Potri.004G037700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G037700.4 pacid=42796160 polypeptide=Potri.004G037700.4.p locus=Potri.004G037700 ID=Potri.004G037700.4.v4.1 annot-version=v4.1
ATGATAATGGCTGACGGGAGGTCGGGGAGTGTTATGCCAGGTGGTGAAATTATGGCCGTTGGTGACGGTAATAGCGGCGATGGAGAGGAGGAGGATTTGT
CAATGGATGTCGATTTGTTTTATACAATTCTCGGAGAAGAACCGTCGCCTGCTTCTCCGAGTGGTAAAGGAGATTTTTTGGATGAGACGGTGCTGGATGC
TGGAAATAGCCGAAACTGGCTGCTCCAAAGTGGGTCTCAGAAGATAGAAGGGGCAGATGGGTTAGGGGGAGAATCCATGGATCATGCTGCCTACTCTTTG
CACAATCCAGAAGGTTCGGATGCTAGGGCAGGGCATTCGGGTGGTTCTATTGATTACACAGGGAGACAAATGAGCTCTCTTATGCATGCTCGCTCTGGAA
GTAGTAGAGAGTTTTGCTTCCCGTTTCAAGAAGATCAGGGAACAATGAGGGCAGGAGCGTCACAATCAGAGATAGCCAGTTGCATTACAGAACCTACTAC
TTTTGCTGATGGAGTGTCTAGCTTTGTTGCAGATCATGCAGGTGGTTTGAACCTAAAGCTTCTTTTGGATGATAATGGAAACCAATTGAGGCATGTGGAA
GGGAATGTTGAATCCAAAGGTTCTTCACATGGTCCTTGGATGGATGGTTTGGATGAGAAATTTGGATCACGTGATGCTTTAGACAATGATAGCTTAGGAA
TTCTAGAACTTAAAACTGATATTCATAGATCGATGGTCATGCCCTTGATGGATACAGATCATGATATGATTTCTACCAAGTCCGCTGACTGGCATTATCC
TGGTTTTAATAGCGAACTAAGGGATCATGATTCAGCTATGCAATTTGGCATGAATGGATATGATGCACATTATACTGATTCTTCAGGTTTTGATTTTTCC
AGTGATTTTAATTTTGGATTATTTCCCATTAATCAAGAAATTAATGAATTTCAACCTGAGAATGCATGCTCTGGACCAGAGATTTCGATGATGCCTTGCT
CTGATGTAAATGGGATGAATTTTAAATCTGAAGGTGATGGCTATATGTTTCCAAAGACCAGAAAATTTTCATCAAGTGCTGATGATGGACTTAACCATGA
CAAAGCATCGGTTATGCCACCCAGTGATATTCAGTTAGGCATCAGTGAGGTCCAAACAGTTTGTGTAGAGGATGAAAAAACCGATGGATTGGTTGCATGT
AGGAACATGACATGGCAGTCTGGTGAAGGTGTTACCGAAGCTGTTGACAGGAAGTGTTCTTGGAGCGATGGAAACAGTACTTTTGTTTACAAAGACAAAC
AGCAGTCACCTTCTGGTGTTTTATCATCTGTGCAAAGTCAGAAGCATGTGATTTACACCAACGATGACAGAGGGGGCATGGCTCTTGGATCTTCAAGAGC
TCAAGTTGAGGGGATTGCTGGTAGGTTTCCCTTCGATAGTGTATATTTGAATTTAAGTGCTTCAGAGCAGTATTTGCCCTTTGCTCCGACATCCCATATA
AGCAAGATGCAGCTGGGTTGTGGCAAGGATGAAAAACAAGGCTTGCCTATTCATTCTAAGGCCTTGGGTTCTCATCTTTCAATAGTCAGCCCTGAATCAA
TTCAGAGTAATTCATCTGGTAGCAAATCCCATGTTGATGATGAACCTGATATATGTATTCTTGACGATATCAGTCAACCTGCTCGCTCAAACCAATGTTT
TGCTCCTAGTAAGCCCATAGTTCCTTTGCTACACCCTACATACAATGATTCACTTCATCATTCCACAGTAGAAGGCACAAGGTTCAAGGCCAATGATGAG
