External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G07100 65 / 4e-13
WRKY
WRKY26
WRKY DNA-binding protein 26 (.1.2)
AT1G29860 64 / 7e-13
WRKY
ATWRKY71, WRKY71
WRKY DNA-binding protein 71 (.1)
AT5G56270 62 / 4e-12
WRKY
ATWRKY2, WRKY2, WRKY23
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 2, WRKY DNA-binding protein 2 (.1)
AT4G18170 62 / 6e-12
WRKY
ATWRKY28, WRKY28
WRKY DNA-binding protein 28 (.1)
AT5G26170 60 / 6e-12
WRKY
ATWRKY50, WRKY50
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 50, WRKY DNA-binding protein 50 (.1)
AT4G26640 61 / 1e-11
WRKY
ATWRKY20, WRKY20
WRKY family transcription factor family protein (.1.2)
AT2G30250 61 / 1e-11
WRKY
ATWRKY25, WRKY25
WRKY DNA-binding protein 25 (.1)
AT2G03340 60 / 2e-11
WRKY
WRKY3
WRKY DNA-binding protein 3 (.1)
AT5G64810 58 / 3e-11
WRKY
ATWRKY51, WRKY51
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 51, WRKY DNA-binding protein 51 (.1)
AT1G55600 60 / 4e-11
WRKY
MINI3, ATWRKY10, WRKY10
MINISEED 3, WRKY DNA-binding protein 10 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027139
66 / 2e-13
AT5G56270 528 / 1e-179
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 2, WRKY DNA-binding protein 2 (.1)
Lus10032887
66 / 3e-13
AT5G56270 540 / 0.0
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 2, WRKY DNA-binding protein 2 (.1)
Lus10004537
61 / 9e-12
AT1G29860 188 / 1e-58
WRKY DNA-binding protein 71 (.1)
Lus10012027
61 / 1e-11
AT4G12020 102 / 2e-24
MAPK/ERK KINASE KINASE 4, MITOGEN-ACTIVATED PROTEIN KINASE KINASE KINASE 11, protein kinase family protein (.1.2.3)
Lus10020215
60 / 4e-11
AT5G56270 276 / 3e-84
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 2, WRKY DNA-binding protein 2 (.1)
Lus10016595
59 / 7e-11
AT4G26640 556 / 0.0
WRKY family transcription factor family protein (.1.2)
Lus10020832
58 / 8e-11
AT4G39410 169 / 3e-51
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 13, WRKY DNA-binding protein 13 (.1)
Lus10042243
58 / 1e-10
AT5G07100 323 / 5e-105
WRKY DNA-binding protein 26 (.1.2)
Lus10036800
58 / 1e-10
AT2G03340 405 / 4e-138
WRKY DNA-binding protein 3 (.1)
Lus10037785
58 / 1e-10
AT5G49520 179 / 4e-53
ARABIDOPSIS THALIANA WRKY DNA-BINDING PROTEIN 48, WRKY DNA-binding protein 48 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0274
WRKY-GCM1
PF03106
WRKY
WRKY DNA -binding domain
Representative CDS sequence
>Potri.003G222501.1 pacid=42785797 polypeptide=Potri.003G222501.1.p locus=Potri.003G222501 ID=Potri.003G222501.1.v4.1 annot-version=v4.1
ATGTACCCGGCCCTACCGCCCTTTAGATTCTCCATCAAAGCCACTATATACGAGATTCATCCAATGGAAACTGATCAGGAGGGTAGAGAAGAGCAGGATA
CAGATCGAGGAGGGCAGAGATTGGTGCTGCCAGAAGATGGGCATGGATGGAAGAAATATGGCCAGAAGTTCATAAAAAAAATTGGAAAATTCCGGAGCTA
TTTTAAATGCCAGAAGCAGAATTGTGTGGCAAAGAAGAGAGTCGAGTGGTCCAACCCTGATTATATTCGAATATGGTACGAGGGATCTCACAATCATGCC
TCATCTATACAAGGGACTTCTTCCTCCGCTGCTGCAAATCAGTACAGCTTGTACGCTCAAGTTTTTGGATCAGATCAACCAGCTGCTTCACGCACGCATC
ATCAAGATGCTTAA
AA sequence
>Potri.003G222501.1 pacid=42785797 polypeptide=Potri.003G222501.1.p locus=Potri.003G222501 ID=Potri.003G222501.1.v4.1 annot-version=v4.1
MYPALPPFRFSIKATIYEIHPMETDQEGREEQDTDRGGQRLVLPEDGHGWKKYGQKFIKKIGKFRSYFKCQKQNCVAKKRVEWSNPDYIRIWYEGSHNHA
SSIQGTSSSAAANQYSLYAQVFGSDQPAASRTHHQDA
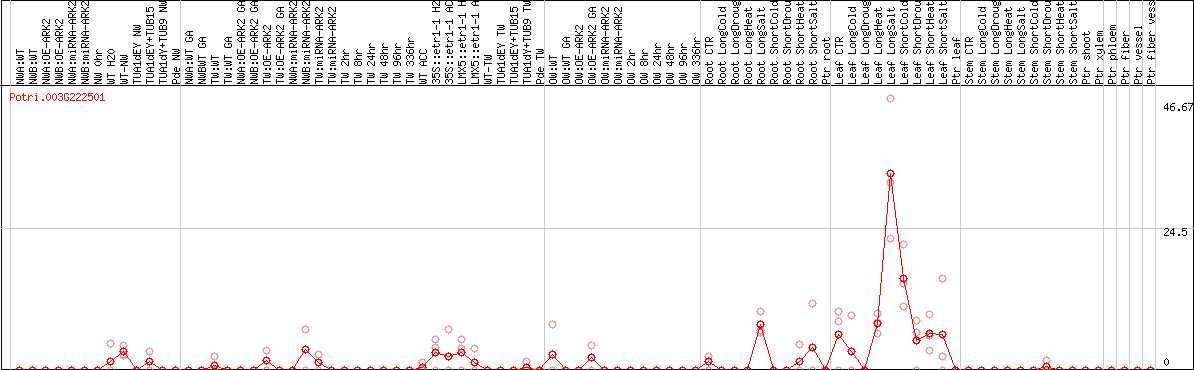
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.003G222501 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.