External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
ATCG00900 250 / 1e-85
ATCG00900.1, RPS7.1
CHLOROPLAST RIBOSOMAL PROTEIN S7, Ribosomal protein S7p/S5e family protein (.1)
ATCG01240 250 / 1e-85
ATCG01240.1, RPS7.2
ribosomal protein S7 (.1)
ATMG01270 76 / 2e-17
ATMG01270.1, RPS7
mitochondrial ribosomal protein S7 (.1)
AT2G07696 76 / 2e-17
Ribosomal protein S7p/S5e family protein (.1)
AT2G37270 44 / 3e-05
ATRPS5B
ribosomal protein 5B (.1.2)
AT3G11940 42 / 7e-05
AML1, ATRPS5A
ARABIDOPSIS MINUTE-LIKE 1, ribosomal protein 5A (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G138900
248 / 1e-84
ATCG00900 310 / 2e-110
CHLOROPLAST RIBOSOMAL PROTEIN S7, Ribosomal protein S7p/S5e family protein (.1)
Potri.011G084201
149 / 2e-46
ATCG00900 186 / 2e-62
CHLOROPLAST RIBOSOMAL PROTEIN S7, Ribosomal protein S7p/S5e family protein (.1)
Potri.009G100700
115 / 3e-32
AT2G15570 193 / 2e-63
THIOREDOXIN-M3, GFP ARRESTED TRAFFICKING 1, Arabidopsis thioredoxin M-type 3, Thioredoxin superfamily protein (.1.2)
Potri.007G061841
77 / 7e-18
ATMG01270 291 / 4e-103
mitochondrial ribosomal protein S7 (.1)
Potri.011G075200
65 / 3e-14
ATCG00900 81 / 7e-22
CHLOROPLAST RIBOSOMAL PROTEIN S7, Ribosomal protein S7p/S5e family protein (.1)
Potri.016G063200
45 / 9e-06
AT2G37270 359 / 4e-128
ribosomal protein 5B (.1.2)
Potri.006G197700
44 / 3e-05
AT3G11940 358 / 2e-127
ARABIDOPSIS MINUTE-LIKE 1, ribosomal protein 5A (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002394
239 / 4e-80
ATCG00900 229 / 3e-77
CHLOROPLAST RIBOSOMAL PROTEIN S7, Ribosomal protein S7p/S5e family protein (.1)
Lus10027894
178 / 4e-56
ATCG00900 167 / 7e-53
CHLOROPLAST RIBOSOMAL PROTEIN S7, Ribosomal protein S7p/S5e family protein (.1)
Lus10008436
49 / 1e-06
AT3G11940 371 / 3e-130
ARABIDOPSIS MINUTE-LIKE 1, ribosomal protein 5A (.1.2)
Lus10013381
47 / 3e-06
AT2G37270 370 / 1e-132
ribosomal protein 5B (.1.2)
Lus10000168
44 / 4e-06
ATCG00900 46 / 5e-08
CHLOROPLAST RIBOSOMAL PROTEIN S7, Ribosomal protein S7p/S5e family protein (.1)
Lus10002395
44 / 4e-06
ATCG00900 46 / 5e-08
CHLOROPLAST RIBOSOMAL PROTEIN S7, Ribosomal protein S7p/S5e family protein (.1)
Lus10000946
45 / 2e-05
AT2G37270 374 / 8e-134
ribosomal protein 5B (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00177
Ribosomal_S7
Ribosomal protein S7p/S5e
Representative CDS sequence
>Potri.004G140500.1 pacid=42793898 polypeptide=Potri.004G140500.1.p locus=Potri.004G140500 ID=Potri.004G140500.1.v4.1 annot-version=v4.1
ATGGCTTCAACTGCTACTTCATTCTGCTCTCCACCGCTCACGAGCTCCCGCGCTTCCGTTCTTCGCCAATGCCAGCAGCTGAACCCTAATAGACTCTCAT
TTCCAATCAACTTGAACACCGCTAAAAGAGCACCGAATCTCACTGTTCAACATGCGCCTCTCCCTCTCAAGGTTCTCTGTGCTCGCGGAAAACGAGGTAC
AACGGCAAAAAAACCTGCCACTTCTGATCCAGTTTATCGTAACCGGTTAGTTAACATGTTTGTCAACCGAATCTTGAAAAAGGGGAAAAAGTCACTGGCT
TATAATATCATCTATCAAGCATTGAAAAATATTCAGCAAAAGACACAATCAAATCCATTAGCTGTTTTACGTGAGGCAGTGAATGGAGTAACTCCTGATG
TAGCGGTAAAAGCAAGACGTGTTGGTGGATCTACCCAACAAGTTCCTATTGAAGTAGGAACATTCCAAGGAAAAGCACTTGCTATCCGTTGGTTGTTGGA
GGCATCCCGAAAACGTTCAGGACGAAGTATGGTTCTCAAATTGAGTTCTGAAGTTATGGATGCTGCCAAAGGGACCGGTGAAGCCATAAAAAAGAGGGAA
ACCACTCATAAGATGGCAGAGGCTAATAGAGCTTTTGTGCATTTTCGTTGA
AA sequence
>Potri.004G140500.1 pacid=42793898 polypeptide=Potri.004G140500.1.p locus=Potri.004G140500 ID=Potri.004G140500.1.v4.1 annot-version=v4.1
MASTATSFCSPPLTSSRASVLRQCQQLNPNRLSFPINLNTAKRAPNLTVQHAPLPLKVLCARGKRGTTAKKPATSDPVYRNRLVNMFVNRILKKGKKSLA
YNIIYQALKNIQQKTQSNPLAVLREAVNGVTPDVAVKARRVGGSTQQVPIEVGTFQGKALAIRWLLEASRKRSGRSMVLKLSSEVMDAAKGTGEAIKKRE
TTHKMAEANRAFVHFR
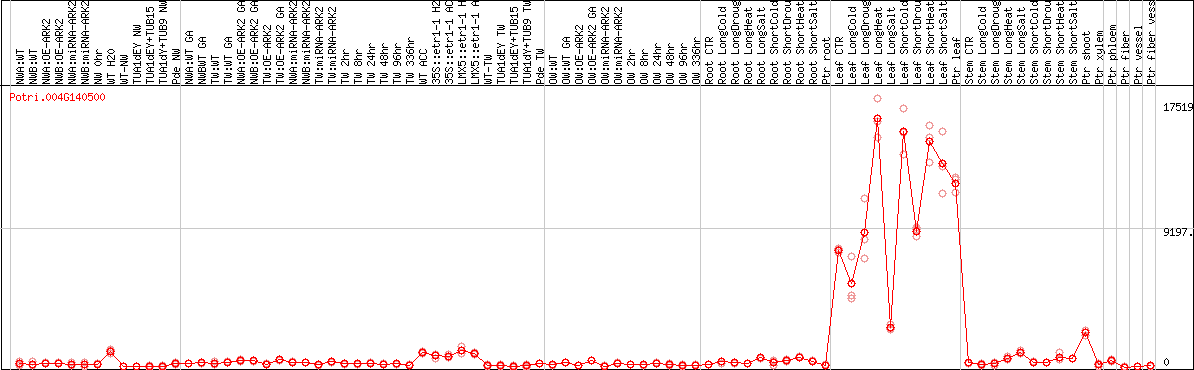
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G140500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.