External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G18400 152 / 5e-45
bHLH
bHLH044, BEE1
BR enhanced expression 1 (.1)
AT1G73830 145 / 3e-42
bHLH
bHLH050, BEE3
BR enhanced expression 3 (.1.2)
AT1G25330 143 / 6e-42
bHLH
HAF, bHLH075, CESTA
HALF FILLED, CESTA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G50915 117 / 3e-31
bHLH
bHLH137
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT4G34530 117 / 7e-31
bHLH
bHLH063, CIB1
cryptochrome-interacting basic-helix-loop-helix 1 (.1)
AT4G36540 116 / 7e-31
bHLH
bHLH058, BEE2
BR enhanced expression 2 (.1.2)
AT5G48560 119 / 1e-30
bHLH
bHLH078
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G68920 119 / 1e-30
bHLH
bHLH049, ACE1
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT3G07340 118 / 1e-30
bHLH
bHLH062
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G59640 115 / 2e-30
bHLH
ZCW32, bHLH031, BPE, BPEUB, BPEP
BIG PETAL UB, BIG PETAL P (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G115300
405 / 3e-144
AT1G25330 151 / 4e-45
HALF FILLED, CESTA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.012G055700
194 / 7e-61
AT1G73830 216 / 1e-69
BR enhanced expression 3 (.1.2)
Potri.015G046300
191 / 7e-60
AT1G18400 192 / 3e-60
BR enhanced expression 1 (.1)
Potri.007G023600
122 / 2e-32
AT2G18300 182 / 3e-54
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.002G235400
120 / 8e-31
AT3G07340 257 / 2e-79
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G148900
120 / 9e-31
AT3G07340 254 / 2e-78
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G248500
119 / 1e-30
AT3G07340 349 / 5e-115
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.012G104900
117 / 2e-30
AT5G50915 177 / 3e-53
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.005G121850
117 / 2e-30
AT4G34530 177 / 3e-52
cryptochrome-interacting basic-helix-loop-helix 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038939
150 / 8e-44
AT1G73830 193 / 9e-61
BR enhanced expression 3 (.1.2)
Lus10009673
145 / 2e-42
AT1G73830 204 / 1e-65
BR enhanced expression 3 (.1.2)
Lus10009034
136 / 7e-39
AT1G73830 186 / 2e-58
BR enhanced expression 3 (.1.2)
Lus10017602
134 / 2e-38
AT1G18400 122 / 7e-34
BR enhanced expression 1 (.1)
Lus10033562
133 / 4e-38
AT1G18400 121 / 5e-34
BR enhanced expression 1 (.1)
Lus10026008
118 / 2e-31
AT2G18300 168 / 8e-50
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10003172
118 / 3e-31
AT5G48560 205 / 4e-62
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10014300
117 / 4e-31
AT2G18300 171 / 3e-51
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10023562
117 / 5e-31
AT1G26260 178 / 9e-54
cryptochrome-interacting basic-helix-loop-helix 5 (.1.2.3)
Lus10041261
116 / 5e-31
AT3G23690 172 / 3e-51
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.004G099400.2 pacid=42796089 polypeptide=Potri.004G099400.2.p locus=Potri.004G099400 ID=Potri.004G099400.2.v4.1 annot-version=v4.1
ATGGCTGAGTTTGCAGAATATCTGCAGAGGTTTGGGGCATCTCAACCTTTGACAGAGATGATGGACATGAACATGGAAATGCTCAAGCATCTTCCAGAAC
TGAATCCAAGCATCTTGGAGAATTTCAGCATCACTCATGACTTCTCAGCAGACTCTTTCTTCGCCCACCAACAACCTGAATTTCCTGCAACATATAATCA
CAACAACCTTTCAAGTACTTCCCATCCTCACATCTTGAGTACAGCACCACTTGTTCATTCTGTCAGCTTGAATCAAAATGTTTTCCATGAAAGGAAGAAG
CCAAAAGCAATGGAACAATCAACCGGCAGCTCCAAGAACGTTTCTCCTACAGCCTCTATAAACATCACAGAGATAAAGAATAACTTAGGAAGAGGTAAGA
AAGGCAAAAACAAGGAGAAAGAAGGGGACAAATCAAAGGAAGTTATTCATGTTAGAGCTAAGAGAGGCCAGGCAACAGACAGTCACAGTATAGCAGAAAG
GATAAGAAGAGAGAAAATCAATAACAAATTAAGATGCTTGCAAGACATTGTTCCTGGATGCCACAAGTCAATGGGGATGGCAGTTATGCTAGAGGAGATA
ATCAATTATGTACACTCATTGCAGAATCAAGTTGAGTTTCTCTCCATGGAGCTTGCCGCTGCCAGTTGTTCTAATGACCTGAAAAATCTCACAGAATCTT
CCAAGAAGGCTCAGGGAACAAATTCAACTGATGATGCACAGGAGACACAGAAATGGTCAAGAGAAAGATATGGAGAAATCACTTGCTTCCACTCTACATG
GTCCATTTGA
AA sequence
>Potri.004G099400.2 pacid=42796089 polypeptide=Potri.004G099400.2.p locus=Potri.004G099400 ID=Potri.004G099400.2.v4.1 annot-version=v4.1
MAEFAEYLQRFGASQPLTEMMDMNMEMLKHLPELNPSILENFSITHDFSADSFFAHQQPEFPATYNHNNLSSTSHPHILSTAPLVHSVSLNQNVFHERKK
PKAMEQSTGSSKNVSPTASINITEIKNNLGRGKKGKNKEKEGDKSKEVIHVRAKRGQATDSHSIAERIRREKINNKLRCLQDIVPGCHKSMGMAVMLEEI
INYVHSLQNQVEFLSMELAAASCSNDLKNLTESSKKAQGTNSTDDAQETQKWSRERYGEITCFHSTWSI
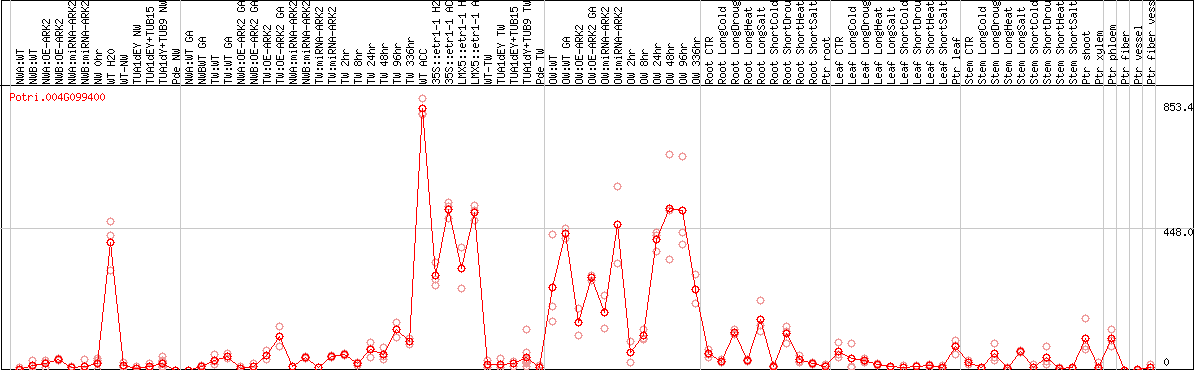
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G099400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.