Potri.004G094200 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G094200.2 pacid=42796062 polypeptide=Potri.004G094200.2.p locus=Potri.004G094200 ID=Potri.004G094200.2.v4.1 annot-version=v4.1
ATGGCCTTTCAATCCGATCACCAAACCCCACAGTCCATAAACTCACAAACCCTATACTACAAATCCCTCACTTCACTTTCCCTCCAAATCCCTCCAACCC
CATCATCCTTCCTCCCTCTCCCTCCTTCTCTCCCTCCTTCAAAACTCATCCCCAAAACCCGTTTCCTCATCGACGGATTCCGCTTTTCCTCCCCTTCCAT
TACTGCCTACTTCCTCTCCCACTTCCATTCTGACCATTACACTGGTCTTTCCTCTCATTGGTCTCAAGGTATCATTTTTTGTTCCCCAATCACTACCTCT
CTTGTTACTAGTATCCTTAATGTCCCTGAATGTTTTGTCTTTTCGCTTCCTCTGAATCGTGCTGTTGATATTGATGGTGTTGAAGTTAGTTTAGTTGATG
CTAATCACTGTCCTGGCGCTGTTCAGTTCTTGTTGAAAGTGCCCATTTGTAAGAATTTAGATAATTTTGAGTTGTATGTTCATACTGGTGATTTTAGGTA
CTCTTGTGAGATGAAAGATGATGTTTTTTTGAGAGGTTTTGTGGGGTGTAATACGGTTTTTTTGGATACTACTTATTGTAATCCGAAATTTGTGTTTCCT
TTACAAGAGGAGTCGGTTGATTATGTTGTTAGTGCGATTGAGAAGATAGGAGGAGAGGGGTTCAGTGGGGGTTTGGAGAAGAGGGTTCTGTTCCTCGTGG
CTACTTATGTGGTGGGGAAAGAAAAGATTTTGATTGAGATTGCACGGAGGTGTAATAGGAAGGTCTATGTGGATGCCAGGAAAATGGAGGTTTTGAGGGT
TTTGGGGTGCGGGGAGAGTGGGGTGTTTACAGAGGATGAGAATGAGAGTGATGTGCATGTTGTTGGCTGGAATGTCTTGGGGGAAACTTGGCCTTATTTT
CGGCCGAATTTTGTGAAAATGAAGGAAATTATGGTGGAAAGAGGTTACAATAAGGTTGTGGGGTTTGTGCCAACTGGGTGGACATACGAAGTGAAACGTA
ATAAGTTTGCAGTGAGATCAAAGGATTCATGTGAGATTCATCTTGTGCCATATAGTGAGCATTCAAATTACAATGAACTCAGGGAGTATGTGAAGTTTTT
GAGACCTAAACGTGTCATTCCTACTGTTGGTGTGGATGTTGAAAAACTCGATAGTAAACATGCTGCTAAAATGCAGAAGCATTTTGCTGGGTTAGTTGAC
GAGATGGCTAATAAGAAGGAATTTTTAATGGGTTTTCTTCGTGGGTCGTCTGAGAATGATAAGAAGGTTGAAATGGATGCTGTCTCAGGCTTGAACGAGG
GGCTGGCGCAGGAGAAAGAGTTAGTGGAGTCAGTTGAAATGAAAGCCCATGAGAATAATGACACTGTTGCTTGTCTCAACTCCTCATCTATACTGCAAGA
ATCTCCTACACACAATTTAAGCATGCTAAATGATGAGGAAAGAGATAAACTCGTTCATGAATTGAGTGATTGTTTGCCCACATGGGTTACCCGAGACCAA
ATGTTAGATTTAATCAGCACTCATGGGAGGAATATTGTTGAAGCAGTTTCTAGCTTTTATGAACGTGAAATGGAATTCCATGATCAAGTAATTTCTTGTA
GAGTAACTGTTTCTTCATCTGAGACAGTTCCATTATATGACTCTGAATCACCTTCAAAACCAGCTTCTATTAATACTGATAGTCGAAGTATGGGTTTTCT
GTCAGGTCAAAACTACAAGTCACCTAGCAAGAATCCTAAACTAAAGGGTGGTAATTCTCCTGGAAAAAGGAAGAGGAGCGTTGGCAATAAGCCTGGCAAG
AAATCAAAAATTAATTCAAAGCTGGAATCTGGCTTGTCGAAGCAATCCACCATTACTAAGTTTTTCAATAAAGTTTTGCCTGATGCTTCTCAAGTTAGTG
