Potri.004G135500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G135500.1 pacid=42795444 polypeptide=Potri.004G135500.1.p locus=Potri.004G135500 ID=Potri.004G135500.1.v4.1 annot-version=v4.1
ATGGGGGGTGCTTGTAATTTGGCAAGTAGTCCACAACTGCTCCTGCAAGTATCCATCATTTGTTCCATCACTCTCATTTCCTTTGGACTTGCAGCTTCTG
CTTCAGCTAAGCTCCACTCCCAAGAAGTTAGGGTATTAAGAGAAATAGGGAAGAAGCTGGGGAAGAAGGACTGGGATTTCAACAAGGATCCATGTAGTGG
AGAAGGGAATTGGAGTATTTTGGATGAGAGGAAAGGGTTCGAGAACAGTGTCACTTGTGACTGCTCTTTCAACAACAATTCCTCTTGCCACCTAGTAAGC
ATAGCTTTAAAGTCTCAGAATCTATCGGGAATCATTCCACCGGAGTTTTCCAAGTTTCGCTACCTTAAACAACTGGATCTTAGCCGCAATCTCTTCACTG
GTGTTATACCTCCCCAATGGGGTACCTTGCGCTTGGAGGAGTTCTCTGTTATGGGGAATCGGTTGTCTGGGCCATTCCCGAAAGTCCTCACCAACATGAC
CACCCTCAGAAACCTGAGTATTGAAGGAAATCATTTTTCAGGACCCATTCCACCTGAAATTGGAAGACTGATCAATTTACAGAAACTTGTTTTCTCCTCA
AATGCATTGACAGGAAACTTGCCTGCAGAACTTGGCAAGTTGGTCAACTTGACAGATGTGAGGATAAATGATAATAATTTCTCAGGAAAGCTACCTACTT
TCATCAGTAAATGGACAAAGGTTCAGAAACTGCATTTACAGGGTACCTCTCTTAAGGGGCCCATACCTTCTAGCATTGCTTCCTTGACCAAACTAAGTGA
TTTGAGGATTAGTGACTTAACAGGAAGAGGGTCTCCCTTCCCACCACTGAGTGATATGGAATCCATGAAAACATTGATATTGAGAAACTGCTTAATATAT
GGAGAGATCCCAGAATACGTTGGGCAGATGGAAAAACTGAAGCACTTAGATGTCAGCTTCAATAACCTCAGAGGAGAAATTCCTTCGACGTTTATCCAAC
TGGCAAGAATAGATTTCCTGTACTTGACAGGAAACAAGCTCACTGGGTCTGTTCCTCCATGGCTTCTGGAAAGAAACAAGAATGTGGATCTTTCTTATAA
CAACTTCACCTGGCAAAGCTCAAGTCCAGATGAATGTGCACGAGGGAGCGTGAACATAGTGGAGAGCTTCTCGCCGTCAACAATTAAATCAAGTAAAGCC
CATTCATGCCTGAAACAAAACTTTCCGTGTTCTGCTTCCCGAAATCAACAGCACTATACCTTGCATATTAACTGTGGAGGTAATGAGATAACTGTCGATG
GCAACACCACTTACCAAGATGATAAAGAACCAAGGGGTGCTTCTATGTTTTACTCCCATCCAAGCCAAGAATGGGCGTTTAGTAGCACCGGCAACTTCAT
GGACGATGACAGTGAGGCAGACGCCTATACTAAAACCAACAAGTCTGCTATATCCAATGTCTCTGCTACCATTGCACAACTATACACAACAGCGCGTGTT
TCTCCCCTTTCTCTCACTTACTACGGACTTTGCCTCATGAATGGGAATTACACAGTAAAACTTCACTTTGCTGAGATTATTTTCACAAATGACAGTTCAT
TAACTAGTCTTGGGAAGCGTATATTTGATGTTTACATCCAGGGAAAGTTGGTCCTGAAAGATTTTAATATTGAAGATGAGGCTGGAGGTGTTGCCATACC
CCTTGTAAAGACATTCATTGCAGCAGTAACACATAATACATTAAAGATCCGCTTGTATTGGGCTGGAAGAGGGACAACAGGCATACCACTTAGAGGAATT
TATGGTCCTCTAATATCAGCTATATCGGTAGACCCGAACTTTAAACCACCTTCAAATGGTAGCAAAAGGAATGTAGTAATCATAGTGACTGGAGCAGTAG
CTGGAGCAATTTTCCTTGCCTTTCTGGTACTAGGTGTCATGTGGAGAAATGGCTGGCTTTGTGGCAAAGCCGCTGCAGATAAAGAGCTTAAAGGCCTAGA
TCTGCAAACAGGACTATTCACGCTAAGACAGATGAAAGCTGCCACCAATAACTTCGATGCGGAAAACAAAGTTGGGGAGGGTGGTTTTGGTTCTGTTTAC
