Potri.004G077600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G077600.1 pacid=42796329 polypeptide=Potri.004G077600.1.p locus=Potri.004G077600 ID=Potri.004G077600.1.v4.1 annot-version=v4.1
ATGGACAAGAGAACTGATCGTGTGAAGTGGGAAAAGGTCAAGGGTAGAGGGTTCCTAGATTTAGTGTTTTCTTGGTCTATTGAAGATGTGCTCAATAAGG
ATCTTTACAAAGACCAGGTGGAGGAGATCCCAAACTCCTTCATGTCAACAGCTCATTACATGAAAGCATTCATTACACCCCTACACGTGGAAACACATGC
TGATTTGCTTTCAAGTACAGAATCACTGGCTGGAGCACCAACTTATAGAATACTTCGTGTTAGAAAATCTAAGGATTACAAGCCACCTAAGGACTTGTTT
TACGAAATTTCGATGGAAGAAACCAGAGGTGGATATGTGCCTTGGGTTGGGGATCTTATAGCCCTGACAAATGTGAAACTGAAGTGCATTGATGATTTAA
GGAAGACCCAGCAATCCTATCATGTTGCTTTCGTTCATGCCGTGAAAAGGGGAAATCGTTTGACACCCTCGATTCTGTCTTCGAAGCCCATAGTGGATGA
AGAAGGCTTGAAGAATGGAACACTCTTTGCTGTTCATTTGATAAATCTGATGACAAATTTGCGAATATGGAGATCGTTGCACTTGGAACTCGAAGGGAGG
AACATGAACGTTATTGAGAAAGTGCTGCAAAATAATTTCAATGATGATGGAGATTGTACTATCTGCTCTTCTCGTAAAAAAAGTGATGCTGCCTCTGCCT
GCATAAGGGATACATTACAGTCTTCTAATTTAAACAGCTCGCAAGAAGCTGCGGTTTTAAGCTGTATTCACACAGCCAGATGCTGGCATCAGTACACTGT
CAAATTAGTACAGGGTCCTCCAGGAACTGGGAAAACAAAGACAGCTAGTTGCTTATTACACGCACTATTAAGAATGAAATGCAGAACACTTACATGTGCT
CCAACTAACATTGCAGTGGTAGAAGTTGCAGCTCGTGTCGTGAGTACAGTTGCCGATTTAGTTGAATATGAAACTTATGGCATGGGAGACATAATTCTGT
TTGGGAATTGGGAGAGAATGAAGTTTGATGGTGATCAGAATGACCTTCTTCATGTATTTCTTGATCATCGTGCTGATATACTTGAGAAGTGCTTTGATCC
ATCAACTGGCTGGAAACGCATTTTAGCTTCATTGATAAGTTTACTTGAAGACTCTGAAGCGCAATACCATTTGTACTTGCAGGATAACATGGGAAAGGAA
GGACTTTTGACTTGTGAGCAATTCGTATGGAAAAGATTTGATTTCTCCGGAAAGCAATTAAAGTTCTGTATTGTGAATTTGTATACACACTTACCGACTA
CTTTGATTTCATTACAAGTGATGAGGATCATGACCAGGGCTCTTGATTTGATGACATCACTTGAAACCTTGTTGCTTAGTCTCAGTGCTGCCGATGAAGG
TTTAAAGCAGATTCTAGGGGAAAATGAAGATGAAGAAAGAAAACTTCATAATCGTATAAAGTTGATAAATGAAAAAAGGGAATGCCTCAATACTCTACGC
TTACTTTCTCTAAAATTTCAGGTTCCAGAATTCGCTGATAAAAATGCTATTGAGAAATTCTGCTTATCAAATGCATGCCTGATTTTCTGTACCGTGTCAA
GCTCTGCCAGGCTGCATAGTATAAGAATGGCACCTCTGCGTTGCTTGGTGATTGACGAAGCTGCTCAGCTCAAAGAATGTGAATCAACCATCCCCTTACA
ACTCTTCGGCCTTCACCATGCTATTCTCATAGGAGATGAACGGCAATTGCCTGCCATCGTTAATAGCGAGATTTCAGGGAAGGCTGGTTTTGGTAGAAGT
TTGTTTGAGAGGCTCGTAAAGCTGGGATGCAAGAGCCATCTTCTAAACATTCAATATCGAATGCATCCATCAATAAGCTTATTTCCGAATACGGAGTTCT
ATGGAAGGCAGGTTTTGGATGCGCCAAACGTCCAAGAAACTGGCTACAGGAGACGGTTCCTCCAAGGAGACATGTTTGAATCCTACTCTTTCATAAATAT
AGCCCATGGAAAAGAGGAATTTGTTGAGAAACAGAGTTTCAAAAATACAGTCGAGGCTGCTGCTGCTGCTGATATAGTTGGGAGACTTTTCAAAGATATT
