External link
Symbol
PEX7.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G29260 555 / 0
PEX7, ATPEX7
ARABIDOPSIS PEROXIN 7, peroxin 7 (.1)
AT5G58230 94 / 2e-21
MSI1, MEE70, ATMSI1
MATERNAL EFFECT EMBRYO ARREST 70, ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, Transducin/WD40 repeat-like superfamily protein (.1)
AT4G35050 93 / 6e-21
NFC3, MSI3
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP C 3, MULTICOPY SUPPRESSOR OF IRA1 3, Transducin family protein / WD-40 repeat family protein (.1)
AT3G49660 86 / 4e-19
AtWDR5a
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
AT2G16780 85 / 2e-18
MSI02, NFC2, NFC02, MSI2
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP C 2, MULTICOPY SUPPRESSOR OF IRA1 2, Transducin family protein / WD-40 repeat family protein (.1)
AT5G25150 83 / 3e-17
TAF5
TBP-associated factor 5 (.1)
AT2G05720 79 / 9e-17
Transducin/WD40 repeat-like superfamily protein (.1)
AT4G18900 77 / 2e-15
Transducin/WD40 repeat-like superfamily protein (.1)
AT1G52360 77 / 2e-15
Coatomer, beta' subunit (.1.2)
AT4G02730 76 / 2e-15
AtWDR5b
human WDR5 \(WD40 repeat\) homolog b, Transducin/WD40 repeat-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G066600
420 / 3e-150
AT1G29260 379 / 3e-134
ARABIDOPSIS PEROXIN 7, peroxin 7 (.1)
Potri.005G148500
92 / 3e-21
AT3G49660 503 / 0.0
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.004G175300
92 / 8e-21
AT2G16780 593 / 0.0
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP C 2, MULTICOPY SUPPRESSOR OF IRA1 2, Transducin family protein / WD-40 repeat family protein (.1)
Potri.014G179700
92 / 2e-20
AT5G58230 790 / 0.0
MATERNAL EFFECT EMBRYO ARREST 70, ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.002G216200
91 / 2e-20
AT5G58230 784 / 0.0
MATERNAL EFFECT EMBRYO ARREST 70, ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.009G135100
89 / 8e-20
AT2G16780 612 / 0.0
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP C 2, MULTICOPY SUPPRESSOR OF IRA1 2, Transducin family protein / WD-40 repeat family protein (.1)
Potri.006G263400
89 / 2e-19
AT5G25150 1018 / 0.0
TBP-associated factor 5 (.1)
Potri.018G020200
88 / 4e-19
AT5G25150 984 / 0.0
TBP-associated factor 5 (.1)
Potri.007G009500
84 / 3e-18
AT3G49660 469 / 4e-168
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029221
558 / 0
AT1G29260 531 / 0.0
ARABIDOPSIS PEROXIN 7, peroxin 7 (.1)
Lus10007274
557 / 0
AT1G29260 529 / 0.0
ARABIDOPSIS PEROXIN 7, peroxin 7 (.1)
Lus10005074
91 / 2e-20
AT4G35050 590 / 0.0
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP C 3, MULTICOPY SUPPRESSOR OF IRA1 3, Transducin family protein / WD-40 repeat family protein (.1)
