Potri.004G077650 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G077650.1 pacid=42794799 polypeptide=Potri.004G077650.1.p locus=Potri.004G077650 ID=Potri.004G077650.1.v4.1 annot-version=v4.1
ATGGAGAAAACTGAAGAAGAATTAGTAGCTGGTAGAAGCTTGGTAGATTTGGTGTTCTCTTGGTCTATCGGGGATGTACTCAACAAAGATCTTTGCAGAA
ACAAGGTGAAGAAGATACCAGAGACGTTCATGTCAACAACACATTACATGAAGTCATTCATTCCCGCACTAATCGAGGAAACTCGTGCAGACTTATGCTC
AAACATGATAATGATCTCCCAAGCACCTACAAGAGAAATCTTTTCAGTGGGAATGGATAAAAAGAATAAGCCTCCGGAAGACTTGTTTTATAACATCTGG
TTTAAAAAAAGGAGAAACAAAGCCAATGGGAAAGAAATATATGAGCCTGACGTTGGAGATCTCCTTGCTTTGACAGATGTAAGACCGAAAGACATTGATG
ACTTGAACAGGCCTGGGTTTAACTATCTTCTTGCATATGTTCATAGGCTATCTGAATGGCAAGATGACGACGATAAATATGTAATTTTGTCAACTCTCAC
ATCCAAACCCATTCAATTTGAAATAGAAGACCAGGAAAATAAAAAGGAGTCTATCATTGCTGGAAAAGGAAGGCGAAAGACCATGAAAGCTAATGTCTAC
GTTGTTTACCTAGTGAACATGATGACAAATATTCGTATATGGAGATCATTGAACTCAGATCTGGAAGGAGGGAACATGAACATTATCCAGAATGTGCTTC
ATACTAGCTCAGCTATGTTGTGTCTGATCCTGCAGGATGGCCAAGATTGCTCTCACTGCTTATCTGAAGTGAATAAAAGTGCTACACTTTCTGGTATGGA
AGAAACCATAATCAGCTCTTCCAATTTAAATGACTCGCAGCAAGATGCAATTGTAAGCTGCATTGGTCTGAGTGAATGCCAACATCAAAGTACTGTGAAA
CTAATTTGGGGCCCTCCGGGGACTGGCAAAACAAAGATGATTGGTTTATTATTGTTCTCTCTCCTTAAATTGAAGTGCAGGACCCTTACATGTGCTCCAA
CCAATATTGCTGTGTTGGAAGTAACATCGAGACTCCTAAGGCTAGTTACAGACTCTCGTGAAGATGACACTTATGGACTTGGAGATATAATTCTATTTGG
GAATGGGAAGCGAATGAAGATCTCTGAGAATGATGATCTTGAAGATATATTTCTTGGCCATCGTGTTAAAGTACTTGAATATTGCTTTAGCCCGTCGAAT
GGGTGGAAGCACACTGTAGACTCATTGATAAATTTGCTTGAAGATCCTGAGAATCAGTACCGCCGGTACTTGGAAAATATGGAGAAAAAAAATGAGGAGG
GAGAGAGAGAGGATCAGGAAGATGAAATGCTTGAAATTGAAGAGATAAACAACAAGAAAGAGAAAGATGAAGTGGTCAATGATCAAAACAAAAAGGGCAG
GAACAGAGTTCTTCTTCAGGCCTTGAAAGATGACATGAAAAAGGAAAAACAAAAACAAAAACAAAAGGTTTTTTCTCATCAAGAGAATCTTACAAAGTGT
