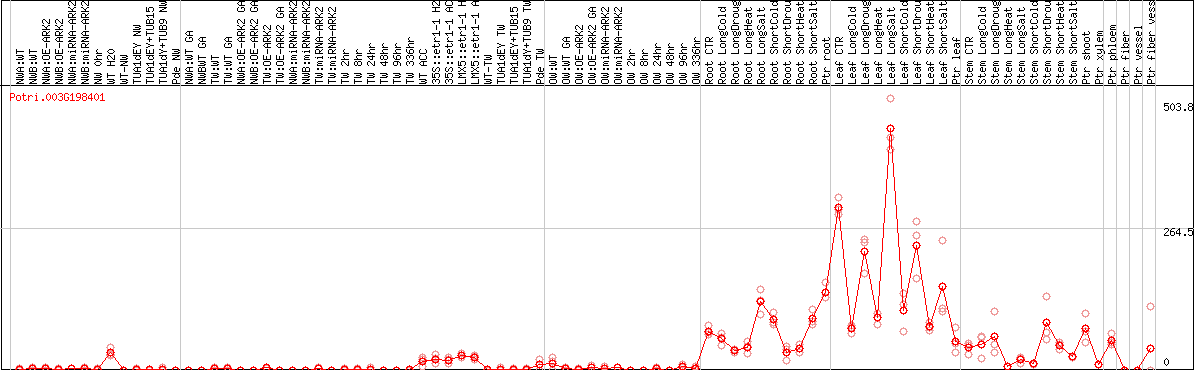
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G198401 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