AATGGCACTGGGAAGAAGGTTAGCATAGGTATCATATCACCATACCAGGCTCAAGTTCATGCAATTCAAGCGAAGATTGGGAAATTCATTTCAGACTCTG
ACAGTGCCTTGTCTGTAAGCGTTGGCACGGTTGATGGTTTTCAGGGTGGAGAGGAGGATCTGATAATTATCTCTACTGTTCGAAGCAATGAGAACGGATC
GGTGGGTTTTGTTTCGAATCCTCAACGAGCAAATGTGGCGCTGACTCGTGCAAGGTATTGCCTCTGGATTTTAGGAAATGAAGCAACTTTAGTTAAGAGT
GGCTCTATTTGGAAGGAGATAGTCAACGATGCGAAGCATCGGCAATGCTTCTATAACGCTGAAGAGGATGAGAGTTTGGCTCAGGCTATTACAGAAAGCT
TGATCGAGCATGGCCGACTTGATGTTCTGCTACGAACTCACTCCCCATTGTTCCGGAATGCAAGATGGATGGTTTTTTTCAGTGATGATTTTCGGAGATC
CGTGGCGAGAGTTAGGAATGTCAGAATCTGTAAGGAAGTGCTTTCCCTATTAGCCAAGCTTTCAAATGGTTGGCGTCAGCATCATTCTCGCAAGAAGAGA
AGCCTACTGGTTCACAATGGCATTTCTTCTCCACTGATAGAGCAGTATAAAGTTAGTGGACAGCTAAATATGATCTGGACTGTGGACATTCTGCAGGAAA
ACTCATTCTGCATTCAAGTTTTGAAGGTTTGGGATATTTTACCATCATCTGATATACCAAAACTAGCACCGCGTCTCGACACCTTGTTTCGGAATTATAC
GGAGGAGCAGATGAATCGCTGCTTATACAAATGTATGGAGGGGAATTTGGTTGTTCCAATGAGATGGACAGTGGACTCGTCTAGTGATCATCAGGACAGC
TGCGGCGAAGCCGATGCTGTGCAGCTGCCGAAATCACTTGCATCACTCTGTCTAGACGATGGACAGTGGACTCGTCTAGTGATCGAGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G077600.1 pacid=42796329 polypeptide=Potri.004G077600.1.p locus=Potri.004G077600 ID=Potri.004G077600.1.v4.1 annot-version=v4.1
MDKRTDRVKWEKVKGRGFLDLVFSWSIEDVLNKDLYKDQVEEIPNSFMSTAHYMKAFITPLHVETHADLLSSTESLAGAPTYRILRVRKSKDYKPPKDLF
YEISMEETRGGYVPWVGDLIALTNVKLKCIDDLRKTQQSYHVAFVHAVKRGNRLTPSILSSKPIVDEEGLKNGTLFAVHLINLMTNLRIWRSLHLELEGR
NMNVIEKVLQNNFNDDGDCTICSSRKKSDAASACIRDTLQSSNLNSSQEAAVLSCIHTARCWHQYTVKLVQGPPGTGKTKTASCLLHALLRMKCRTLTCA
PTNIAVVEVAARVVSTVADLVEYETYGMGDIILFGNWERMKFDGDQNDLLHVFLDHRADILEKCFDPSTGWKRILASLISLLEDSEAQYHLYLQDNMGKE
GLLTCEQFVWKRFDFSGKQLKFCIVNLYTHLPTTLISLQVMRIMTRALDLMTSLETLLLSLSAADEGLKQILGENEDEERKLHNRIKLINEKRECLNTLR
LLSLKFQVPEFADKNAIEKFCLSNACLIFCTVSSSARLHSIRMAPLRCLVIDEAAQLKECESTIPLQLFGLHHAILIGDERQLPAIVNSEISGKAGFGRS
LFERLVKLGCKSHLLNIQYRMHPSISLFPNTEFYGRQVLDAPNVQETGYRRRFLQGDMFESYSFINIAHGKEEFVEKQSFKNTVEAAAAADIVGRLFKDI
NGTGKKVSIGIISPYQAQVHAIQAKIGKFISDSDSALSVSVGTVDGFQGGEEDLIIISTVRSNENGSVGFVSNPQRANVALTRARYCLWILGNEATLVKS
GSIWKEIVNDAKHRQCFYNAEEDESLAQAITESLIEHGRLDVLLRTHSPLFRNARWMVFFSDDFRRSVARVRNVRICKEVLSLLAKLSNGWRQHHSRKKR
SLLVHNGISSPLIEQYKVSGQLNMIWTVDILQENSFCIQVLKVWDILPSSDIPKLAPRLDTLFRNYTEEQMNRCLYKCMEGNLVVPMRWTVDSSSDHQDS
CGEADAVQLPKSLASLCLDDGQWTRLVIE
|
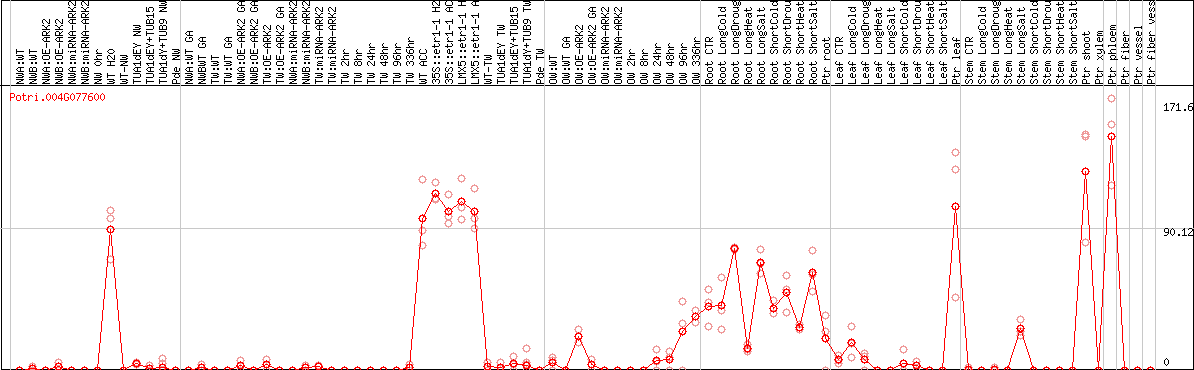
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G077600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