CAACTTGTACTTCGAGTTGCATTGCAGGATCTTGCTCAACCAAAGTCAGAAGCTGTTCCACCCGATGGTTTTTTGGCAGTACCACTTTTGAGACACCAGC
GAATTGCTTTGTCATGGATGGTTCAGAAGGAAACATCTAGCTTGCACTGTTCTGGAGGAATTCTTGCAGATGATCAGGGATTGGGAAAGACAGTATCAAC
CATTGCGCTTATACTCAAGGAAAGAGCTCCTTTATGTCGAGTGGATGCTGTGGCTGTTAAGAAAGAAGAGTGCGAGACCTTAAATTTGGATGATGATGAC
GATGGGGTTATTGAGATTGATAGATTGAAAAAAGGTGCTGATGGTTCTCAAGTTAAGTCAAATCGAAGTTCAACAAAGAGCCTGAATTCTCCTGGGCAAT
CCAAGGGAAGGCCAGCTGCTGGAACTCTCATTGTTTGCCCCACAAGTGTCTTGCGACAGTGGGCTGATGAGCTGCATACGAAGGTAACTACTGAAGCCAA
TCTCTCTGTTCTAGTATACCATGGAAGCAATCGAACAAAAGATCCTTCCGAGGTGGCAAAGTATGATGTTGTTGTTACAACATATTCAATTGTCAGCATG
GAGGTCCCGAAGCAGCCTCTGGCTGATGAAGATGAAGAGAAACAAAGAATGGAAGGTGATGATGTTCCGCACCTAGGACTTTCATATGGTAAGAAAAGGA
AATATCCTCCCACTTCTGGCAAAAAAGGTCTGAAGAATAAGAAGGGAATGGACAGTGCAATGCTTGAGTCCATTGCCCGGCCACTTGCAAAGGTGGCATG
GTTTAGGGTTGTCCTTGATGAAGCCCAGAGCATCAAGAATCACAGAACTCAAGTGGCTAGGGCCTGTTGGGGTCTTCGGGCAAAACGCAGGTGGTGCTTG
TCCGGTACACCTATCCAAAATGCAATCGATGATCTCTATAGCTATTTCAGATTCCTCAGATATGAACCTTATGCGGTCTACAAGCTATTCTGCTCAGCAA
TAAAGGTCCCAATTCAAAAGAATCCAGCAAAAGGGTACAGAAAGTTACAAGCTGTTTTGAAGACAGTAATGCTTCGTCGAACCAAAGGCACTCTTCTTGA
CGGGGAACCTATTATTAACCTTCCACCCAAAGTAGTAGAATTGAAAAAAGTGGACTTTACAGAGGAAGAACGTGATTTCTACACCAGACTAGAAATTGAT
TCACGAGCTCAGTTTAAAGAATATGCAGCTGCTGGAACTGTTAAACAAAATTATGTTAACATCTTATTGATGCTGTTGCGTCTTCGACAAGCTTGTGATC
ATCCTCTTCTTGTCAAGGGTTTAGATTCAAATTCTCTAGGTGGATCCTCAATTGAGATGGCCAAGAAGCTTCCTCAGGAGAAACAATTATGCCTTCTCAA
ATGCTTAGAAGCATCTTTGGCAATCTGTGGCATCTGTAGTGATCCGCCTGAAGATGCTGTTGTTTCAGTCTGTGGTCATGTATTCTGCAAACAATGTATC
TGTGAGCATCTTACTGGTGACGACAACCAATGCCCTGTGTCAAACTGCAAAGTTCGCCTGAATGTGTCTTCGGTGTTTTCCAAAGCCACATTAAATAGTT
CTCTATCTGACGAGCCTGATCAAGATTCTTCTGGTTCTGGTTCTGAGCTTGTTGCTGCAGTTAGCTCATCCTCTGACAACCGTCCCCATAATTCATCAAA
AATCAGGGCAACTCTTGAAGTCCTGCAATCGCTCACTAAACCAAAAGATTGTTTGTCTAAATGCAATTTGTCAGAGAATTCTGCTGATGGAAATGTTGCT
TGTCATGAAACCTCATCTGGTTCTACAGGCTCACTTAATGATGGAACTGATAAAAGACACCCCCCAGCTAAGGTTGTTGGGGAGAAAGCCATAGTGTTTT
CCCAGTGGACAGGGATGCTAGATTTGCTTGAAGCTTGTCTTAAGAGTTCTTCCATTCAGTACAGAAGACTGGATGGAACAATGTCAGTTGTTGCCAGAGA