TGGTTGCATCTATATCTGAGCAATGCCCTGGAGATGAAAATTTGTTGCAAAATGATGATGTTACAGAGTCATACAGGGAGGAGGTAGATCAGTTTATTCA
GATAATTGATGGCAATGAGTCAACAAGAAGTTATGCTGCCACCATCCTCAAGAAGACAGAAGGAGATATTAACAAGGCTCTGGACATGCATTATGGTGAT
CCCATGGGTAACCTTGGCAAGAGTATAGAGGCTTTGGTGGTTTCTGGCAATATGGTTGAGCGCCAGTGCGAAACTGGCTCCTCTTCTGCCAGGGAGAAAG
AATTGTTTGGAGAAATAGAAAATATGGTTGATCTATCCGTTCAGGGATCATTAATAAAAAACGTGGATGCAACTCTTGTATCACTACCAACAGAAAAATA
TAATCCCATAGAGCATGCATGCTGGAATGGTGGACAACCTGCACCATATATCCATCTTGCACGTGCTTTTGACCTGGTCGAGGCAGAAAAAGGAAAGATC
AAAGTTACTTCGTTATTGTGCAATATGTTTAGAAGTTTGCTGGCATTGTCTCCTGAGGATGTGCTGCCTGCTGCCTATCTGTGCACAAATAAGATTGCTG
CTGACCATGAAAATGTGGAACTAAACATAGGTGGAACCTTGGTTACATCGGCTTTGGAAGAGGCATGTGGAACAAAGCGGTCTAAAATAAGGGAAATGTA
CAATAGTATGGGGGATCTTGGTGACGTTGCTCAAGTATGCCGACAAACACAGACATTACTGGCTCCTCCTCCTCCCCTTCTTATTAAAGATGTATTTTCT
GCGCTGCAGAAGATAAGTGTACAAACAGGTAGTGGAAGTACTGGTCGAAAGAAGAGCCTCATTGTGAATCTTATGCGTTCTTGTAGGGAGAAGGAGATGA
AGTTTATTGTCAGAACTTTGGTAAGGAATTTACGGATTGGAGCAATGATGAGAACTATCCTACCTGCTTTAGCTCAAGCTGTTGCTTTGAATTCCTTTTC
TTCTGATGAATGTAAAGCTGAGAATGTGAAGGAGAAACTTCAGTACATTTCCACAGCAGTGGTTGAAGCTTATAACATCCTTCCTACTCTGGATTTGGTT
GTTCCTTCACTCATAAATGAAGGCGTTGGATTTTCATCGTCAACTTTATCAATGGTTGCTGGGATACCTCTTAAACCAATGCTTGCAAAAATTACAAATG
GAGTTGCTCAAGTACTAAAGCTCTTTGAAAATAAAGCTTTTACATGTGAATACAAATACGATGGTCAGCGTGCCCAAATTCACAAAATGCCAAATGGCAC
TGTACGTATATATTCACGAAATGGGGATGAAACAACTTCTAGGTTTCCTGATTTGATCAAGATAATTGAGGAATCTTGTAAACCTGCTGCAGCGACTTTA
ATAGTGGATGCAGAGGTAGTTGCAGTTGATAGAAAGAATGGCTGCAAGCTCATGTCCTTTCAAGAGCTATCTTCAAGGGAGAGAGGGAGCAAAGATTCTT
CCATTGCTGTGAATAAGATAAAGGTTGATATATGCGTGTTTGTCTTTGATATCATGTTTGCTAATGGAGAACAGCTGTTGGAGCTTCCTCTCCGCCAAAG
GCGACAATACTTGAAAGACTTGTTTTGTGATGAGAGATTAGGTTGTTTTGAATATGCGAAGGAGATGACGGTTGAAGCACAGGATGCTTGTTTGACCAAT
GATGCAACTTTGACCAAGATGAAATCTTTCCTTGAAGATGCATTGCGGTCCTCGTGTGAAGGGATTATGGTGAAATCTCTAGATATCGATGCTGGATACT
GTCCATCAAAACGGACTGACGCATGGTTAAAGGTCAAGAAGGATTATGTTGAAGGACTGAATGATTCTCTTGATTTAGTCCCGATTGGTGCTTGGCATGG
AAATGGGAGAAAAGCAGGATGGTATAGTCCATTCCTCATGGCATGCTACAACCCTGAGACTGAGGAATTCCAAAGTGTGTGCCGTGTCATGTCTGGGTTC
