External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G18800 243 / 2e-81
NRP2
NAP1-related protein 2 (.1)
AT1G74560 235 / 7e-78
NRP1
NAP1-related protein 1 (.1.2.3)
AT2G19480 46 / 1e-05
NFA2, NFA02, NAP1;2
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
AT3G13782 45 / 1e-05
NFA4, NFA04, NAP1;4
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 4, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 04, nucleosome assembly protein1;4 (.1)
AT4G26110 43 / 8e-05
NAP1;1, ATNAP1;1
ARABIDOPSIS THALIANA NUCLEOSOME ASSEMLY PROTEIN 1;1, nucleosome assembly protein1;1 (.1.2)
AT5G56950 40 / 0.001
NFA3, NFA03, NAP1;3
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A3, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 03, nucleosome assembly protein 1;3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G080700
323 / 8e-113
AT1G18800 292 / 4e-100
NAP1-related protein 2 (.1)
Potri.012G069100
248 / 9e-83
AT1G18800 325 / 7e-113
NAP1-related protein 2 (.1)
Potri.015G062700
239 / 2e-79
AT1G18800 324 / 2e-112
NAP1-related protein 2 (.1)
Potri.006G148600
50 / 4e-07
AT2G19480 447 / 1e-157
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
Potri.018G067800
47 / 3e-06
AT2G19480 462 / 4e-163
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
Potri.003G040700
43 / 9e-05
AT2G19480 287 / 7e-93
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
Potri.003G021400
43 / 0.0001
AT2G19480 286 / 2e-94
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
Potri.003G040250
43 / 0.0001
AT2G19480 280 / 9e-87
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
Potri.003G036500
41 / 0.0004
AT2G19480 280 / 5e-90
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023599
266 / 4e-90
AT1G18800 334 / 2e-116
NAP1-related protein 2 (.1)
Lus10024229
263 / 7e-89
AT1G18800 336 / 4e-117
NAP1-related protein 2 (.1)
Lus10008620
213 / 7e-70
AT1G18800 272 / 1e-92
NAP1-related protein 2 (.1)
Lus10027615
44 / 7e-05
AT2G19480 385 / 3e-134
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
Lus10011957
43 / 9e-05
AT2G19480 449 / 2e-158
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
Lus10011956
43 / 0.0001
AT2G19480 422 / 8e-148
NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 2, NUCLEOSOME/CHROMATIN ASSEMBLY FACTOR GROUP A 02, nucleosome assembly protein 1;2 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00956
NAP
Nucleosome assembly protein (NAP)
Representative CDS sequence
>Potri.004G128701.1 pacid=42793965 polypeptide=Potri.004G128701.1.p locus=Potri.004G128701 ID=Potri.004G128701.1.v4.1 annot-version=v4.1
ATGGTGGCTGACAAAGGAAAGAAATCAAAGGTCACTGGAGAAGAAGAGACTGATCACATTGACGGAGAACTTGTTCTTTCCGTTGAGAAGTTGCAGGAGC
TCCAAGATGAGCTTGAGAAGATTAATGAACAAGCAAGCGATGAAGTCCTGGGAATTGAGCAGAAGTACAACGATGTTCGCAGACCTGTCAACACTAAAAG
GACCGATATCATCAAATCTATTCCTGATTTCTGGTTAACTGCAGAGAGTTTCCTGAGTCATCCTGCTCTTTGTGATCTTTTGACCGAGGAGGACCAAAAG
ATCTTCAAGTATCTAGATTCTCTAGACATGGAGGATTCCAAAGATGTGATGTCAGGGTATTCTATCACATTCAATTTCAAAGAGAATGCTCATTTTGAAG
ATACAAAGCTGAAAAAAACCTTTACATTCTTTGAGGAGGGTACAGCTAAGATAACTAGTACATACATCAAGCGCTGCAAATGGAGTCAATCACGAGAAAA
AGGAGACAAACGACCTTTGGCTGAGTTGAGCTTCTTCAGTTGGTTTGCTGAAACTGAACAGGAGGAGATCTCTGAGCTTCATGATGAGGTTGCGGAGATA
ATTAAGGAAGACTTGTGGCCCAATCCTCTAAAGTACTTTAACAATGAAGCTGATGAAGAGGATTCTGATGGAAGATGGAGATGGAGATGA
AA sequence
>Potri.004G128701.1 pacid=42793965 polypeptide=Potri.004G128701.1.p locus=Potri.004G128701 ID=Potri.004G128701.1.v4.1 annot-version=v4.1
MVADKGKKSKVTGEEETDHIDGELVLSVEKLQELQDELEKINEQASDEVLGIEQKYNDVRRPVNTKRTDIIKSIPDFWLTAESFLSHPALCDLLTEEDQK
IFKYLDSLDMEDSKDVMSGYSITFNFKENAHFEDTKLKKTFTFFEEGTAKITSTYIKRCKWSQSREKGDKRPLAELSFFSWFAETEQEEISELHDEVAEI
IKEDLWPNPLKYFNNEADEEDSDGRWRWR
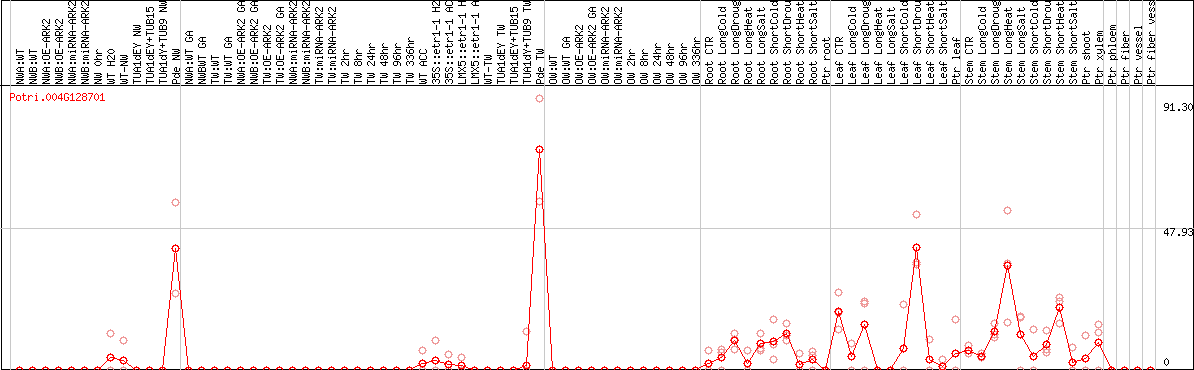
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G128701 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.