TAAAGCTGTGAAGGATTTCAATACCCTTCCAGAGGTATCTGTTATGATAATGTCTCTGAAAGCAGCTAGTCTTGGACTGAACATGGTTGCAGCTTGCCAT
GTGTTGCTTCTGGATCTCTGGTGGAATCCTACAACTGAGGATCAGGCAATTGATAGGGCACATCGTATTGGGCAAACTCGTAAAGTTACAGTTTTGCGAT
TAACTGTAAAAAACACTGTTGAAGATCGTATCTTAGCTCTCCAGCAAAAGAAGAGAGAGATGGTGGCATCTGCCTTTGGAGAGGATGAAAATGGTGGTCG
TCAGACTCGCCTAACCGTGGATGACTTGAATTACCTATTTATGGTGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G037700.4 pacid=42796160 polypeptide=Potri.004G037700.4.p locus=Potri.004G037700 ID=Potri.004G037700.4.v4.1 annot-version=v4.1
MIMADGRSGSVMPGGEIMAVGDGNSGDGEEEDLSMDVDLFYTILGEEPSPASPSGKGDFLDETVLDAGNSRNWLLQSGSQKIEGADGLGGESMDHAAYSL
HNPEGSDARAGHSGGSIDYTGRQMSSLMHARSGSSREFCFPFQEDQGTMRAGASQSEIASCITEPTTFADGVSSFVADHAGGLNLKLLLDDNGNQLRHVE
GNVESKGSSHGPWMDGLDEKFGSRDALDNDSLGILELKTDIHRSMVMPLMDTDHDMISTKSADWHYPGFNSELRDHDSAMQFGMNGYDAHYTDSSGFDFS
SDFNFGLFPINQEINEFQPENACSGPEISMMPCSDVNGMNFKSEGDGYMFPKTRKFSSSADDGLNHDKASVMPPSDIQLGISEVQTVCVEDEKTDGLVAC
RNMTWQSGEGVTEAVDRKCSWSDGNSTFVYKDKQQSPSGVLSSVQSQKHVIYTNDDRGGMALGSSRAQVEGIAGRFPFDSVYLNLSASEQYLPFAPTSHI
SKMQLGCGKDEKQGLPIHSKALGSHLSIVSPESIQSNSSGSKSHVDDEPDICILDDISQPARSNQCFAPSKPIVPLLHPTYNDSLHHSTVEGTRFKANDE
QLVLRVALQDLAQPKSEAVPPDGFLAVPLLRHQRIALSWMVQKETSSLHCSGGILADDQGLGKTVSTIALILKERAPLCRVDAVAVKKEECETLNLDDDD
DGVIEIDRLKKGADGSQVKSNRSSTKSLNSPGQSKGRPAAGTLIVCPTSVLRQWADELHTKVTTEANLSVLVYHGSNRTKDPSEVAKYDVVVTTYSIVSM
EVPKQPLADEDEEKQRMEGDDVPHLGLSYGKKRKYPPTSGKKGLKNKKGMDSAMLESIARPLAKVAWFRVVLDEAQSIKNHRTQVARACWGLRAKRRWCL
SGTPIQNAIDDLYSYFRFLRYEPYAVYKLFCSAIKVPIQKNPAKGYRKLQAVLKTVMLRRTKGTLLDGEPIINLPPKVVELKKVDFTEEERDFYTRLEID
SRAQFKEYAAAGTVKQNYVNILLMLLRLRQACDHPLLVKGLDSNSLGGSSIEMAKKLPQEKQLCLLKCLEASLAICGICSDPPEDAVVSVCGHVFCKQCI
CEHLTGDDNQCPVSNCKVRLNVSSVFSKATLNSSLSDEPDQDSSGSGSELVAAVSSSSDNRPHNSSKIRATLEVLQSLTKPKDCLSKCNLSENSADGNVA
CHETSSGSTGSLNDGTDKRHPPAKVVGEKAIVFSQWTGMLDLLEACLKSSSIQYRRLDGTMSVVARDKAVKDFNTLPEVSVMIMSLKAASLGLNMVAACH
VLLLDLWWNPTTEDQAIDRAHRIGQTRKVTVLRLTVKNTVEDRILALQQKKREMVASAFGEDENGGRQTRLTVDDLNYLFMV
|
DESeq2's median of ratios [POPLAR]
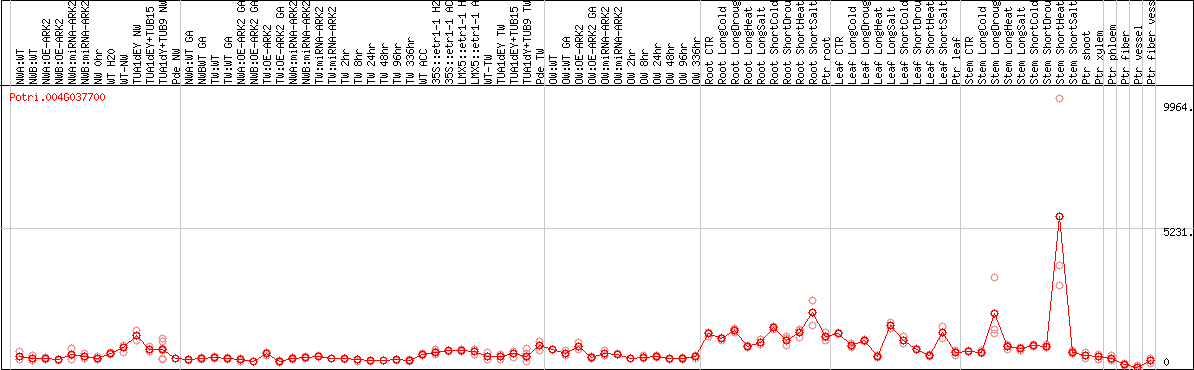
Coexpressed genes
Potri.004G037700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
