ALPHA.13 (Potri.004G152400) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ALPHA.13 | |||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G152400.1 pacid=42793887 polypeptide=Potri.004G152400.1.p locus=Potri.004G152400 ID=Potri.004G152400.1.v4.1 annot-version=v4.1
ATGGAGTCGTTGATCGAGCTATGCGACCTAATTTCACAGAACCCTGCACAATTCGCTGACAAATTGACTTGGCTATGCAATCGCTGTCCTCAACCGGAGG
CTCTCTTAGCCGGATCGCCTAGGGTTTCTCACTCACAGATCAATGCAATCCTTGCGATTTCTCGATTCCTCTCCAAAACCCTCGACCACACCGATAACCG
TCCAAAATCACTAATCCTCACCTTCTTTCGCTCAATTCCGACCTCGTTTCACCCTTCCTTCTGGCCACAATCATTCCCCAACGACTCGATCGCGTCCTTT
TTTACCGATTTCCTGGCGTACGTCTCAAAGTCGGCCGAATTGGATCCGGATTTCGCTGTCGATGTTGCCGGACTTGTTGGAGAAGTTGTCGTGGCTGCGA
TCGGGAACAATGCCGGTGAGAATTGGGAGAGTTCGGCGATTTCTAGGGTTTTCTTGATTGCGTTAACGAAGAATTTTGTTCCAATTTTGCCGGAGGATGG
AGAGAAGTTGATTACGTGTCTTTTAGATCAGTTTAACCTGCCCGTACAGGTGCCGTCGTCACCTAGTGAGCGAATTGGGATGAATTCTGGAACATCTTCG
TCACAGAGTTCGCCATTGAGCAATAATGTGAATAGTCATAATAGTAGTTACAGTGCCCATAATGAGATTTCGAGTATGGTGAATGATTTGAGTCAAATGT
CAGTATCTTCGTCGAGTGCATCGACGACAGTTGTGGTGAATGGGAGTGGAGTCACGTGGAAGAGTGGATTGGAGACAATGGGGGTGGGACTTGATGGAGG
TGGGGTATTGAGTAGGCAACAAGTGGCTTCATTCGAGGAAGAGAGTGTGGAGGGATTGGAGAAGCAGGAGATCGCATACAAGTTGATTGGACTTGTTTTG
GATTGTGCTAGGATAGACAACAAGCTTTTGGATCAAGTTAGGTTAATTGCTAAGAAGCAGCTTCAATCTCTCTCTGCATTTTTGAAGATACGGAAACGGG
ACTGGAATGAACAGGGACAACTTTTAAAAGCAAGAGTCAGTGCAAAACTTTCTGTCTATCAAGCTGCAGCCAGGATGAAAGTTCAGAGTCTTGCATCTCT
TGATGTGGATGGCAAGACATCCAAGAGGTTGGTGCTTGAGACTCTGGCATTGTTGATGGATGCTGCAGAAGCCTGTTTGTTCTCAGTGTGGCGGAAGCTG
AGAGTTTGTGAAGAGCTGTTCAGCTCATTGCTTGGTGGGATTGCACAAATTGCGGTAACCAGGGGAGGTCAGCCAATGCGAGTCCTGCTTATTCGCCTTA
AGCCGCTTGTGCTGGCTGCATGTGCACAGGCTGATACCTGGGGTGGCAGTCAGGGAGTAATGTTTGAGATTGTCATGAAGACAAGTTGTCAGATAATTGA
ATCTGGCTGGACAAAGGACCGGGCTCCTGTAGACACTTTCATCTCTGGTTTGGCCTCCAGCATACGTGAACGAAATGATTATGATGAACAGGTTGAGAAG
AAGCAGGGAGTTCCTGCTGTGCAGCTTAATGTTATTCGCTTGCTTGCTGATTTGACTGTTTCCGTTAACAAGTCTGAAGTGGTAGACATGATTTTGCCAC
TTTTTATTGAAAGCTTGGAAGAGGGCGAAGCTTCAACTCCTGGTTTATTGCGACTTCGACTTCTTGATGCTGTTTCTCGAATTGCAAGTTTAGGTTTTGA
GAAGTCATATCGTGAGACAGTTGTTCTTATGACAAGAAGTTACTTAAGTAAACTATCAAGTGTAGGGTCTGCTGAAAGCAAAATATTGGCAGCAGAAGCC
ACCACAGAACGTGTTGAGACTCTTCCTGCAGGATTTCTTCTGATTGCTAGCAGGCTTGAAAATAAGAAATTGCGTTCAGACTATCGTCATCGATTGCTAT
