External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G28500 355 / 3e-123
NAC
ANAC073, SND2, NST7
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2, NAC domain containing protein 73 (.1)
AT1G28470 346 / 2e-119
NAC
ANA010, ANAC010, SND3, NST8
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 3, NAC domain containing protein 10 (.1)
AT4G29230 298 / 3e-98
NAC
ANAC075, NST9
NAC domain containing protein 75 (.1)
AT5G56620 271 / 6e-89
NAC
ANAC099
NAC domain containing protein 99 (.1)
AT1G25580 219 / 3e-68
NAC
ANAC008, SOG1
SUPPRESSOR OF GAMMA RADIATION 1, Arabidopsis NAC domain containing protein 8, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT5G14490 181 / 1e-54
NAC
ANAC085
NAC domain containing protein 85 (.1)
AT3G01600 173 / 3e-51
NAC
ANAC044
NAC domain containing protein 44 (.1)
AT5G18270 85 / 1e-18
NAC
ANAC087
Arabidopsis NAC domain containing protein 87 (.1.2)
AT3G04060 84 / 2e-18
NAC
ANAC046
NAC domain containing protein 46 (.1)
AT5G61430 77 / 9e-16
NAC
ANAC100, ATNAC5
NAC domain containing protein 100 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G058400
540 / 0
AT4G28500 344 / 4e-118
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2, NAC domain containing protein 73 (.1)
Potri.017G016700
382 / 1e-133
AT4G28500 344 / 4e-119
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2, NAC domain containing protein 73 (.1)
Potri.007G135300
354 / 7e-123
AT4G28500 337 / 3e-116
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2, NAC domain containing protein 73 (.1)
Potri.018G068700
295 / 1e-97
AT4G29230 511 / 2e-179
NAC domain containing protein 75 (.1)
Potri.006G152700
295 / 2e-97
AT4G29230 489 / 9e-171
NAC domain containing protein 75 (.1)
Potri.008G116600
238 / 9e-76
AT1G25580 509 / 4e-180
SUPPRESSOR OF GAMMA RADIATION 1, Arabidopsis NAC domain containing protein 8, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.010G129700
234 / 5e-74
AT1G25580 476 / 5e-167
SUPPRESSOR OF GAMMA RADIATION 1, Arabidopsis NAC domain containing protein 8, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.001G343800
219 / 1e-68
AT3G01600 266 / 5e-86
NAC domain containing protein 44 (.1)
Potri.015G046800
79 / 2e-16
AT3G18400 326 / 1e-111
NAC domain containing protein 58 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013967
358 / 3e-124
AT4G28500 320 / 2e-109
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2, NAC domain containing protein 73 (.1)
Lus10015392
347 / 2e-120
AT4G28500 308 / 2e-105
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2, NAC domain containing protein 73 (.1)
Lus10039873
327 / 6e-112
AT4G28500 355 / 5e-123
