External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G34360 385 / 7e-137
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT3G60910 116 / 3e-31
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT3G17365 110 / 6e-29
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT2G31740 73 / 2e-14
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT1G66680 55 / 2e-08
AR401
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G055600
123 / 1e-33
AT3G60910 378 / 1e-133
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.001G261000
121 / 3e-33
AT3G60910 387 / 2e-137
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.010G153200
100 / 3e-25
AT3G17365 313 / 1e-108
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.001G317700
81 / 3e-17
AT2G31740 886 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019750
115 / 9e-31
AT3G60910 394 / 3e-140
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10016376
100 / 9e-25
AT3G60910 327 / 2e-113
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10016375
73 / 3e-16
AT3G60910 164 / 7e-52
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10016656
78 / 6e-16
AT2G31740 895 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10017619
49 / 1e-06
AT1G66680 471 / 1e-167
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10033577
49 / 1e-06
AT1G66680 385 / 3e-132
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10037890
42 / 0.0002
AT5G54400 449 / 2e-160
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10038599
42 / 0.0003
AT5G54400 455 / 8e-163
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF13847
Methyltransf_31
Methyltransferase domain
Representative CDS sequence
>Potri.004G150200.2 pacid=42794527 polypeptide=Potri.004G150200.2.p locus=Potri.004G150200 ID=Potri.004G150200.2.v4.1 annot-version=v4.1
ATGCAAGAAGAAAGCGAGCCTAAGCTGAGAAACCCTGAAATGGAAGGGTCCCGAAAGCCGACCACCGAGCCTCCCTCTACTGCTCTGGCTTACCAGGATC
CTCACTACTGGAACGAGAGATTCTCTAAGGAGGAACATTACGAATGGTTCAAAGACTACTCTCATTTTCGCCATCTTATTCAAGCCCACATCCCTCCAAC
CTCCTCTGTGTTGGAGCTTGGTTGTGGGAATTCACAGTTGTGTGAAGAAATGTACAGAGATGGAATTACTGAGGTTACTTGCATTGATTTGTCCGCTGTT
GCGGTCGAGAAGATGCAGAAACGGTTGGAAGCTAAGGGGTATAAAGAAATAAAAGTGCTGGAAGCTGACATGCTAGACCTGCCCTTTAATGATGAATGTT
TTGATGTGGTTATTGAGAAAGGAACCATGGATGTATTATTCGTCAACAGTGGTGACCCATGGAATCCAAGGCCTGAAACAGTAGCCCAAGTGAAGGCAAT
GCTTGAAGGAGTTCACAGGGTTCTGAAACCAGATGGCATTTTTATCTCGATCTCTTTTGGCCAGCCACATTTTAGGCGTCCTTTATTTGATGCTCCGGAC
TTCACTTGGTCTGTAGAGTGGAGCACTTTTGGTGATGGATTTCACTACTTTTTCAATGTCTTGAGGAAGGGGAAGAGATCATCAAGTGACGAAGGGTCTA
GTGGAAAGAATGAAATACCATCCATTTGTCTATTTCAAGAAGAATTAGAGGGAGAAGATTTTATTTTTCGAACCAACATAGATGAAAATTCTTAG
AA sequence
>Potri.004G150200.2 pacid=42794527 polypeptide=Potri.004G150200.2.p locus=Potri.004G150200 ID=Potri.004G150200.2.v4.1 annot-version=v4.1
MQEESEPKLRNPEMEGSRKPTTEPPSTALAYQDPHYWNERFSKEEHYEWFKDYSHFRHLIQAHIPPTSSVLELGCGNSQLCEEMYRDGITEVTCIDLSAV
AVEKMQKRLEAKGYKEIKVLEADMLDLPFNDECFDVVIEKGTMDVLFVNSGDPWNPRPETVAQVKAMLEGVHRVLKPDGIFISISFGQPHFRRPLFDAPD
FTWSVEWSTFGDGFHYFFNVLRKGKRSSSDEGSSGKNEIPSICLFQEELEGEDFIFRTNIDENS
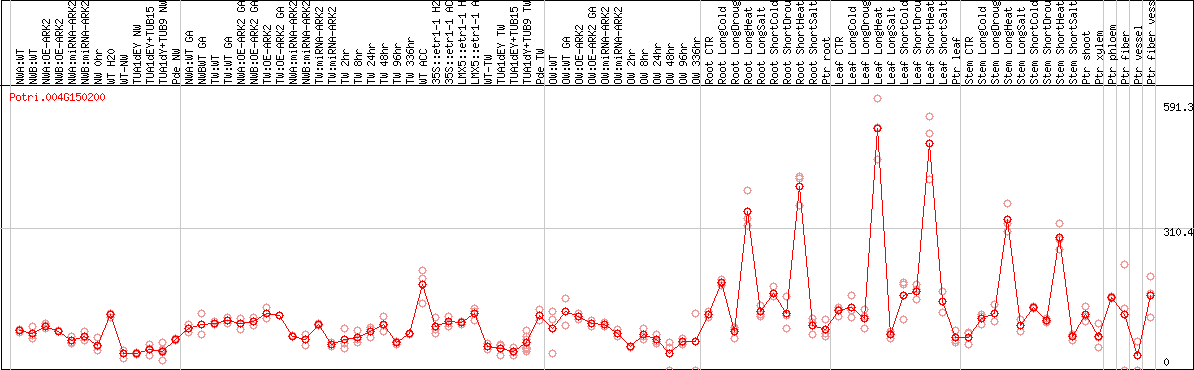
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G150200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.