CTCTGTGTTCAGATGTAGGCCTGGCTGCTGAATCTAAAAGTGGAAGGAGTGGAGCAGATTTTCTAGGACCTCTGCTTCTTGCTGTTGCTGAAATATGCTC
TGATTTCAATCCCGCTGTGGATGTGGAACCATCACTCTTGAAATTGTTCCGCAACCTGTGGTTCTATGTTGCTCTTTTTGGTCTAGCACCTCCAATTCAG
AAAATTCAACAACCAACAAAGTCAGTTTCAACTACACTGAACAGTGTGGGAAGCATGGGCACAATTGCTCTTCAAGCAGTGGGTGGACCATACATGTGGA
ATGCACAGTGGTCGTCTGCTGTTCAGCGAATTGCTCAAGGGACTCCTCCCCTAGTTGTTAGCTCAGTAAAATGGCTTGAAGATGAATTGGAACTCAATGC
CCTTCACAATCCTGGAAGTCGTCGGGCCAGTGGCAATGAGAAAGCAGCTTCAACCCAAAGATCTGCTCTTTCAGCTGCTCTAGGTGGCAGAGTTGATATT
GCAGCTATGAGCACAATTTCAGGTGTGAAGGCAACCTATCTTCTTGCTGTAGCATTTTTGGAGATAATACGCTTCAGCAGCAATGGCGGCATCCTTAATG
GTGTTGCTAGTTTGAGTGCTTCTAGAAGTTCCTTCAGCTGTGTCTTTGAGTATCTAAAAACTCCAAACCTCATACCGGCTGTGTTCCAATGTTTAACAGC
TATTGTTCATAGGGCGTTTGAAGCAGCAGTATTCTGGCTGGAAGATCGAATAACTGAAACAGGCAATGAAGCCAATGTTAGAGAATCAACTCTTTTCTCC
CATGCCTGTTTTCTTATAAAAAGCATGTCTCAGAGGGAGGAACACATTCGAGATATTTCTGTAAGCTTGCTGACTCAGCTTAGAGATAAGTTTCCTCAGG
TTTTGTGGAATTCATCTTGTTTAGACTCCCTGCTATTTTCAGTTCACAACGACTCACCTTCTACAGTTATTAATGATCCTGCTTTGATAGCGTCTATTCG
TTCTTTGTACCAAAGGATTGTGAGGGAATGGATCAGCATATCACTTTCATATGCTCCATGCACTAGCCAAGGTCTGCTTCAGGAAAAGCTTTGCAAGGCA
AATACGTGGCAGAGAACGCAACATACAACTGATGTTGTTTCTCTTTTAACTGAGATACAGATTGGGAATGGTAAAAATGATTGGACTGGCATACGAACAG
CTAACATTCCTGCAGTCATGGCTGCTGCAGCAGCTGCTTCAGGAGCAAACTTCAAATCGACGGAAGCTTTCAACTTGGAGGTGCTCAGTATTGGTATCGT
CAGTGCAACAGTTAAGTGCAACCATACTGGAGAAATTGCAGGCATGAGAAGGTTGTATAATAGCATTGGTGGATTTCAATCTGGTGGCACACCAACAGGT
TTTGGTGGTGGTCTTCAAAGGTTGATATCTGGAGCATTTTCTCAACAACCACCAGCCGAGGATGATGCATTTAATGAGATGCTACTTAATAAGTTTGTGC
ATCTTCTTCAGCAATTTGTCAGTATTGCAGAAAAAGGTGGGGAAGTGGACAAGTCACAGTTTCGTGATACATGTTCTCAGGCCACTGCATTTCTTCTCTC
AAATCTGGCTTCTGAGTCAAAATCAAACGTAGAAGGCTTTGCTCAACTTCTACGTCTTCTTTGCTGGTGTCCAGCGTATATTTCTACACCTGATTCGATG
GAAACTGGTGTTTTCATTTGGACTTGGTTGGTTTCTGCAGCTCCGCAACTCGGTTCTCTTGTGCTTGCAGAGCTAGTGGATGCCTGGTTATGGACAATTG
ATACGAAACGAGGAGTTTTTGCTCATGAAGTGAAGTATTCTGGACCAGCTGCAAAACTGAGACCTCAGCTAGCCCCCGGGGAACCTGAATCGCAGCCTGA
AATTGATCCTGTCGAACAAATAATGGCGCATAGAATATGGGTTGGATTCTTCATTGACCGTTTTGAGGTAGTTCGACACAATAGTGTTGAGCAACTCTTG
CTTCTTGGCCGGTTGTTGCAGGGGACAACCAAGTCTCCCTGGAACTTTTCATGCCATCCCGCAGCTACTGGCACTTTTTTCACCATCATGCTTCTTGGTC