GAGGAAAAGGAATATAAAGATGGAAAGGTAAATAAGGAAGATATTCTCTCTTTCGAGGAATTCGTAAAGGAATGGTTCAAATTCCTTAGTGCGAAGTTGG
ATATTTTAATTGCAGGTTTATATACGCACTTGCCAACATCTATCATTTCATTAGAAGTGGTGAAGAGCATGACAAGAGCTGTTGATTCACTCAGCTGCCT
CAAACCCTTGTTGTATAGTGTTAGTGTTGGAGATGAAGGTTTAAAACAAGTCCTCAACGATTTTGAAAATGAAGGAAGCAGCGCTGGTCAATTTTCCCGG
TTGTCCTTTATGAGAAACTATTGTATTCAGACACTGAATTCACTTCCTCGAGAATTTGAAGTTCCCAATTTTTTTGATAATAGGGCAGCCAGATACTTTT
GCTTGGGAAATGCGTGTCTGGTTTTCTGTACTGCTTCAAGCTCTGCCAAGTTGCACACAGAAGGAGTGACACCGATAAAATTGTTGGTTATTGATGAAGC
TGCCCAGCTTAAAGAATGTGAATCGACTATTCCATTGCAACTTTCTGGTCTTCGCCATGCCATTCTTATAGGTGACGAGCGCCAACTTCCTGCCATGGTT
CAAAGCAAGATTTCCGAGGAAGCTGAATTCGGGAGGAGTTTGTTTGAGAGATTGGTAATACTGGAACACGAGAAACACCTTCTGAATACGCAGTATAGGA
TGCACCCATCTATAAGCTTATTCCCAAATAAAGAGTTCTATGATATGCTAATTCAGGATGCTTCAAATGTCAAAGAAAGAAACTATCAGAAACAATTCCT
TCAAGGCAATATGTATGGCCCTTACTCATTTATTAATGTAGCCAATGGGAAAGAGCAATCCAACGACGGCCGCAGCAAGAAAAATTTGGTCGAGGTTGCT
GTAGTTTCAGCAATAGTTGCAGGCCTTTTTAAAGAATTTAAAAGGGCAAGAAAGAGGATGAGCATAGGAGTCATATCACCATACAATGCTCAAGTGTATG
CAATTCAACAAAAAATTGGAAATACTTACAGTACATTTAGTGACTTTGCTGTAAATGTTCGGTCTGTCGATGGATTTCAAGGCAGCGAGGAGGATGTTAT
CATTATCTCTACAGTCAGATGCAATGCAAGTGGATCAGTGGGTTTTCTTTCCAATCGCCAAAGGGCAAATGTTGCGCTGACCCGTGCACGGTACTGCCTC
TGGATACTGGGAAATGGAGCAACTCTTGTCAACAGTGGCTCTATTTGGAAGAAATTGGTTACTGACGCGAAGGAACGAGGGTGTTTCTACAATGCTGATG
AGGATAAAAGCTTGTCCAAGGCAATAATGGATGCCTTGTTAGAGTTGGACCAACTTGATGATTTGTTGAATGTCAATTTTCTACTGTTCAGAAATGCAAG
ATGGAAGTTTTGCTTCAGTGAAAACTTTCGGAAATCTATTATGAAAGTGGGAAATGAGGCTCGTCAGGAAGTGATTTCTTTGTTGGCAAAGCTTTCAAGT
GGCTGGCGTCAATCTCCTGAAGAGAGAAACATCATTGTCCTACACGGAACTTCTTCTGAACTGTTAGAAAATTACAGAGTCAATGACCAGCTCAGTCTCA
TTTGGACAGTGAATATAATCAAGGAAAACAAAAACGACACTCAAATTCTTAAGGTTTGGGATGTTTTATCATTACATGATTTACCGAAACTAGCAAGGAG
CCTCGATGCTGTCGTTGGAAATTATACAGTGAATAAGATGAACCGCTGCAGACACAAATGCACAGAAGGGGATTTGGTTGTTCCAATGAGATGGTCAATA
