External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G16770 151 / 4e-45
bZIP
bZIP23
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT4G35040 147 / 2e-43
bZIP
bZIP19
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT3G51960 140 / 4e-41
bZIP
ATBZIP24
basic leucine zipper 24 (.1.2)
AT1G22070 46 / 1e-05
bZIP
TGA3
TGA1A-related gene 3 (.1)
AT1G77920 40 / 0.0008
bZIP
bZIP transcription factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G020200
287 / 2e-98
AT2G16770 120 / 9e-33
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.004G175200
160 / 2e-48
AT4G35040 267 / 6e-90
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.009G134900
149 / 3e-44
AT4G35040 291 / 2e-99
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Potri.005G231300
41 / 0.0002
AT1G75390 172 / 2e-55
basic leucine-zipper 44 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014121
154 / 2e-46
AT2G16770 265 / 8e-90
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10027847
155 / 4e-46
AT2G16770 295 / 1e-100
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10005073
155 / 9e-46
AT4G35040 294 / 2e-99
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10016368
122 / 4e-35
AT4G35040 135 / 5e-40
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10019788
113 / 1e-30
AT2G16770 209 / 5e-68
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10019756
83 / 2e-20
AT4G35040 79 / 4e-19
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10001347
40 / 0.0004
AT1G75390 159 / 4e-50
basic leucine-zipper 44 (.1.2)
Lus10018662
40 / 0.0005
AT3G49760 82 / 2e-20
basic leucine-zipper 5 (.1)
Lus10018009
41 / 0.0006
AT1G77920 424 / 2e-148
bZIP transcription factor family protein (.1)
Lus10006489
40 / 0.0009
AT1G45249 297 / 1e-97
ABSCISIC ACID RESPONSIVE ELEMENTS-BINDING PROTEIN 2, abscisic acid responsive elements-binding factor 2 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0018
bZIP
PF07716
bZIP_2
Basic region leucine zipper
Representative CDS sequence
>Potri.003G204400.3 pacid=42786000 polypeptide=Potri.003G204400.3.p locus=Potri.003G204400 ID=Potri.003G204400.3.v4.1 annot-version=v4.1
ATGAAAGATTATAGAATGGATGATGGGGAGGTAGAGCTTTCGGATCATGTTTTGTTACCGGATCCCGGTACTTCTGGAAGTTTGCAGTGTTCTGGGTCTG
TGGATTCACTTCTTGATGAATTATTGAAGAATACGAGAACGTGTACTCACACTCACACTTGCAACCCTCATGGCCCCGATGCAATCCATACACATACATG
CTACCACACACACACCCAAGTCATTGCTTCTGAAGAAGATGATAACCCAGACAATAGAGAGCATTCAAGAAAAAGGCCAGCAGGAAATAGGGAAGCAGTT
CGAAAGTATAGGGAAAAGAAGAAGGCGCATACGGCTTACTTAGAAGAGGAAGTAAAGAAATTGCATTTGTTGAACCAGCAGCTGGTGAGGAAAATACAAC
GGCAGGCAATTCTTGAAGCCGAGGTTTCGAGGCTAAGAAGCATTCTGGTGGACCTGAGGGGAAAGATCGACACTGAGTTGGGTGTTTTTCCATTCCAAAA
CCATTGCAACACCACCACTGTTTTGAAGGAAGGTGAGTGTGGAGCGCAGTCCACTGGTGGGCTAATGAATCTACAGTGTCAGGCAGATTTAGCATGCTTC
CATACCCCTACAATAGTAAATTGTCAAGCTAACACAAATAATACGGCAAATGCTGAAGGGCATGCAATGGACATGGTGGAAGCCTTCCTGCCATCTGCCT
CTCAAGCTCATTAA
AA sequence
>Potri.003G204400.3 pacid=42786000 polypeptide=Potri.003G204400.3.p locus=Potri.003G204400 ID=Potri.003G204400.3.v4.1 annot-version=v4.1
MKDYRMDDGEVELSDHVLLPDPGTSGSLQCSGSVDSLLDELLKNTRTCTHTHTCNPHGPDAIHTHTCYHTHTQVIASEEDDNPDNREHSRKRPAGNREAV
RKYREKKKAHTAYLEEEVKKLHLLNQQLVRKIQRQAILEAEVSRLRSILVDLRGKIDTELGVFPFQNHCNTTTVLKEGECGAQSTGGLMNLQCQADLACF
HTPTIVNCQANTNNTANAEGHAMDMVEAFLPSASQAH
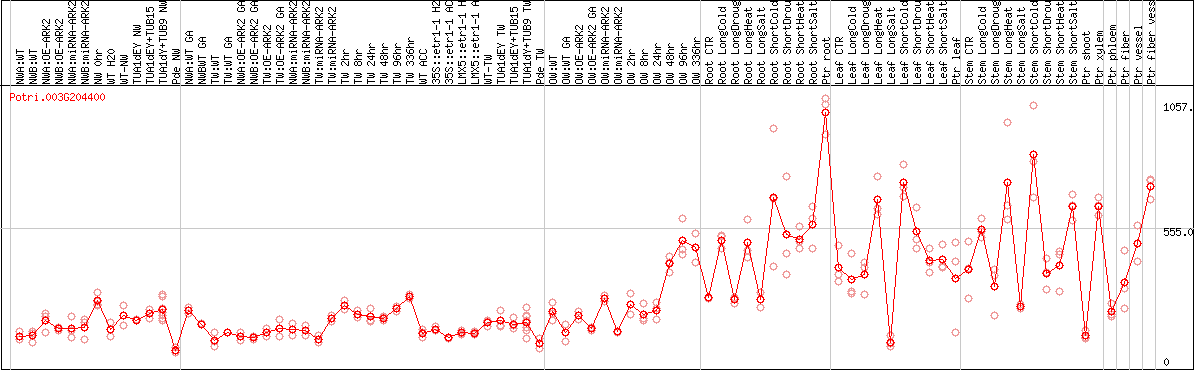
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G204400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.