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2, NAC domain containing protein 73 (.1)
Lus10018637
325 / 2e-111
AT4G28500 350 / 2e-121
SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2, NAC domain containing protein 73 (.1)
Lus10034999
282 / 1e-91
AT4G29230 388 / 3e-130
NAC domain containing protein 75 (.1)
Lus10012927
275 / 7e-90
AT4G29230 346 / 3e-115
NAC domain containing protein 75 (.1)
Lus10021708
228 / 1e-71
AT1G25580 462 / 4e-161
SUPPRESSOR OF GAMMA RADIATION 1, Arabidopsis NAC domain containing protein 8, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10041492
212 / 1e-65
AT1G25580 486 / 7e-171
SUPPRESSOR OF GAMMA RADIATION 1, Arabidopsis NAC domain containing protein 8, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10003668
207 / 3e-63
AT5G14490 273 / 2e-88
NAC domain containing protein 85 (.1)
Lus10026879
81 / 8e-17
AT5G18270 325 / 3e-110
Arabidopsis NAC domain containing protein 87 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Potri.004G049300.1 pacid=42794902 polypeptide=Potri.004G049300.1.p locus=Potri.004G049300 ID=Potri.004G049300.1.v4.1 annot-version=v4.1
ATGACATGGTGCAATAATAACTCAGAAGATGAACGAGTCATTCAATTAGTCACAAGTAACCCGAAAGAAGATACTTTTGTCGCTGCAAAAACAGAAGAAA
TTCGAAGGATAACTTGCCCTTCATGTGGCCATAATTTTGAAATTCAAGATCAGGGTGGTATAATTCATGATTTGCCGGGACTACCAGCGGGAGTGAAGTT
TGATCCGACGGACCAAGAAATACTTGAACATTTAGAAGCAAAGGTGTTATCAGATAGGCGTAAACTTCATCCTCTAATTGATGAATTTATCCCAACAATA
GAGGGGGAGAATGGAATTTGCTACTCTCATCCTGAGAAGCTACCAGGAGTAAGCAATGATGGCCAAATCCGCCACTTCTTTCATCGACCATCAAAGGCAT
ACACAACCGGAACTAGAAAACGCAGAAAGGTGCACACTGATGACGATGGAAGTGAAACCAGATGGCACAAAACAGGCAAGACCAGGCCAGTTTTTGCTGG
CGGGACAGTGAAGGGATTCAAAAAAATTCTGGTGCTTTACACCAATTATGGTAGGCAAAGGAAACCTGAAAAGACAAACTGGGTAATGCACCAATACCAT
CTTGGAAACAACGAAGAAGAGAAAGATGGAGAGTTAGTGGTTTCAAAAGTTTTCTATCAAACTCAGCCTAGACAATGTAGTTCAAGCATTAAGGATTCAA
TTGACAATAAATCGACAAATCAAAGTGGTGATCATATTGATAATATTCACCCCCTAGCCAAGAACAGTACAGGCCTCCTTGATCAGTTCTATAATCGAGC
TTATATTTCTTATGATCATGGGAATCATAGCAGTGAAATCCCACCTCAATTTCTCCCCAATTTGGTTGTTCAGGGTGACGGGTCTTCCTATATTAGGTTA
GCTGCAGAGACAAGCAAAGGAAAGCTTCAGAGAAAGCAGTGA
AA sequence
>Potri.004G049300.1 pacid=42794902 polypeptide=Potri.004G049300.1.p locus=Potri.004G049300 ID=Potri.004G049300.1.v4.1 annot-version=v4.1
MTWCNNNSEDERVIQLVTSNPKEDTFVAAKTEEIRRITCPSCGHNFEIQDQGGIIHDLPGLPAGVKFDPTDQEILEHLEAKVLSDRRKLHPLIDEFIPTI
EGENGICYSHPEKLPGVSNDGQIRHFFHRPSKAYTTGTRKRRKVHTDDDGSETRWHKTGKTRPVFAGGTVKGFKKILVLYTNYGRQRKPEKTNWVMHQYH
LGNNEEEKDGELVVSKVFYQTQPRQCSSSIKDSIDNKSTNQSGDHIDNIHPLAKNSTGLLDQFYNRAYISYDHGNHSSEIPPQFLPNLVVQGDGSSYIRL
AAETSKGKLQRKQ
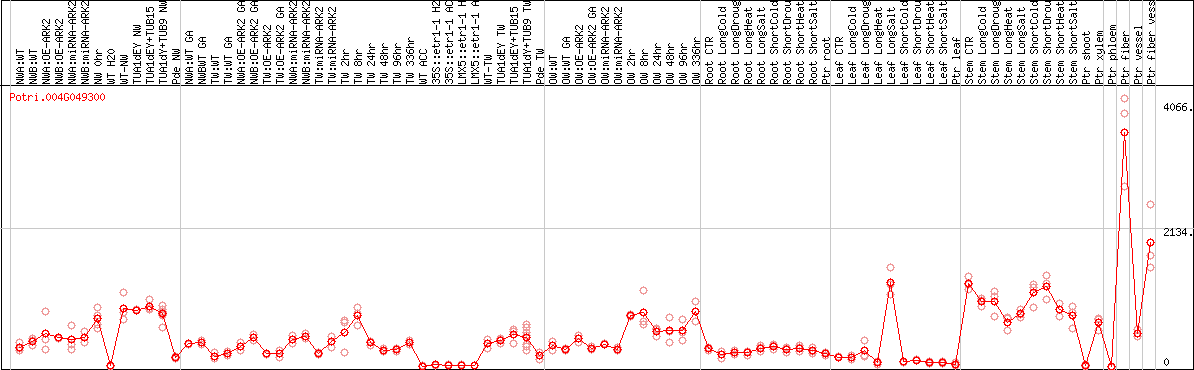
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G049300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.