DMT901,Pt-MET1.2 (Potri.004G134000) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | DMT901,Pt-MET1.2 | ||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G134000.2 pacid=42794457 polypeptide=Potri.004G134000.2.p locus=Potri.004G134000 ID=Potri.004G134000.2.v4.1 annot-version=v4.1
ATGGGATCTTCATCTATATTGGATATAACAACAAATGACCCAGTTGATAATTCTACTTTTTCCGCATCATCAGGAATGAGGAAGAACAAGAAAGGAAAAC
AAAAATCTTCAGTTTCCAATGCAAAGAAGGAAGTGCCAGAAAAAACCGCAAAAGGAAAGAAGAGAAACTTCTCTGATACTAACAAGGAAGATCCTGCTGG
TGGGTCGTTGACAAGGCCTAGGAGAGCTGCTGCATGCAAAGACTTCAAGGAGAAATCTCTACGTTTGCATGAAGAGAAGTCTTCTGTTGTTGAATCAAAG
AAAGAACAAGTTGTCAACGAAGAGATTTTAGCTCTTCGTTTGACTCAAGGACAAGAAGAGGGTCGCCCCAACAGGAGATTGATTGATTTTGTTGTGCATG
ATGCAAATGGTAACCCTCAACCCTTGGAGATGATTGAAGTTGATGATATGTTTATCTCTGGTGTTATAATGCCTCTTGAGGAGAGCCTTGATAAGGAAAA
AGAAGTGCCTGTTAGGTGTGAAGGCTTTGGGAGGATAGAAGCATGGAATATCTCTGGTTACGAAGATGGATCCCCAGTCATTTGGCTGACTACTGAAGTA
GCAGATTATGATTGCATCAAGCCTTCCGGTGGCTACAAGAAGTTCTTTGATCGTTTCTTTCAGAAGGCACTCGCTTGCATAGAAGTTTACAAAAAACTCT
CAAGATTTTCTGGAGGAAACCCTGAATTCACCCTTGATGAGTTGCTTGCTGGCGTTGTGCGAGCAATGAGCGGAAATAAGTGTTTCTCTGGTGCTCCTTC
CGTAAAGAATTTCCTCGTTTCTCAGGGTGAATTTATTTATCAACAAATAACTGGTTTAGACCAGACCTCCAAAAAGAACGATAAATTTTTTTCAGACTTG
CCTGCTCTTGTTGCCTTGAGAGATGAAAGTCGCAATCATGGAAGTGTTCTGCTAGCAAAAGCAGCAAATCCTGGTGGCAACTTGGTGATTGATCCAAAGT
CTGTGGATGGTGCTATTGTGAATCAGTCTAACCAGTCCAGCACCATAGCTGAGGAAGATGAAGATGCAAAGTTAGCAAGATTATTACAGGAAGAAGAGTA
TTGGCACTCTAATATGAGGCAAAAGAAAAGTCGGGGTTCAGCTTCTGCATCGAACACAATCTACATCAAAATTAATGAAGATGAGATTGCAAATGATTAC
CCTCTCCCTGTGTTTTACAAACACTCTGATGAAGAAACCGATGAGTATGTAGTTGTTGCAAGTGATGATGTAATTGACCACCCTGATGATCTCCCAAGAA
AAATGCTTCATAACTGGTCACTTTATAACTCAGACTCAAGGTTGATTTCTTTGGAGCTTCTTCCAATGAAACCTTGTGAAGATATTGATGTCACCATCTT