AGTTCCGGTGCCTCTCTGGAGAGTAGTAATCCTGAAACCGATCCTGCACAGCTTCTGTCACAACCATTAGCTTCACTTGTTATCAGGGATGAATCAGAAG
CACCAGCAACAACTAGTAGACAGCCCTTGAGGTCTAAGAAAGATGGATTTAGCAGTGGAACAAGGGGATCAAAACCTACGTGGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G077650.1 pacid=42794799 polypeptide=Potri.004G077650.1.p locus=Potri.004G077650 ID=Potri.004G077650.1.v4.1 annot-version=v4.1
MEKTEEELVAGRSLVDLVFSWSIGDVLNKDLCRNKVKKIPETFMSTTHYMKSFIPALIEETRADLCSNMIMISQAPTREIFSVGMDKKNKPPEDLFYNIW
FKKRRNKANGKEIYEPDVGDLLALTDVRPKDIDDLNRPGFNYLLAYVHRLSEWQDDDDKYVILSTLTSKPIQFEIEDQENKKESIIAGKGRRKTMKANVY
VVYLVNMMTNIRIWRSLNSDLEGGNMNIIQNVLHTSSAMLCLILQDGQDCSHCLSEVNKSATLSGMEETIISSSNLNDSQQDAIVSCIGLSECQHQSTVK
LIWGPPGTGKTKMIGLLLFSLLKLKCRTLTCAPTNIAVLEVTSRLLRLVTDSREDDTYGLGDIILFGNGKRMKISENDDLEDIFLGHRVKVLEYCFSPSN
GWKHTVDSLINLLEDPENQYRRYLENMEKKNEEGEREDQEDEMLEIEEINNKKEKDEVVNDQNKKGRNRVLLQALKDDMKKEKQKQKQKVFSHQENLTKC
EEKEYKDGKVNKEDILSFEEFVKEWFKFLSAKLDILIAGLYTHLPTSIISLEVVKSMTRAVDSLSCLKPLLYSVSVGDEGLKQVLNDFENEGSSAGQFSR
LSFMRNYCIQTLNSLPREFEVPNFFDNRAARYFCLGNACLVFCTASSSAKLHTEGVTPIKLLVIDEAAQLKECESTIPLQLSGLRHAILIGDERQLPAMV
QSKISEEAEFGRSLFERLVILEHEKHLLNTQYRMHPSISLFPNKEFYDMLIQDASNVKERNYQKQFLQGNMYGPYSFINVANGKEQSNDGRSKKNLVEVA
VVSAIVAGLFKEFKRARKRMSIGVISPYNAQVYAIQQKIGNTYSTFSDFAVNVRSVDGFQGSEEDVIIISTVRCNASGSVGFLSNRQRANVALTRARYCL
WILGNGATLVNSGSIWKKLVTDAKERGCFYNADEDKSLSKAIMDALLELDQLDDLLNVNFLLFRNARWKFCFSENFRKSIMKVGNEARQEVISLLAKLSS
GWRQSPEERNIIVLHGTSSELLENYRVNDQLSLIWTVNIIKENKNDTQILKVWDVLSLHDLPKLARSLDAVVGNYTVNKMNRCRHKCTEGDLVVPMRWSI
SSGASLESSNPETDPAQLLSQPLASLVIRDESEAPATTSRQPLRSKKDGFSSGTRGSKPTW
|
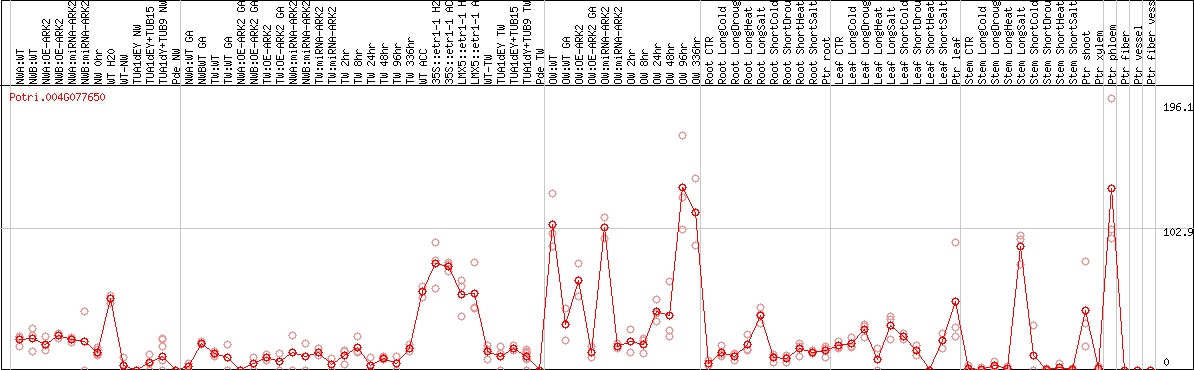
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G077650 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
