Potri.004G040000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G040000.5 pacid=42796614 polypeptide=Potri.004G040000.5.p locus=Potri.004G040000 ID=Potri.004G040000.5.v4.1 annot-version=v4.1
ATGTATAAATCAGTGGTGTATAAAGGAGATGAGTTACTGGGAGAAGTAGAAATATACGCACAAGAACAACAGCAAGAAGAAGAAGAGAACAAGAACAAGA
AGAAGAGAGTGATTGATGAGATAGTGAAGGAAATCAGAATAAGCCATTTCTCACAAACAAGTGAGAGATGTCCACCACTTGCAGTGCTTCATACAATTAC
ATCTATTGGTGTTTGCTTCAAAATGGAAGAGTCAACTTCATCATCAACAACAAAGATCTCTCAACAAGAATCGCCGCTTCATCTTTTGCACTCCTCCTGT
ATCCAAGAAAACAAGACTGCAGTGATGCATCTGGGAGGGGAGGAGCTCCATTTGGTTGCAATGCCCTCCAGGAGTAATGAAAGACAGCATCCATGTTTCT
GGGGATTCAGTGTTGCACCAGGACTTTATGACTCTTGTCTTGTCATGCTGAACCTCAGATGCCTGGGCATTGTATTTGATCTTGATGAAACTCTCATAGT
TGCAAATACAATGCGCTCATTTGAAGATAGAATTGATGCTTTGCAGCGGAAAATAAGCACTGAGGTGGACCCGCAGCGCATTCTTGGTATGCTGTCAGAG
GTCAAGAGGTATCACGATGACAAGAACATTTTGAAGCAATATGTGGAAAATGATCAGGTTGTTGAAAATGGAAAGGTGATCAAAACACAGTCTGAGGTTG
TTCCAGCCCTGTCTGACAACCATCAACCTATGGTTCGCCCCTTGATACGGTTGCAAGAGAAGAACATTATTTTGACTCGTATCAATCCGCAGATTCGTGA
TACAAGTGTGCTTGTGAGGTTGAGACCTGCTTGGGAGGATCTGAGAAGCTACTTGACTGCAAGAGGGCGCAAGCGCTTTGAGGTTTATGTTTGCACAATG
GCTGAAAGAGATTATGCTTTAGAAATGTGGAGGCTTTTGGATCCAGAGTCAAACTTGATAAACTCAAAAGAATTGCTAGATCGGATTGTGTGTGTCAAGT
CTGGTTTAAGGAAGTCCTTGTTCAATGTCTTTCAAGATGGAATATGCCATCCTAAGATGGCATTGGTGATAGATGATCGCTTGAAAGTGTGGGATGAGAG
GGATCAATCCCGTGTGCATGTTGTTCCTGCATTTGCTCCATACTATGCTCCTCAAGCTGAAGTAAATAATGCTGTTCCAGTATTGTGCGTTGCGAGAAAT
GTTGCTTGCAATGTCAGAGGTGGTTTCTTCAAAGAATTTGATGAGGGCCTTCTGCAGAAGATACCTGAGGTTGCATACGAAGATGATACTGATAATATTC
CTTCTCCTCCTGATGTGAGCAACTATTTAGTTTCAGAGGATGATGCTTCTGCTGTAAATGGAAATAGAGATCAACTTTCTTTTGATGGCATGGCAGATGC
TGAGGTTGAAAGACAGCTAAAGGAAGCAGTTTCAGCTTCTTCAGCAATCCTCTCAACTATTCCTTCAACAGTCAGTAGCTTAGATCCAAGGCTTCTTCAG
TCTCTCCAGTACACAATAGCATCTTCTTCAAGCTCGATGCCAACATCACAACCTTCAATGCTGGCATCACAACAACCAATGCCAGCATTACAACCACCAA
AGCCTCCATCACAACTATCAATGACGCCATTTCCCAACACACAGTTTCCTCAGGTGGCTCCATCAGTTAAACAATTGGGCCAGGTTGTCCCTCCAGAACC
AAGCTTGCAAAGTTCTCCTGCTCGAGAAGAGGGTGAAGTACCCGAATCTGAATTAGATCCTGATACAAGGAGGCGGCTTCTTATATTACAACATGGGCAT
GATTCAAGAGACAATGCACCAAGTGAATCTCCATTTCCTGCAAGACCTTCAACACAAGTTTCAGCTCCACGTGTGCAATCTGTTGGAAGCTGGGTTCCAG
TGGAGGAAGAGATGAGCCCAAGGCAACTGAACCGAACACCTAGAGAATTTCCTCTGGATTCAGATCCAATGAATATTGAAAAGCACCGAACTCACCATCC
ATCCTTTTTCCATAAGGTTGAGAGTAATATTCCGTCTGATAGGATGATTCATGAGAACCAAAGACAGCCAAAAGAGGCAACATATAGAGATGATCGGATG
