External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G08350 197 / 2e-63
GRAM domain-containing protein / ABA-responsive protein-related (.1)
AT5G23370 194 / 2e-62
GRAM domain-containing protein / ABA-responsive protein-related (.1)
AT5G23350 193 / 4e-61
GRAM domain-containing protein / ABA-responsive protein-related (.1)
AT5G23360 184 / 1e-58
GRAM domain-containing protein / ABA-responsive protein-related (.1)
AT5G13200 147 / 2e-43
GRAM domain family protein (.1)
AT4G01600 142 / 9e-42
GRAM domain family protein (.1.2)
AT2G22475 133 / 1e-37
GEM
GL2-EXPRESSION MODULATOR, GRAM domain family protein (.1.2)
AT1G28200 131 / 3e-37
FIP1
FH interacting protein 1 (.1)
AT4G40100 88 / 5e-21
GRAM domain family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G088400
226 / 9e-75
AT5G23370 241 / 8e-81
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Potri.005G088700
225 / 2e-74
AT5G08350 234 / 5e-78
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Potri.005G088300
224 / 7e-74
AT5G23370 244 / 4e-82
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Potri.007G075700
223 / 2e-73
AT5G08350 240 / 3e-80
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Potri.007G075200
222 / 4e-73
AT5G08350 240 / 3e-80
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Potri.005G088200
217 / 2e-71
AT5G08350 230 / 1e-76
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Potri.004G068100
151 / 5e-46
AT5G23370 111 / 1e-30
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Potri.004G068200
147 / 2e-44
AT5G23350 112 / 3e-30
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Potri.004G068150
146 / 5e-44
AT5G23350 106 / 4e-28
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013445
228 / 3e-75
AT5G08350 246 / 1e-82
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Lus10017372
226 / 2e-74
AT5G08350 238 / 2e-79
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Lus10010166
223 / 2e-73
AT5G08350 233 / 1e-77
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Lus10040999
216 / 2e-70
AT5G08350 234 / 5e-78
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Lus10010167
212 / 5e-69
AT5G08350 247 / 4e-83
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Lus10017373
209 / 7e-68
AT5G08350 240 / 3e-80
GRAM domain-containing protein / ABA-responsive protein-related (.1)
Lus10035154
139 / 7e-40
AT5G13200 308 / 9e-106
GRAM domain family protein (.1)
Lus10031986
136 / 7e-39
AT5G13200 302 / 3e-103
GRAM domain family protein (.1)
Lus10040539
135 / 1e-38
AT1G28200 338 / 6e-118
FH interacting protein 1 (.1)
Lus10010279
133 / 4e-38
AT2G22475 196 / 3e-62
GL2-EXPRESSION MODULATOR, GRAM domain family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0266
PH
PF02893
GRAM
GRAM domain
Representative CDS sequence
>Potri.004G068000.1 pacid=42795632 polypeptide=Potri.004G068000.1.p locus=Potri.004G068000 ID=Potri.004G068000.1.v4.1 annot-version=v4.1
ATGAGCATGAAGGGTCAGTTTGCAGACAAGTTCAATGGAAGTCCATTGATGATCAATAAAACTTTTGAGACAACCCCAGAAAGATCATCTTCTTTATCTG
AGCCTGTCAGCTTGCTGCAAAGTCCATATTCCTCTCAGGGTTCTCCAAGCTCCTCCAAAACTAGTAGTACTAAATGGGATTCACAGCCAGGAAAAACAAA
AACACTAGGAAGGAAAAGCAGTACTTTTGCTTACAGGATTCACGACCACGTGAAGTTGGGGGCTACCTTCTCTGAAACAGTGAAAGGCAAGCTGAGACTG
GGGGCAAAGATAATTCAAGAAGGAGGGAGGGAGAATATTTTCAAGCAGGTTTTTGGTGTCAGAGAAGGAGAAGAGCTATTAAAGGCATCACAGTGCTATT
TATCAACCACAGCTGGTCCTCTTCCAGGACTTCTTTTTATCTCCACTGAAAAGGTTGCCTTCTGCAGTGAGAGATCAATTACCTTCCCTTCTCCTAATGG
ACAATTTGTCAGAAAACCCTACAAGGTGGTGATACCAGTGAGGAAGATTGAAAGAGCCAACAGGAGTGAGAACATGGACAAGCCGCAACAGAAGTACATA
GAAATAGTCACGCAAGACAATTTTGAGTTCTGGTTCATGGGATTCTTGCGTTACGAAAAAGCTTTCAAGAATCTACACAAAGCAATTTCTTTGGCGAATT
CTGCCTCTTAA
AA sequence
>Potri.004G068000.1 pacid=42795632 polypeptide=Potri.004G068000.1.p locus=Potri.004G068000 ID=Potri.004G068000.1.v4.1 annot-version=v4.1
MSMKGQFADKFNGSPLMINKTFETTPERSSSLSEPVSLLQSPYSSQGSPSSSKTSSTKWDSQPGKTKTLGRKSSTFAYRIHDHVKLGATFSETVKGKLRL
GAKIIQEGGRENIFKQVFGVREGEELLKASQCYLSTTAGPLPGLLFISTEKVAFCSERSITFPSPNGQFVRKPYKVVIPVRKIERANRSENMDKPQQKYI
EIVTQDNFEFWFMGFLRYEKAFKNLHKAISLANSAS
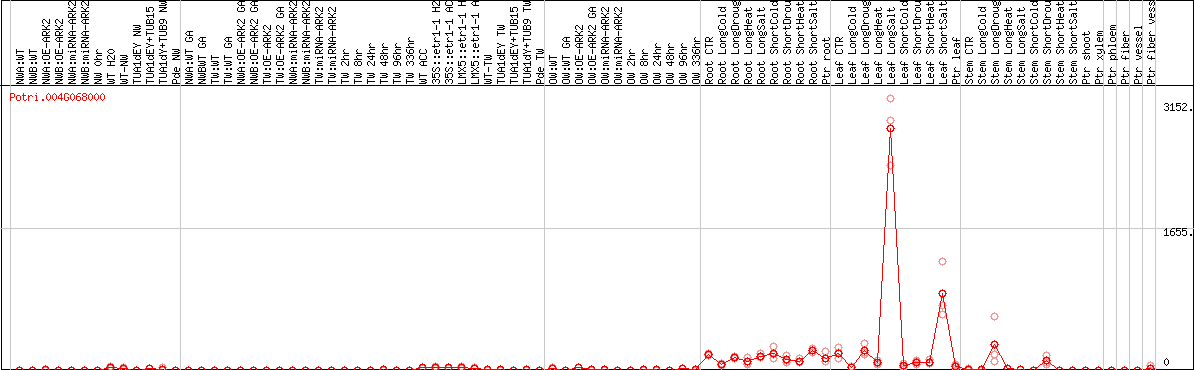
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G068000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.