TCAGATGCTTTTTATATAGAGATGAAAGAATTCTTCTCTGGAGATAGGATCTTGGCCAAAAAGCCACCATACTATCGAACTGTGGAGGCGCCGGATATGT
GGTTTTCTCCAGAAGTTGTTTGGGAAATAAGAGGTGCTGACTTTACAATTTCACCTGTTCACCAGGCTGCTGTAGGTTTAGTTCATCAATCTCGTGGCAT
CTCAATCAGATTTCCAAGATTTATTCACTCCATATCGGATAGAAATCCAGAGGAATGTAGCACAGCAGCAGATATAGCTGAAATGTTTAATTCTCAAACA
AGGAAGATGGACGTGACAGCTGAAAGATGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G094200.2 pacid=42796062 polypeptide=Potri.004G094200.2.p locus=Potri.004G094200 ID=Potri.004G094200.2.v4.1 annot-version=v4.1
MAFQSDHQTPQSINSQTLYYKSLTSLSLQIPPTPSSFLPLPPSLPPSKLIPKTRFLIDGFRFSSPSITAYFLSHFHSDHYTGLSSHWSQGIIFCSPITTS
LVTSILNVPECFVFSLPLNRAVDIDGVEVSLVDANHCPGAVQFLLKVPICKNLDNFELYVHTGDFRYSCEMKDDVFLRGFVGCNTVFLDTTYCNPKFVFP
LQEESVDYVVSAIEKIGGEGFSGGLEKRVLFLVATYVVGKEKILIEIARRCNRKVYVDARKMEVLRVLGCGESGVFTEDENESDVHVVGWNVLGETWPYF
RPNFVKMKEIMVERGYNKVVGFVPTGWTYEVKRNKFAVRSKDSCEIHLVPYSEHSNYNELREYVKFLRPKRVIPTVGVDVEKLDSKHAAKMQKHFAGLVD
EMANKKEFLMGFLRGSSENDKKVEMDAVSGLNEGLAQEKELVESVEMKAHENNDTVACLNSSSILQESPTHNLSMLNDEERDKLVHELSDCLPTWVTRDQ
MLDLISTHGRNIVEAVSSFYEREMEFHDQVISCRVTVSSSETVPLYDSESPSKPASINTDSRSMGFLSGQNYKSPSKNPKLKGGNSPGKRKRSVGNKPGK
KSKINSKLESGLSKQSTITKFFNKVLPDASQVSVVASISEQCPGDENLLQNDDVTESYREEVDQFIQIIDGNESTRSYAATILKKTEGDINKALDMHYGD
PMGNLGKSIEALVVSGNMVERQCETGSSSAREKELFGEIENMVDLSVQGSLIKNVDATLVSLPTEKYNPIEHACWNGGQPAPYIHLARAFDLVEAEKGKI
KVTSLLCNMFRSLLALSPEDVLPAAYLCTNKIAADHENVELNIGGTLVTSALEEACGTKRSKIREMYNSMGDLGDVAQVCRQTQTLLAPPPPLLIKDVFS
ALQKISVQTGSGSTGRKKSLIVNLMRSCREKEMKFIVRTLVRNLRIGAMMRTILPALAQAVALNSFSSDECKAENVKEKLQYISTAVVEAYNILPTLDLV
VPSLINEGVGFSSSTLSMVAGIPLKPMLAKITNGVAQVLKLFENKAFTCEYKYDGQRAQIHKMPNGTVRIYSRNGDETTSRFPDLIKIIEESCKPAAATL
IVDAEVVAVDRKNGCKLMSFQELSSRERGSKDSSIAVNKIKVDICVFVFDIMFANGEQLLELPLRQRRQYLKDLFCDERLGCFEYAKEMTVEAQDACLTN
DATLTKMKSFLEDALRSSCEGIMVKSLDIDAGYCPSKRTDAWLKVKKDYVEGLNDSLDLVPIGAWHGNGRKAGWYSPFLMACYNPETEEFQSVCRVMSGF
SDAFYIEMKEFFSGDRILAKKPPYYRTVEAPDMWFSPEVVWEIRGADFTISPVHQAAVGLVHQSRGISIRFPRFIHSISDRNPEECSTAADIAEMFNSQT
RKMDVTAER
|
DESeq2's median of ratios [POPLAR]
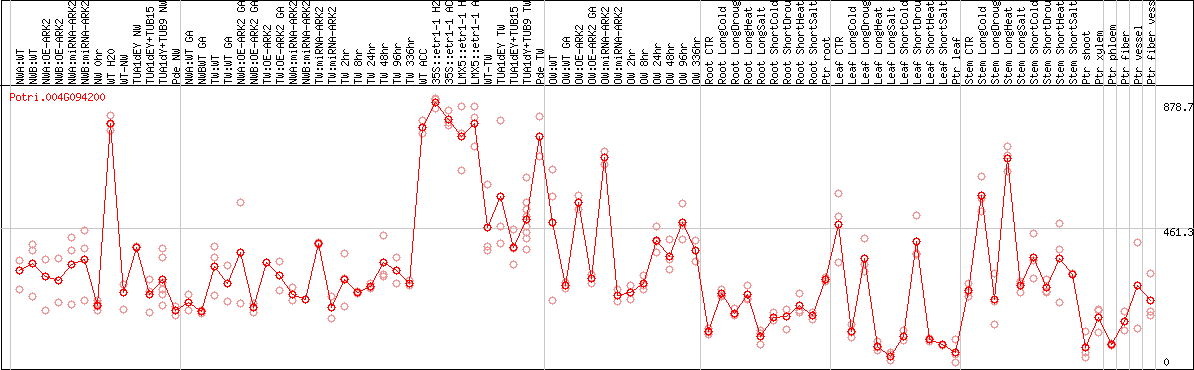
Coexpressed genes
Potri.004G094200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