Lus10027848
90 / 5e-20
AT4G35050 585 / 0.0
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP C 3, MULTICOPY SUPPRESSOR OF IRA1 3, Transducin family protein / WD-40 repeat family protein (.1)
Lus10020917
90 / 7e-20
AT5G58230 696 / 0.0
MATERNAL EFFECT EMBRYO ARREST 70, ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10039636
89 / 9e-20
AT3G49660 502 / 0.0
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10033459
89 / 1e-19
AT5G58230 723 / 0.0
MATERNAL EFFECT EMBRYO ARREST 70, ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10033460
89 / 1e-19
AT5G58230 781 / 0.0
MATERNAL EFFECT EMBRYO ARREST 70, ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10020916
89 / 2e-19
AT5G58230 782 / 0.0
MATERNAL EFFECT EMBRYO ARREST 70, ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10024447
85 / 6e-18
AT5G25150 1006 / 0.0
TBP-associated factor 5 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
>Potri.004G057600.1 pacid=42794614 polypeptide=Potri.004G057600.1.p locus=Potri.004G057600 ID=Potri.004G057600.1.v4.1 annot-version=v4.1
ATGCCAGTCTTCAAAACCCCATTCAATGGCTACTCCGTCAAATTCAGCCCATTTTACGAATCCCGTCTCGCCGTCGCCACTGCCCAGAACTTCGGAATCC
TCGGCAACGGTCGTCTCCACGTCCTCTCCTTACCTCCAGCACCCTCCTCCCCTCTAACCGAACTCATCTCCTTCGACACTGCTGATGGCATCTACGACCT
TGCCTGGTCTGAATCCCACGATTCCCTCTTAATCGCAGCTGTGGCCGATGGGTCCGTCAAGCTTTACGACACCGCCTTACCCCCAACACAAAACCCAATT
CGTTCCCTCCAGGAACACACGCGTGAAGTCCACTCAGTCGATTACAATCCCACGCGCCGTGATTCTTTTATTACCGCTTCCTGGGATGACACGATTAAAT
TATGGACCCTTGACAGGCCGGCGAGTATTCGTACGTTTAAAGAACACGCGTATTGTGTTTATTCTGCTGCGTGGAACCCGAGGCATACGGATGTTTTTGC
ATCTGCTTCCGGGGACTGTACTGTGCGAATCTGGGACGTGCGTGAGCCGGGGAGTACGATGATTATTCCAGGTCATGATTTTGAGATTTTGTGTTGTGAT
TGGAATAAATATGATGATTGTATTATTGCTACTGCATCGGTGGATAAGAGTATTAAAGTATGGGATGTAAGGAGTTTCAGAGCTCCGATTTCGGTTTTGA
ATGGACATGGGTATGCTGTTAGGAAAGTTAAGTTTTCGCCCCATCATAGGAATTTGATGGTTTCGTGTTCGTATGATATGAGTGTTTGTATGTGGGATTT
TATGGTTGAAGATGCATTGGTTGGGAGGTATGATCATCATACTGAGTTTGCTGTTGGAGTTGATATCAGTGTGCTTGTTGATGGGTTGATGGCAAGCACT
GGGTGGGATGAGTTGGTTTATGTTTGGCAGCATGGGACGGACCCGAGAGCACCTTGA
AA sequence
>Potri.004G057600.1 pacid=42794614 polypeptide=Potri.004G057600.1.p locus=Potri.004G057600 ID=Potri.004G057600.1.v4.1 annot-version=v4.1
MPVFKTPFNGYSVKFSPFYESRLAVATAQNFGILGNGRLHVLSLPPAPSSPLTELISFDTADGIYDLAWSESHDSLLIAAVADGSVKLYDTALPPTQNPI
RSLQEHTREVHSVDYNPTRRDSFITASWDDTIKLWTLDRPASIRTFKEHAYCVYSAAWNPRHTDVFASASGDCTVRIWDVREPGSTMIIPGHDFEILCCD
WNKYDDCIIATASVDKSIKVWDVRSFRAPISVLNGHGYAVRKVKFSPHHRNLMVSCSYDMSVCMWDFMVEDALVGRYDHHTEFAVGVDISVLVDGLMAST
GWDELVYVWQHGTDPRAP
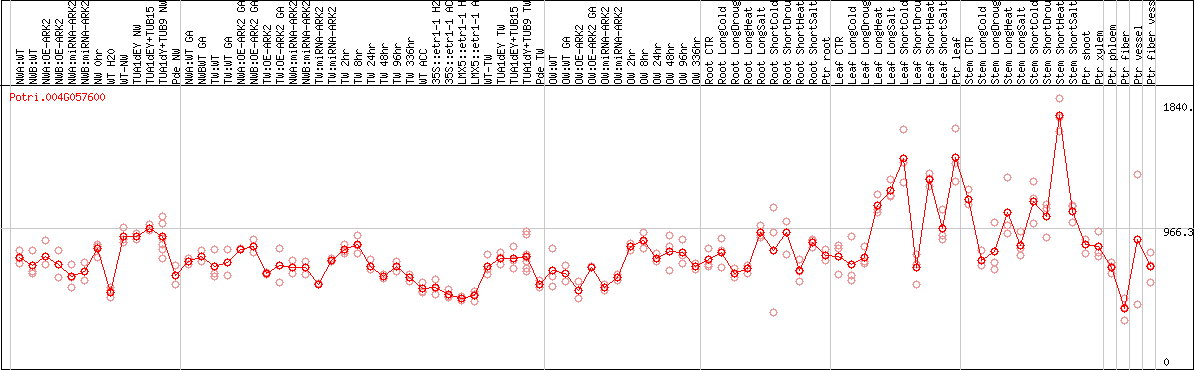
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G057600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.