PHO1.2 (Potri.004G123600) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | PHO1.2 | ||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G123600.6 pacid=42796095 polypeptide=Potri.004G123600.6.p locus=Potri.004G123600 ID=Potri.004G123600.6.v4.1 annot-version=v4.1
ATGGCTACTACTAGTTCACATTTCTCACCGACGTCGCATTGGTGCAGCAATGGCAGTTCAATATCAAGATTGGTCGATTTTGGATCAAAATGGAGGAGGA
AGCAGCAGCTCTTCTCCATGAATTCGCGTCGTGTGGTAAAAAGGAGGTCGGTCTCGGTCAGTATTAAGAATGTGTCAAGTAGCGAACCTAAGCAGAAACT
CAAAGATGATGCTCTCATCGAAGAAGAAGAAGTCCCAAGAATTTTGAATCCTTCCACTCCAAATGCTTCATCTATTGCCTCGAGTATCAAATACCATGCA
GAATTCACACCGTTGTTTTCTCCGGAGCGATTTGAGCTTCCCAAGGCTTACTATGCTACTGCACAAAGTGTCCGTGATGCTCTCATTATAAATTGGAATT
CAACATACGAGTCTTATGAGAGGTTAAATGCAAAACAGGCATATTATCTATCAATGGAATTTCTACAGGGTAGAGCACTGTTGAATGCCATTGGTAATTT
AGAGCTGACTGGTGCTTATGCAGAGGCTTTGAGCAAGCTTGGGCACAGCCTAGAAAATGTGGCTTGCCAGGAGCCTGATGCTGCACTTGGAAATGGGGGT
CTTGGGCGACTTGCGTCCTGCTTTTTGGACTCGTTGGCAACGTTAAATTACCCAGCATGGGGCTATGGATTAAGATACAAGTATGGCTTATTTAAACAGC
AGATTACAAAAGATGGTCAGGAGGAAGTTGCTGAAGATTGGCTTGAGATGGGCAATCCCTGGGAAATTCTTAGAAATGATATCTCATATCCTATAAAATT
TTACGGCAAGGTTGTCTCTGGATCAGATGGGAAAAAACACTGGATTGGTGGAGAAGATATTAAGGCTGTGGCATATGATGTCCCAATACCGGGATATAAA
ACTAAAACCACTATTAACCTTAGACTTTGGTCAACAAAAGCTCCATCAGAAGATTTGGACTTATATGCTTTTAATGCAGGAGACCACACCAAAGCATATG
AGGCCCTCTCAAATGCTGAAAAGATTTGCCACGTTCTTTACCCTGGGGATGACTCACTGGAGGGCAAGATTCTTCGGTTGAAGCAACAATATACCTTGTG
TTCAGCTTCTCTCCAAGATATCATATCATGTTTTGAGAGAAGATCAGGATCGAACATAGATTGGGAAAAATTTCCAGAGAAAGTTGCAGTGCAGATGAAT
GACACTCATCCCACACTGTGCATCCCTGAGTTGATGAGAATTTTGATAGATTTGAAAGGTTTAAGCTGGAAGGAGGCCTGGAATATCACTCAAAGAACTG
TGGCATACACAAACCATACTGTTCTACCAGAGGCTTTGGAGAAATGGAGTCTAGAACTTATGCAGAAACTACTACCACGACATGTTGAGATCATTGAATT
GATTGATGAAGAGCTGATTTGCACCATAGTATCAGAGTATGGTACCGAAGATTCCGATTTGTTGGAGAAGAAACTGAAGGAAATGAGAATATTAGAAAAT
GTTGATCTCCCTTCAGCTTTTGCAGAGTTGATTGTTAAGCCTAAGCAAAGCTCTGTGGGTGTTACCAGTGAGGAATTAGAAAATTCTGATGAAGAAACCA
AGCTAGTTTATAATTCTGAAGAAGAAACTCTCTCTAAGAGAGCTAATGATTTTGAAGAAGAAACTAAGAGAGCTAATGATTTGGAAGAAGAAACTAAGAG
AGCTAATGATTTGGAAGAAGAAACTAAGAGAGCTAATGATTTTGAAGAAGAAATGGAGCTGGTTGATGAAAAAGATGAATCCAAAAGCAAAGTAACTCAG
AAGAAGGAGAAGATAATGGCAGAACCACCCCCTAAACCACCAAAGATGGTCCGTATGGCTAATCTTGCTGTTGTGGGTGGTCATGCAGTAAATGGAGTTG
CTGAGATACATAGTGAAATAGTCAAGGATGAAGTATTTAATGCATTTTATAAGTTGTGGCCAGATAAGTTTCAAAATAAGACAAATGGGGTTACACCAAG
AAGATGGATTCATTTCTGCAATCCAGGTTTGAGTAAAATTATTACTGACTGGATTGGCATGGATGATTGGGTTTTAAATACTGAAAAACTGGCAGAACTA