TGGATCAGGACGCATGACTGAAGATGATGGAAGCGGGTTCTGTCTTGATGATGATCCTGATCAGTCTTCTTCTAGGGGTTCGGAGGCTCAGGATGACATG
GGTTTGCCAATATTTTTGAGTGCGATAAAGGAGTGGATGATTGAGTTTGGATCATCAATGATTTTTATATCAATACGCACAGACATGGCCTGGTATAGAC
TTGGGAAGCCATCAAAACAATATGGTTCCTGGTACAAGCCAGTCCTAAAAACGGTAAAGCTTGCAAGAAGCATCATTACATTGTTGAAGGAGCAAAGTCG
AGTATCACGGCTTTCTTTCGCAGATGTAATCCGGAAAGTGTCTGAGTTCAAGAAGGATCATCATGCTTATATTTCATCTGATCCAGCAGCTATTGAGAGG
TACGTAGTTGTTCATGGACAGATAATACTGCAACTTTTTGCCGAGTTTCCAGATCAGAAGATCAAGAAATGTGCTTTTGTGGTCGGTTTGACTCGTAAAA
TGGAGGAGAGGCACCATACTAAATGGGTAGTGAACAAAAAAGCCATTGTGCAAAAGTTTCAATCCAATTTGAATCCTCGAGCAGCAATGGATACTGTGGC
TCCTGGATCCAAGAGGAAGCTCATGCAGGCAACAACCACAAGGCTGATCAACAGAATTTGGGGGGAGTACTATTCGAACTATTCTCCAGAGGATTTGGAA
GAGGGAGCTGAATGTGAAGTGAAAGAAGAGGATGAAGCTGAGGAGCAGTATGAAAATGAAGATGATGATAAAGAGGAGGTGGTAGAGAAAACACTAAAAC
CTCGTTCAGTGTCTGAACGCACTAAATCACATACTTCACAAAAAGAAGTAAGATGGGATGGGAATCCTGTAAGCAAAACATCTTCTGGAGAAGCCATTTA
CAAGCGAGCCATTGTTTGTGGAGAGGTCATTGTGGTGGGGGATGCTGTCTTAGTGGAGGTTGATGAATCGGATGAACTTCCTGCCATCTATTTTGTGGAG
TACATGTTTGAAACACGGAATGGAAGCAGAATGTTTCATGGGAGAATGATGAAGCGAGGATCTGAGACAGTTCTTGGTAACACTGCCAATGATAGAGAGG
TTTTCTTAACAACCGAGTGCATGAATTATAAGTTACAGGATGCTAAGCAAGCAATTATTCTGGAAGTTCTAAAAAGACCTTGGGGACATGATCACAGGAA
AGATAACATCAATGCTGATAGAATTGATAGGGAAAAGGCAGAAGAGAGGAAAAAGAAAGGATTGCAAGTAGAATATTACTGTAAAAGCTTGTACTGGCCG
GAAAGGGGTGCTTTCTTTACTCTCCCGCTTGATACTATGGGTCTTGGGTCTGGTGTCTGCCACTCTTGTAACTTAAAAATAGCTGAAGAAGACAAGGATA
TTTTCAGAGTAAATTCTTCTCAGACAGGATTTTCATACAAGGGAACTGAGTACTCTGTCCACGATTTTGTCTACGTAAGCCCTCATCAATTTGCTTCAGA
AAGGGGGGAGAACGAAACTTTTAAGGGTGGAAGGAATGTTGGGTTGAAGCCTTATGTTGTGTGCCAGTTGTTGGAGGTTGTTCTAAAGGAACCAAAACAA
GCCGAAACAAGATCCACCCAGGTCAATGTTCAAAGATTTTTCAGACCAGATGATATTTCACCTGAGAAGGCATATTGTTCTGATATTAGGGAGATATATT
ACAGTGAAGAAACACATCTTTTATCAGTCGAGACAATTGAAGGGAAATGTGAAGTCAGAAAGAAAAATGACATTCCAACATGCAGTGCTCCTGCAATCTT
