External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G41130 120 / 1e-32
bHLH
bHLH106
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G56770 111 / 1e-29
bHLH
bHLH107
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT1G68810 113 / 4e-29
bHLH
bHLH030
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G25710 112 / 9e-29
bHLH
TMO5, BHLH32, bHLH032, ATAIG1
TARGET OF MONOPTEROS 5, basic helix-loop-helix 32 (.1)
AT2G40200 104 / 9e-27
bHLH
bHLH051
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G38070 57 / 4e-09
bHLH
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G46760 44 / 5e-05
bHLH
bHLH005, ATR2, MYC3
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT1G32640 44 / 5e-05
bHLH
JIN1, JAI1, ZBF1, RD22BP1, ATMYC2
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT1G01260 42 / 0.0003
bHLH
bHLH013, INU4
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G065500
343 / 2e-120
AT1G68810 113 / 1e-29
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.019G089000
211 / 8e-68
AT1G68810 122 / 2e-32
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.013G117600
198 / 1e-62
AT3G56770 112 / 8e-30
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.006G037100
120 / 9e-33
AT2G41130 229 / 2e-75
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.010G130000
122 / 1e-32
AT1G68810 269 / 1e-87
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.008G116000
119 / 2e-31
AT1G68810 259 / 5e-84
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.016G035400
115 / 8e-31
AT2G41130 231 / 2e-76
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.008G070800
101 / 2e-25
AT2G40200 132 / 2e-37
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.010G186700
89 / 6e-21
AT2G40200 132 / 3e-37
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013917
257 / 8e-86
AT2G40200 122 / 2e-33
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10001885
222 / 4e-72
AT1G68810 124 / 1e-33
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10038306
180 / 4e-56
AT3G56770 102 / 2e-26
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10036175
128 / 1e-36
AT3G25710 75 / 3e-16
TARGET OF MONOPTEROS 5, basic helix-loop-helix 32 (.1)
Lus10035056
121 / 6e-32
AT1G68810 321 / 6e-108
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10021711
119 / 4e-31
AT1G68810 310 / 1e-103
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10018020
111 / 6e-29
AT2G41130 206 / 1e-65
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10042017
110 / 1e-28
AT2G41130 204 / 2e-65
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10007502
106 / 1e-27
AT2G40200 163 / 8e-50
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10041493
107 / 2e-27
AT3G25710 201 / 2e-62
TARGET OF MONOPTEROS 5, basic helix-loop-helix 32 (.1)
PFAM info
Representative CDS sequence
>Potri.004G055700.1 pacid=42794797 polypeptide=Potri.004G055700.1.p locus=Potri.004G055700 ID=Potri.004G055700.1.v4.1 annot-version=v4.1
ATGTTTGACTTCTGTTTTAACTCTAACTCTTCATCTGGGTCCGGTTATATGAACACGTTTGATCCGTTTCCGCGTAGTTTAGAGGGTTTTAGTGATGTTT
TGCAAGGTAGGCCAATGGTTTCTCAAACTTTGGTGTTGGATGCTGAGATGGGAGAGCTAGTAAAGGCACCTGCAAGAGTTGGGAATAAGGGGATATCTGA
GGCCAAGGCTTTGGCAGCTTTGAAGAGCCATAGCGAGGCCGAGAGGAGGAGGAGGGAGAGGATCAATGCCCATCTTGATACTCTTCGCGGTCTTGTGCCC
TGCACTGAAAAGATGGACAAAGCCACGTTGCTTGCCGCAGTCATTAGCCAAGTCAATGAACTCAAAAGGAATGCCTTGGAATCCTGTAAAGGCCTTCTTA
TTCCAACAGCTGATGACGAAGTAAAAGTTGAGACATATTTTGATGGAACTAAAGAAGGAACGTTGTACTTCAAGGCATCTATCTGTTGTGACTATCGACC
TGAGCTTCTATCCGATATAAGACAAGCTGTTGATGCACTTCCTTTGAAAATGGTGAACGCAGAAATATCAACCTTGGGAAACCGGTTAAAGAATGAGTTT
GTATTCACCAGCAACAGAAATAAGAATGCTGTAGATGATGCAGAGGCAATGCAGCATCTTACAAAATCTATTCACCATGCCCTAACATCTGTCTTGGAGA
AAGGTTCTGCCTCACTAGAATACTCACCAAGGACAACACTTCCAAACAAGAAAAGGAGGGTGACTTTCTTTGATTCCTCGAGCTCGTCTTCTTAA
AA sequence
>Potri.004G055700.1 pacid=42794797 polypeptide=Potri.004G055700.1.p locus=Potri.004G055700 ID=Potri.004G055700.1.v4.1 annot-version=v4.1
MFDFCFNSNSSSGSGYMNTFDPFPRSLEGFSDVLQGRPMVSQTLVLDAEMGELVKAPARVGNKGISEAKALAALKSHSEAERRRRERINAHLDTLRGLVP
CTEKMDKATLLAAVISQVNELKRNALESCKGLLIPTADDEVKVETYFDGTKEGTLYFKASICCDYRPELLSDIRQAVDALPLKMVNAEISTLGNRLKNEF
VFTSNRNKNAVDDAEAMQHLTKSIHHALTSVLEKGSASLEYSPRTTLPNKKRRVTFFDSSSSSS
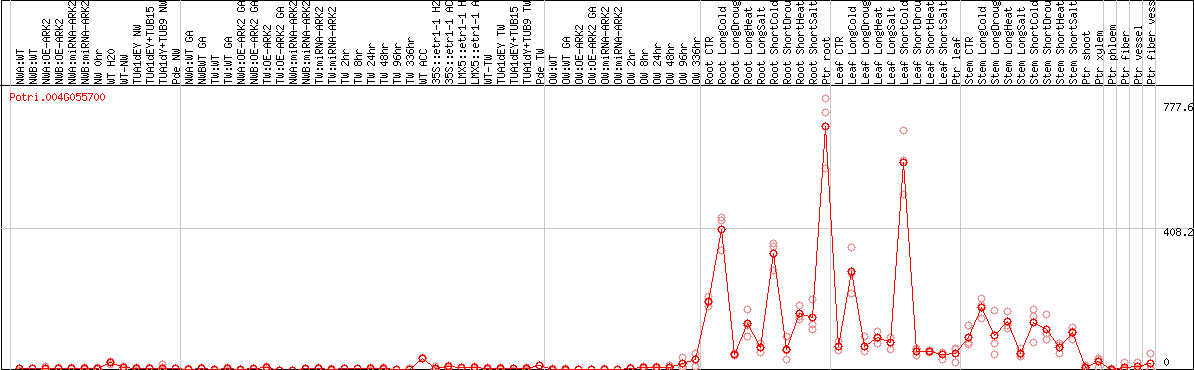
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G055700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.