External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G11950 300 / 9e-105
LOG8
LONELY GUY 8, Putative lysine decarboxylase family protein (.1.2)
AT2G28305 294 / 3e-102
ATLOG1
LONELY GUY 1, Putative lysine decarboxylase family protein (.1)
AT2G37210 289 / 3e-100
LOG3
LONELY GUY 3, lysine decarboxylase family protein (.1.2)
AT3G53450 285 / 7e-99
LOG4
LONELY GUY 4, Putative lysine decarboxylase family protein (.1)
AT2G35990 277 / 1e-95
LOG2
LONELY GUY 2, Putative lysine decarboxylase family protein (.1.2.3)
AT5G06300 273 / 4e-94
LOG7
LONELY GUY 7, Putative lysine decarboxylase family protein (.1)
AT4G35190 270 / 2e-92
LOG5
LONELY GUY 5, Putative lysine decarboxylase family protein (.1)
AT5G03270 234 / 1e-78
LOG6
LONELY GUY 6, lysine decarboxylase family protein (.1)
AT5G26140 169 / 5e-54
ATLOG9
LONELY GUY 9, Putative lysine decarboxylase family protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026131
304 / 3e-106
AT5G11950 350 / 4e-124
LONELY GUY 8, Putative lysine decarboxylase family protein (.1.2)
Lus10026513
295 / 2e-102
AT2G37210 394 / 1e-141
LONELY GUY 3, lysine decarboxylase family protein (.1.2)
Lus10016096
294 / 3e-102
AT2G28305 384 / 7e-138
LONELY GUY 1, Putative lysine decarboxylase family protein (.1)
Lus10021462
293 / 5e-102
AT2G28305 386 / 1e-138
LONELY GUY 1, Putative lysine decarboxylase family protein (.1)
Lus10002226
283 / 1e-97
AT2G37210 379 / 2e-135
LONELY GUY 3, lysine decarboxylase family protein (.1.2)
Lus10022638
279 / 6e-96
AT4G35190 365 / 2e-129
LONELY GUY 5, Putative lysine decarboxylase family protein (.1)
Lus10003335
278 / 1e-95
AT4G35190 359 / 2e-127
LONELY GUY 5, Putative lysine decarboxylase family protein (.1)
Lus10012436
277 / 2e-95
AT5G06300 332 / 7e-117
LONELY GUY 7, Putative lysine decarboxylase family protein (.1)
Lus10024328
276 / 4e-95
AT5G06300 328 / 2e-115
LONELY GUY 7, Putative lysine decarboxylase family protein (.1)
Lus10025117
276 / 8e-95
AT4G35190 373 / 1e-132
LONELY GUY 5, Putative lysine decarboxylase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0349
SLOG
PF03641
Lysine_decarbox
Possible lysine decarboxylase
Representative CDS sequence
>Potri.003G219300.1 pacid=42785677 polypeptide=Potri.003G219300.1.p locus=Potri.003G219300 ID=Potri.003G219300.1.v4.1 annot-version=v4.1
ATGGGAAAGTTCAAAAGTGTTTGTGTCTTTTGTGGAAGCAAGTCAGGGAATAAAAAGATATTCAGTGATGCAGCCCTTGATCTTGGAAGGGAGCTGGTTG
AAAGGAAGATGGATTTAGTGTATGGAGGAGGGAGTATTGGACTGATGGGGCTTGTTTCTCAGACTGTTTATGATGGTGAATGCCATGTCTTAGGTGTTAT
CCCAAGAGCTCTTGTACCGATTGAGATTTCAGGTCACACAGTTGGAGAAGTGTTGATTGTATCCGATATGCATGAAAGGAAGGCTGAGATGGCTCGGCGA
GCTGATGCTTTCATTGCTCTTCCTGGAGGGTATGGAACCTTTGAGGAGTTGCTTGAGATGATAACATGGTCTCAGCTTGGAATCCATAACAAGCCAGTGG
GTCTATTAAATGTTGATGGGTACTATGATAGCTTGCTTGGGTTTTTTGACAAAGGCGTAGAAGAAGGATTTATTGGACCTTCAGCCAGGAACATTGTCAT
ATCTGCAAGGACGGCTACAGAGCTAATCCAGAAAATGGAGGATTATATACCTCTCCATGAACAAGTAGCACCAAGTCATAGCTGGAAAGTAGAAGGATGC
AATGGAAATTTGTAG
AA sequence
>Potri.003G219300.1 pacid=42785677 polypeptide=Potri.003G219300.1.p locus=Potri.003G219300 ID=Potri.003G219300.1.v4.1 annot-version=v4.1
MGKFKSVCVFCGSKSGNKKIFSDAALDLGRELVERKMDLVYGGGSIGLMGLVSQTVYDGECHVLGVIPRALVPIEISGHTVGEVLIVSDMHERKAEMARR
ADAFIALPGGYGTFEELLEMITWSQLGIHNKPVGLLNVDGYYDSLLGFFDKGVEEGFIGPSARNIVISARTATELIQKMEDYIPLHEQVAPSHSWKVEGC
NGNL
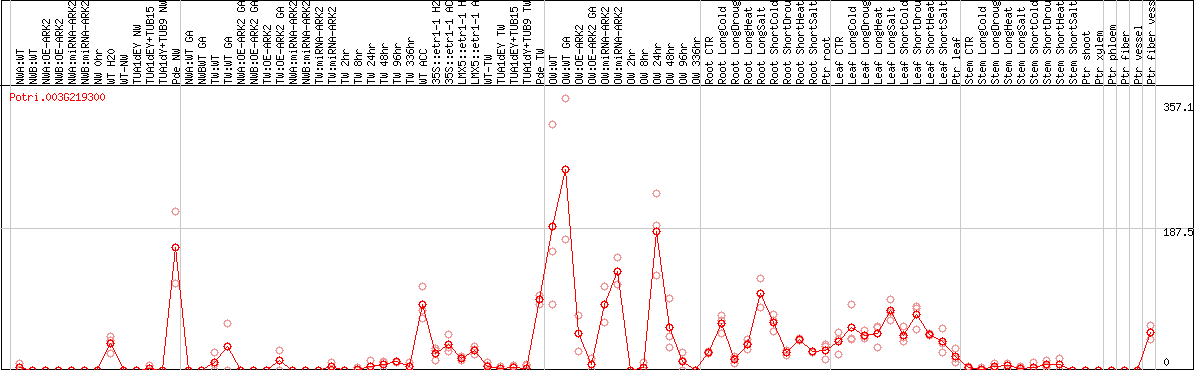
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G219300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.