TCAAGTTCTGTTCATGTCATTCCCAAGGCAATTTGCAGAATTTCAAAACAGGGCTTCAACTGCTGGAAGATCGTATTTATAGGGCCTGTTTGGGTTGGTT
TGCTTTTGAGCCTGAATGGTTTGATGCAAACAATGTGAATTTTGCCCATAGTGAGGCTCAATCTGTCTCCCTATTTGTTCACTATATTTCTAATGATGGT
CAGTCTGATGCAAGAGGAAGGGGGCATGAGAATGGGACTTATTCGGTTGATATGAATGATCAGTATCACCCTGTATGGGGCCAGATGGAGAACTATGCTG
CAGGTAGAGAAAAACGGAGGCAGCTTCTTTTAATGCTATGCCAGAATGAAGCTGATAGGCTTGAAGTTTGGGCACAACCTACTAACTCTAAGGAAAATAC
ATCTTGGCCAAAAATCAGCTCAGAGAAATGGATTGAATATGCTAGGACTGCCTTCTCTGTGGATCCAAGGATTGCCTTATGCTTGGTCTCGAGGTTCCCA
ACAAATACTAATCTGAAAGCTGAAGTAACTCAACTAGTTCAGTCACATATTTTGGACCTCCGCTGCATACCTGAAGCATTGCCATATTTTGTCACCCCAA
AGGCTGTTGATGAAGATTCAGTGCTTTTGCAACAACTGCCTCACTGGGCTGCTTGTTCAATCACACAAGCACTCGAGTTCCTGACTCCTGCATACAAGGG
TCATCCACGTGTGATGGCATATGTTCTTAGAGTTTTGGAGTCTTATCCTCCAGAACGAGTGACCTTCTTTATGCCTCAGCTAGTGCAGTCTTTGCGATAT
GATGACGGGAGGTTAGTAGAAGGATATTTACTCAGAGCAGCTCATAGAAGTGATGTATTTGCCCATATTCTGATATGGAATTTGCAGGGTGAAACGTTTA
CATCAGAATCAAAGGAAGCTTCTTCTGGAAAGAATGTTTCCTTTCAAGCAATGTTGCCAGTTGTTCGTCAGCATATTATTGATGGTTTCACTCCCAAGGC
CCTGGACTTGTTTCGAAGGGAATTTGATTTCTTTGATAAAGTTACATCTATATCTGGTGTTCTTTATCCACTTCCTAAAGAAGAACGCCGGGCTGGTATT
CAGAGGGAGTTGGAGAAAATTGAATTGGAGGGGGAAGACCTTTACCTACCAACTGCTCCTAACAAGCTTGTGAGGGGTATCCGGGTGGATAGTGGAATAC
CTTTACAATCTGCAGCAAAAGTTCCCATCATGGTGACATTTAATGTGGTTGATAGGTGTGGGGACCGGAATGATGTAAAACCCCAAGCTTGCATTTTCAA
GGTAGGAGATGATTGTCGACAGGATGTTCTTGCCCTTCAAGTGATAGCACTCCTTAGAGATATATTTGAAGCAGTTGGAGTTAATCTCTACTTGTTTCCT
TATGATGTCCTCCCAACTGGCCCGGAGAGAGGTATAGTTGAGGTTGTGCCTAAAACACGAAGCAGAAGTCAGATGGGTGAAACAACAGATGGCGGTTTGT
ATGAGATCTTTCAACAGGATTATGGGCCCGTCGGCTCTCCTAGTTTTGAAGCTGCACGCAAGAATTTTATCATTAGCAGTGCTGGTTATGCAGTTGCCAG
TCTTCTACTTCAGCCGAAGGATAGACACAATGGGAATCTTCTTTTTGACAACGTGGGGAGGCTTGTCCACATTGATTTTGGTTTTATCTTGGAAACATCA
CCTGGTGGAAACATGCGCTTTGAGAGTGCGCACTTCAAGCTGAGCCATGAGATGACTCAATTACTAGATCCCTCTGGTGTTATGAAGAGCGAAACTTGGT
TGCAATTTGTAAGCTTGTGTGTGAAGGGGTACCTGGCGGCCCGCCGCTACATGGATGGGATAATCAACACTGTGATGTTGATGCTTGACAGTGGATTACC
TTGCTTCAGTAGGGGTGACCCAATTGGAAATCTTCGCAGGAGATTTCACCCTGAGATGAGCGAGCGTGAAGCTGCCAATTTCATGATCCGTGTGTGCACA
GATGCCTACAATAAGTGGACCACTGCTGGCTATGACTTGATACAATATATTCAACAGGGCATTGAGAAGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G152400.1 pacid=42793887 polypeptide=Potri.004G152400.1.p locus=Potri.004G152400 ID=Potri.004G152400.1.v4.1 annot-version=v4.1