TGATAATATCTTCTTCTGCGAGCACATGTATGATCCTTCTAAAGGGTCCCTCAAGCAGCTACCAGCTCAAGTCAAATCGAAGTTTTCTGCAGTGAGTAGA
GATGGTGATGTTGCAAGTAGAAAGAGAAAGGGAAAGTCTAAAGAAGGAGAAAATGATATAGAAGCTGACAAACAAAGGGAAGCATCTCCAGAGAACCGTT
TGGCCACGTTGGATATATTTGCCGGTTGTGGTGGATTGTCTGAAGGATTGCAGCAAGCTGGTGTCTCATCAACCAAGTGGGCTATTGAATATGAAGAGCC
TGCTGGGGAAGCATTTAAACTCAACCATGCAGGGTCATTGATGTTCATTAATAATTGTAATGTGATCCTGAGGGCTGTCATGGAGAAATGTGGAGATGCA
GATGACTGTATTTCTACTTCTGAGGCTGGTGAATTGGCTTCATCACTTGATGCAAAGGTGATTGATGGTTTGCCATTGCCAGGGCAGGTGGATTTCATCA
ATGGAGGGCCTCCATGCCAGGGCTTCTCTGGAATGAATAGGTTTAATCAGAGCACTTGGAGTAAAGTTCAATGTGAGATGATCTTAGCATTCTTATCCTT
CGCTGATTATTTCCGCCCAAAGTATTTCCTCTTAGAGAATGTGAGGAACTTCGTATCTTTCAATAAAGGACAGACTTTCCGTTTAACTATCGCTTCTCTT
CTTCAGATGGGTTATCAGGTGAGGTTTGGTATTTTGGAAGCTGGAGCATATGGAGTATCCCAGTCACGTAAACGGGCCTTTATATGGGCAGCCTCCCCCG
AAGAGATACTCCCAGAATGGCCAGAGCCAATGCATGTTTTTGCTGCCCCAGAGTTAAAAATCACATTGTCTGAGAAATCACAATATTCTGCTGTTAGAAG
TACTGCTTATGGAGCCCCTTTCCGTGCAATAACAGTCAGAGATACAATTGGTGATCTCCCTGATGTAGGGAATGGTGCTTCGAAGACTAATTTGGAGTAC
GGAAATGATCCTGTCTCTTGGTTTCAAAAGAAGATTCGAGGCGACATGGTTGTCTTGACTGACCATATATCAAAAGAAATGAATGAGCTGAATCTTATCA
GATGCAAGAAGATCCCAAAGCGGCCAGGTGCTGACTGGCGTGATCTTCCAGATGAAAAGGTCAAATTGTCTACTGGACAAATGGTTGATTTGATACCATG
GTGCTTGCCAAACACAGCTAAACGACACAATCAGTGGAAGGGGCTGTTTGGAAGGTTGGATTGGGAAGGGAACTTCCCAACATCAATAACAGATCCTCAG
CCAATGGGTAAGGTGGGGATGTGCTTTCACCCTGAACAGGACAGGATTCTTACTGTTCGTGAATGTGCACGATCTCAAGGATTCCCAGATAGCTATCAGT
TTTCTGGTAACATTCATCACAAGCATCGGCAAATCGGAAATGCTGTCCCTCCTCCTTTATCATATGCACTGGGAAGGAAACTTAAGGAAGCACTGGATAG
TAAGAGGCGGAAATAG
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G134000.2 pacid=42794457 polypeptide=Potri.004G134000.2.p locus=Potri.004G134000 ID=Potri.004G134000.2.v4.1 annot-version=v4.1