AAGGGTTCACTATCAGATGGAACCGTGATTGCAGTCAAGCTACTCTCCTCAAAATCCAAGCAAGGAAATCGTGAATTTGTGAATGAGATAGGAATGATAT
CTGCACTGCAACATCCGAATCTTGTAAAGTTGTATGGATGTTGTGTTGAAGGAAACCAATTGATGATTGTATATGAGTACATGGAAAATAATTGCTTATC
CCGTGCTCTTCTCGGGAAGGAATCCAAGTTCAGAATGAAACTGGACTGGCCTACCAGGCAGAAAATTTGCCTAGGTGTAGCTAAAGGTTTGATGTACCTA
CATGAAGAGTCAATAATAAAAATTGTGCATAGGGATATCAAAACTAGCAATGTGTTGCTTGATAAGGAACTCAATGCCAAGATTTCTGATTTTGGTTTGG
CAAAGCTAAATGAAGATGACGATACCCACATCAGCACTCGCATTGCTGGAACCATTGGTTATATGGCTCCTGAGTACGCAATGCGTGGCTACTTGACCAA
CAAAGCAGATGTTTATAGCTTTGGAGTTGTTGCTTTGGAAATTGTCAGTGGAAAGAGCAACACAAACTACAGGCCAAAGGAGGAGTTTGTTTACCTTCTT
GATTGGGCCTATGTTTTGCAAGAGAGAGGAAGTCTTTTGGAGTTGGTCGATCCAGAATTGGGGTCAGAGTATTCTTCAGAAGAGGCTATGGTGATGCTAA
ATGTCGCTCTCTTATGCACCAATGCATCCCCAACTCTTAGACCGACAATGTCTCAAGTTGTGAGCATGCTCGAAGGCCGAACTCCAGTTCAGGACCTTCT
TTCTGATCCTGGGTTTTCAGCAATCAACACAAAATACAAGGCTATAAGAAACCACTTCTGGCAGAATCCAAGCCAAACCTATAGCATGTCTATAAATGAG
TCGTACCGTACTGATTCCACAAGCTCAGGCGTAGAACCAGAAGATGCCGGCCGTCTTCTGAGAGTTAGTTCGGTAAAATCTAATTCGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G135500.1 pacid=42795444 polypeptide=Potri.004G135500.1.p locus=Potri.004G135500 ID=Potri.004G135500.1.v4.1 annot-version=v4.1
MGGACNLASSPQLLLQVSIICSITLISFGLAASASAKLHSQEVRVLREIGKKLGKKDWDFNKDPCSGEGNWSILDERKGFENSVTCDCSFNNNSSCHLVS
IALKSQNLSGIIPPEFSKFRYLKQLDLSRNLFTGVIPPQWGTLRLEEFSVMGNRLSGPFPKVLTNMTTLRNLSIEGNHFSGPIPPEIGRLINLQKLVFSS
NALTGNLPAELGKLVNLTDVRINDNNFSGKLPTFISKWTKVQKLHLQGTSLKGPIPSSIASLTKLSDLRISDLTGRGSPFPPLSDMESMKTLILRNCLIY
GEIPEYVGQMEKLKHLDVSFNNLRGEIPSTFIQLARIDFLYLTGNKLTGSVPPWLLERNKNVDLSYNNFTWQSSSPDECARGSVNIVESFSPSTIKSSKA
HSCLKQNFPCSASRNQQHYTLHINCGGNEITVDGNTTYQDDKEPRGASMFYSHPSQEWAFSSTGNFMDDDSEADAYTKTNKSAISNVSATIAQLYTTARV
SPLSLTYYGLCLMNGNYTVKLHFAEIIFTNDSSLTSLGKRIFDVYIQGKLVLKDFNIEDEAGGVAIPLVKTFIAAVTHNTLKIRLYWAGRGTTGIPLRGI
YGPLISAISVDPNFKPPSNGSKRNVVIIVTGAVAGAIFLAFLVLGVMWRNGWLCGKAAADKELKGLDLQTGLFTLRQMKAATNNFDAENKVGEGGFGSVY
KGSLSDGTVIAVKLLSSKSKQGNREFVNEIGMISALQHPNLVKLYGCCVEGNQLMIVYEYMENNCLSRALLGKESKFRMKLDWPTRQKICLGVAKGLMYL
HEESIIKIVHRDIKTSNVLLDKELNAKISDFGLAKLNEDDDTHISTRIAGTIGYMAPEYAMRGYLTNKADVYSFGVVALEIVSGKSNTNYRPKEEFVYLL
DWAYVLQERGSLLELVDPELGSEYSSEEAMVMLNVALLCTNASPTLRPTMSQVVSMLEGRTPVQDLLSDPGFSAINTKYKAIRNHFWQNPSQTYSMSINE
SYRTDSTSSGVEPEDAGRLLRVSSVKSNS
|
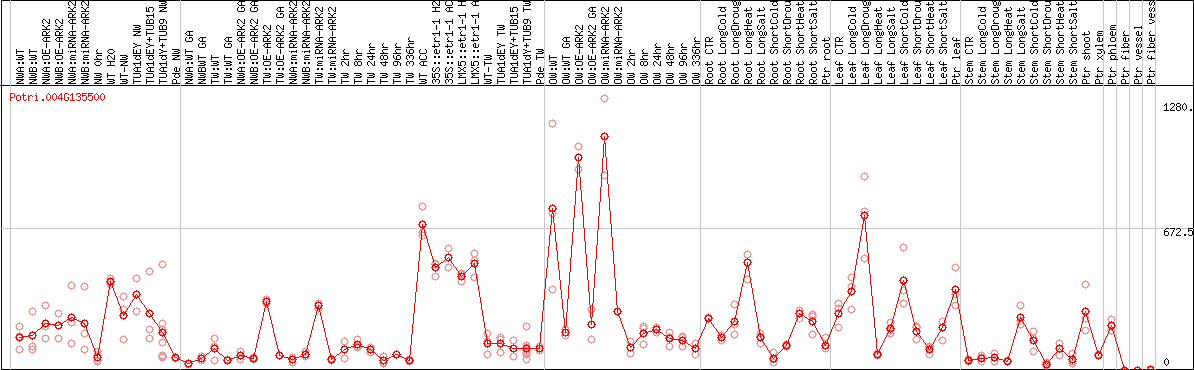
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G135500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