AAGCTAAACCATTCAACATCTAATTATCCTTCCTTCCAAGGGGAGGAGAGTCCATTGAGTAGATCATCCAGTAACAGGGATCTTGACTTGGAATCTGAAC
GAGCTTTTTCAAGCACAGAAACTCCAGTTGAAGTGTTACAGGAAATTGCGATGAAGTGTGGAACCAAGGTGGAATTCAGGCCTGCTTTGATTGCTACTAG
TGATCTGCAGTTTTCCATTGAGACTTGGTTTGTAGGAGAGAAAGTTGGTGAAGGAACTGGTAAAACAAGAAGGGAAGCCCAACGCCAGGCAGCTGAGGGT
TCTATTAAGAAGTTAGCTGGCATATATATGTCACGTGTGAAGCCTGATTCTGGGCCCATGCTTGGGGATTCAAGTAGATATCCTAGTGCAAATGATAATG
GTTTCTTGGGTGATATGAATTCTTTTGGGAATCAACCATTGCTGAAAGATGAGAACATTACGTATTCAGCTACTTCAGAGCCATCCAGACTTCTGGATCA
AAGGCTGGAAGGCTCTAAGAAATCAATGGGTTCGGTCACTGCACTCAAAGAATTTTGCATGACGGAGGGTCTTGGGGTAAATTTTCTGGCACAGACTCCA
CTTTCAACTAATTCAATACCGGGAGAGGAGGTACATGCACAGGTTGAAATAGATGGACAAGTTCTGGGGAAGGGTATTGGGTTGACATGGGATGAAGCTA
AGATGCAGGCTGCTGAAAAGGCCCTTGGAAGTCTAAGAACAATGTTTGGTCAATACACACCAAAACGTCAGGGTTCCCCTAGGCTAATGCAAGGGATGCC
AAATAAACGTCTAAAGCAAGAGTTCCCACGGGTTCTGCAGCGCATGCCCTCTTCTGCTAGATATCACAAGAATGCTTCTCCTGTTCCTTGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G040000.5 pacid=42796614 polypeptide=Potri.004G040000.5.p locus=Potri.004G040000 ID=Potri.004G040000.5.v4.1 annot-version=v4.1
MYKSVVYKGDELLGEVEIYAQEQQQEEEENKNKKKRVIDEIVKEIRISHFSQTSERCPPLAVLHTITSIGVCFKMEESTSSSTTKISQQESPLHLLHSSC
IQENKTAVMHLGGEELHLVAMPSRSNERQHPCFWGFSVAPGLYDSCLVMLNLRCLGIVFDLDETLIVANTMRSFEDRIDALQRKISTEVDPQRILGMLSE
VKRYHDDKNILKQYVENDQVVENGKVIKTQSEVVPALSDNHQPMVRPLIRLQEKNIILTRINPQIRDTSVLVRLRPAWEDLRSYLTARGRKRFEVYVCTM
AERDYALEMWRLLDPESNLINSKELLDRIVCVKSGLRKSLFNVFQDGICHPKMALVIDDRLKVWDERDQSRVHVVPAFAPYYAPQAEVNNAVPVLCVARN
VACNVRGGFFKEFDEGLLQKIPEVAYEDDTDNIPSPPDVSNYLVSEDDASAVNGNRDQLSFDGMADAEVERQLKEAVSASSAILSTIPSTVSSLDPRLLQ
SLQYTIASSSSSMPTSQPSMLASQQPMPALQPPKPPSQLSMTPFPNTQFPQVAPSVKQLGQVVPPEPSLQSSPAREEGEVPESELDPDTRRRLLILQHGH
DSRDNAPSESPFPARPSTQVSAPRVQSVGSWVPVEEEMSPRQLNRTPREFPLDSDPMNIEKHRTHHPSFFHKVESNIPSDRMIHENQRQPKEATYRDDRM
KLNHSTSNYPSFQGEESPLSRSSSNRDLDLESERAFSSTETPVEVLQEIAMKCGTKVEFRPALIATSDLQFSIETWFVGEKVGEGTGKTRREAQRQAAEG
SIKKLAGIYMSRVKPDSGPMLGDSSRYPSANDNGFLGDMNSFGNQPLLKDENITYSATSEPSRLLDQRLEGSKKSMGSVTALKEFCMTEGLGVNFLAQTP
LSTNSIPGEEVHAQVEIDGQVLGKGIGLTWDEAKMQAAEKALGSLRTMFGQYTPKRQGSPRLMQGMPNKRLKQEFPRVLQRMPSSARYHKNASPVP
|
DESeq2's median of ratios [POPLAR]
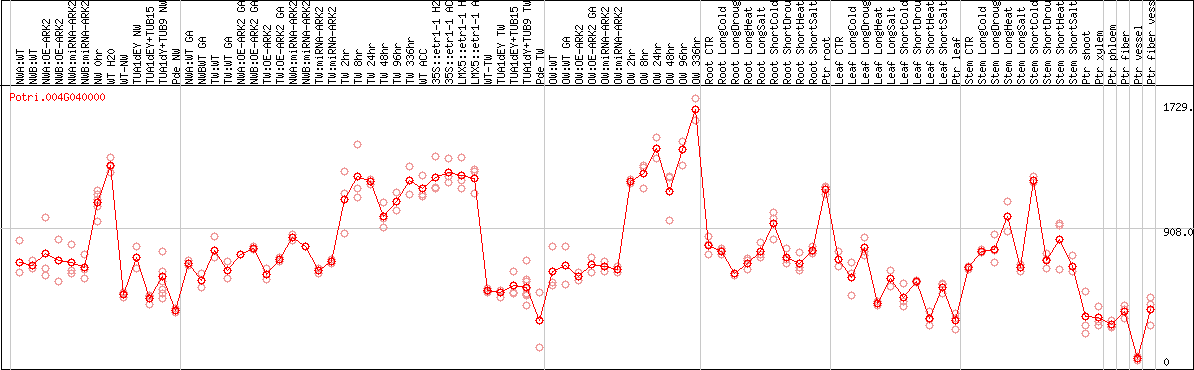
Coexpressed genes
Potri.004G040000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