MGSSSILDITTNDPVDNSTFSASSGMRKNKKGKQKSSVSNAKKEVPEKTAKGKKRNFSDTNKEDPAGGSLTRPRRAAACKDFKEKSLRLHEEKSSVVESK
KEQVVNEEILALRLTQGQEEGRPNRRLIDFVVHDANGNPQPLEMIEVDDMFISGVIMPLEESLDKEKEVPVRCEGFGRIEAWNISGYEDGSPVIWLTTEV
ADYDCIKPSGGYKKFFDRFFQKALACIEVYKKLSRFSGGNPEFTLDELLAGVVRAMSGNKCFSGAPSVKNFLVSQGEFIYQQITGLDQTSKKNDKFFSDL
PALVALRDESRNHGSVLLAKAANPGGNLVIDPKSVDGAIVNQSNQSSTIAEEDEDAKLARLLQEEEYWHSNMRQKKSRGSASASNTIYIKINEDEIANDY
PLPVFYKHSDEETDEYVVVASDDVIDHPDDLPRKMLHNWSLYNSDSRLISLELLPMKPCEDIDVTIFGSGRMTEDDGSGFCLDDDPDQSSSRGSEAQDDM
GLPIFLSAIKEWMIEFGSSMIFISIRTDMAWYRLGKPSKQYGSWYKPVLKTVKLARSIITLLKEQSRVSRLSFADVIRKVSEFKKDHHAYISSDPAAIER
YVVVHGQIILQLFAEFPDQKIKKCAFVVGLTRKMEERHHTKWVVNKKAIVQKFQSNLNPRAAMDTVAPGSKRKLMQATTTRLINRIWGEYYSNYSPEDLE
EGAECEVKEEDEAEEQYENEDDDKEEVVEKTLKPRSVSERTKSHTSQKEVRWDGNPVSKTSSGEAIYKRAIVCGEVIVVGDAVLVEVDESDELPAIYFVE
YMFETRNGSRMFHGRMMKRGSETVLGNTANDREVFLTTECMNYKLQDAKQAIILEVLKRPWGHDHRKDNINADRIDREKAEERKKKGLQVEYYCKSLYWP
ERGAFFTLPLDTMGLGSGVCHSCNLKIAEEDKDIFRVNSSQTGFSYKGTEYSVHDFVYVSPHQFASERGENETFKGGRNVGLKPYVVCQLLEVVLKEPKQ
AETRSTQVNVQRFFRPDDISPEKAYCSDIREIYYSEETHLLSVETIEGKCEVRKKNDIPTCSAPAIFDNIFFCEHMYDPSKGSLKQLPAQVKSKFSAVSR
DGDVASRKRKGKSKEGENDIEADKQREASPENRLATLDIFAGCGGLSEGLQQAGVSSTKWAIEYEEPAGEAFKLNHAGSLMFINNCNVILRAVMEKCGDA
DDCISTSEAGELASSLDAKVIDGLPLPGQVDFINGGPPCQGFSGMNRFNQSTWSKVQCEMILAFLSFADYFRPKYFLLENVRNFVSFNKGQTFRLTIASL
LQMGYQVRFGILEAGAYGVSQSRKRAFIWAASPEEILPEWPEPMHVFAAPELKITLSEKSQYSAVRSTAYGAPFRAITVRDTIGDLPDVGNGASKTNLEY
GNDPVSWFQKKIRGDMVVLTDHISKEMNELNLIRCKKIPKRPGADWRDLPDEKVKLSTGQMVDLIPWCLPNTAKRHNQWKGLFGRLDWEGNFPTSITDPQ
PMGKVGMCFHPEQDRILTVRECARSQGFPDSYQFSGNIHHKHRQIGNAVPPPLSYALGRKLKEALDSKRRK
|
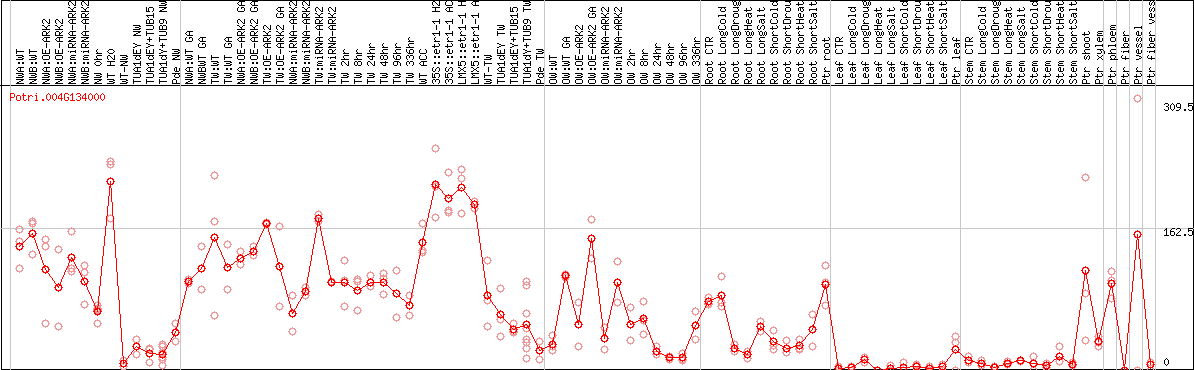
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G134000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