AGAAAGTTTTCCGATAATGAGGACCTCCAAGTGCAGTGGAAGGCAGCCAAAAGGAGCAACAAGATGAAGGTGATATCATTCCTCAAAGAAAAGACAGGAT
ATTCTGTTAGCCCTGATGCAATGTTTGATATCCAGGTGAAACGTATTCACGAGTACAAGCGACAATTACTGAATATTTTGGGAATTGTTTACCGCTACAA
GAAGATGAAAGAAATGACTGCTGCAGAAAGGAAAGCTAAGTATGTTCCACGAGTTTGTATATTTGGGGGAAAAGCATTTTCTACATATGTGCAAGCCAAG
AGAATAGTGAAATTTATCACAGATGTTGGGGCTACAGTTAATCATGATCCAGAAATAGGTGATTTACTGAAGGTTGTTTTTGTCCCTGATTACAATGTCA
GTGTTGCTGAATTGCTTATTCCTGCAAGTGAGCTCTCACAGCATATCAGTACTGCTGGGATGGAAGCCAGTGGAACAAGCAATATGAAGTTTGCAATGAA
TGGTTGTATCCTTATTGGGACCTTAGATGGCGCCAATGTTGAAATCCGAGAAGAAGTAGGAGAAGACAACTTCTTTCTCTTTGGTGCTAGAGCTCATGAA
ATTGCAGGGCTAAGGAAAGAGCGAGCTGATGGGGAGTTTGTCCCAGACCCAAGTTTTGAAGAAGTGAAAGATTTTGTCAAAAGTGGAGTTTTTGGGCCTT
GCAACTATGATGAACTTATAGGATCATTAGAAGGAAATGAAGGTTTTGGCCGTGCAGATTATTTCCTAGTGGGCAAGGACTTCCCCAGTTACATAGAATG
CCAAGAGGAGGTCGATAAAGCATACCATGACCAAAAGACATGGACGAAGATGTCAATCATGAATACAGCAGGCTCGTACAAGTTCAGCAGCGACAGGACC
ATTCATGAATATGCCCGAGAGATTTGGAATATCGAACCTGTCGAATTGCCATAA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G123600.6 pacid=42796095 polypeptide=Potri.004G123600.6.p locus=Potri.004G123600 ID=Potri.004G123600.6.v4.1 annot-version=v4.1
MATTSSHFSPTSHWCSNGSSISRLVDFGSKWRRKQQLFSMNSRRVVKRRSVSVSIKNVSSSEPKQKLKDDALIEEEEVPRILNPSTPNASSIASSIKYHA
EFTPLFSPERFELPKAYYATAQSVRDALIINWNSTYESYERLNAKQAYYLSMEFLQGRALLNAIGNLELTGAYAEALSKLGHSLENVACQEPDAALGNGG
LGRLASCFLDSLATLNYPAWGYGLRYKYGLFKQQITKDGQEEVAEDWLEMGNPWEILRNDISYPIKFYGKVVSGSDGKKHWIGGEDIKAVAYDVPIPGYK
TKTTINLRLWSTKAPSEDLDLYAFNAGDHTKAYEALSNAEKICHVLYPGDDSLEGKILRLKQQYTLCSASLQDIISCFERRSGSNIDWEKFPEKVAVQMN
DTHPTLCIPELMRILIDLKGLSWKEAWNITQRTVAYTNHTVLPEALEKWSLELMQKLLPRHVEIIELIDEELICTIVSEYGTEDSDLLEKKLKEMRILEN
VDLPSAFAELIVKPKQSSVGVTSEELENSDEETKLVYNSEEETLSKRANDFEEETKRANDLEEETKRANDLEEETKRANDFEEEMELVDEKDESKSKVTQ
KKEKIMAEPPPKPPKMVRMANLAVVGGHAVNGVAEIHSEIVKDEVFNAFYKLWPDKFQNKTNGVTPRRWIHFCNPGLSKIITDWIGMDDWVLNTEKLAEL
RKFSDNEDLQVQWKAAKRSNKMKVISFLKEKTGYSVSPDAMFDIQVKRIHEYKRQLLNILGIVYRYKKMKEMTAAERKAKYVPRVCIFGGKAFSTYVQAK
RIVKFITDVGATVNHDPEIGDLLKVVFVPDYNVSVAELLIPASELSQHISTAGMEASGTSNMKFAMNGCILIGTLDGANVEIREEVGEDNFFLFGARAHE
IAGLRKERADGEFVPDPSFEEVKDFVKSGVFGPCNYDELIGSLEGNEGFGRADYFLVGKDFPSYIECQEEVDKAYHDQKTWTKMSIMNTAGSYKFSSDRT
IHEYAREIWNIEPVELP
|
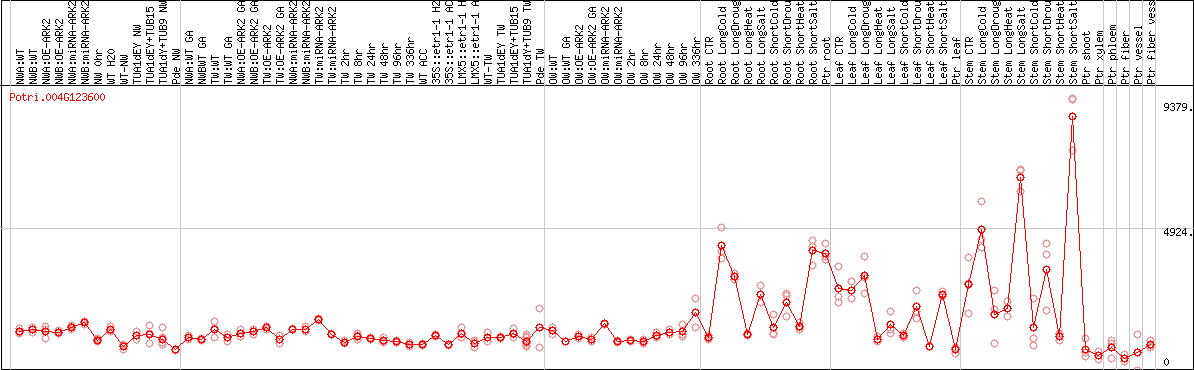
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G123600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