MESLIELCDLISQNPAQFADKLTWLCNRCPQPEALLAGSPRVSHSQINAILAISRFLSKTLDHTDNRPKSLILTFFRSIPTSFHPSFWPQSFPNDSIASF
FTDFLAYVSKSAELDPDFAVDVAGLVGEVVVAAIGNNAGENWESSAISRVFLIALTKNFVPILPEDGEKLITCLLDQFNLPVQVPSSPSERIGMNSGTSS
SQSSPLSNNVNSHNSSYSAHNEISSMVNDLSQMSVSSSSASTTVVVNGSGVTWKSGLETMGVGLDGGGVLSRQQVASFEEESVEGLEKQEIAYKLIGLVL
DCARIDNKLLDQVRLIAKKQLQSLSAFLKIRKRDWNEQGQLLKARVSAKLSVYQAAARMKVQSLASLDVDGKTSKRLVLETLALLMDAAEACLFSVWRKL
RVCEELFSSLLGGIAQIAVTRGGQPMRVLLIRLKPLVLAACAQADTWGGSQGVMFEIVMKTSCQIIESGWTKDRAPVDTFISGLASSIRERNDYDEQVEK
KQGVPAVQLNVIRLLADLTVSVNKSEVVDMILPLFIESLEEGEASTPGLLRLRLLDAVSRIASLGFEKSYRETVVLMTRSYLSKLSSVGSAESKILAAEA
TTERVETLPAGFLLIASRLENKKLRSDYRHRLLSLCSDVGLAAESKSGRSGADFLGPLLLAVAEICSDFNPAVDVEPSLLKLFRNLWFYVALFGLAPPIQ
KIQQPTKSVSTTLNSVGSMGTIALQAVGGPYMWNAQWSSAVQRIAQGTPPLVVSSVKWLEDELELNALHNPGSRRASGNEKAASTQRSALSAALGGRVDI
AAMSTISGVKATYLLAVAFLEIIRFSSNGGILNGVASLSASRSSFSCVFEYLKTPNLIPAVFQCLTAIVHRAFEAAVFWLEDRITETGNEANVRESTLFS
HACFLIKSMSQREEHIRDISVSLLTQLRDKFPQVLWNSSCLDSLLFSVHNDSPSTVINDPALIASIRSLYQRIVREWISISLSYAPCTSQGLLQEKLCKA
NTWQRTQHTTDVVSLLTEIQIGNGKNDWTGIRTANIPAVMAAAAAASGANFKSTEAFNLEVLSIGIVSATVKCNHTGEIAGMRRLYNSIGGFQSGGTPTG
FGGGLQRLISGAFSQQPPAEDDAFNEMLLNKFVHLLQQFVSIAEKGGEVDKSQFRDTCSQATAFLLSNLASESKSNVEGFAQLLRLLCWCPAYISTPDSM
ETGVFIWTWLVSAAPQLGSLVLAELVDAWLWTIDTKRGVFAHEVKYSGPAAKLRPQLAPGEPESQPEIDPVEQIMAHRIWVGFFIDRFEVVRHNSVEQLL
LLGRLLQGTTKSPWNFSCHPAATGTFFTIMLLGLKFCSCHSQGNLQNFKTGLQLLEDRIYRACLGWFAFEPEWFDANNVNFAHSEAQSVSLFVHYISNDG
QSDARGRGHENGTYSVDMNDQYHPVWGQMENYAAGREKRRQLLLMLCQNEADRLEVWAQPTNSKENTSWPKISSEKWIEYARTAFSVDPRIALCLVSRFP
TNTNLKAEVTQLVQSHILDLRCIPEALPYFVTPKAVDEDSVLLQQLPHWAACSITQALEFLTPAYKGHPRVMAYVLRVLESYPPERVTFFMPQLVQSLRY
DDGRLVEGYLLRAAHRSDVFAHILIWNLQGETFTSESKEASSGKNVSFQAMLPVVRQHIIDGFTPKALDLFRREFDFFDKVTSISGVLYPLPKEERRAGI
QRELEKIELEGEDLYLPTAPNKLVRGIRVDSGIPLQSAAKVPIMVTFNVVDRCGDRNDVKPQACIFKVGDDCRQDVLALQVIALLRDIFEAVGVNLYLFP
YDVLPTGPERGIVEVVPKTRSRSQMGETTDGGLYEIFQQDYGPVGSPSFEAARKNFIISSAGYAVASLLLQPKDRHNGNLLFDNVGRLVHIDFGFILETS
PGGNMRFESAHFKLSHEMTQLLDPSGVMKSETWLQFVSLCVKGYLAARRYMDGIINTVMLMLDSGLPCFSRGDPIGNLRRRFHPEMSEREAANFMIRVCT
DAYNKWTTAGYDLIQYIQQGIEK
|
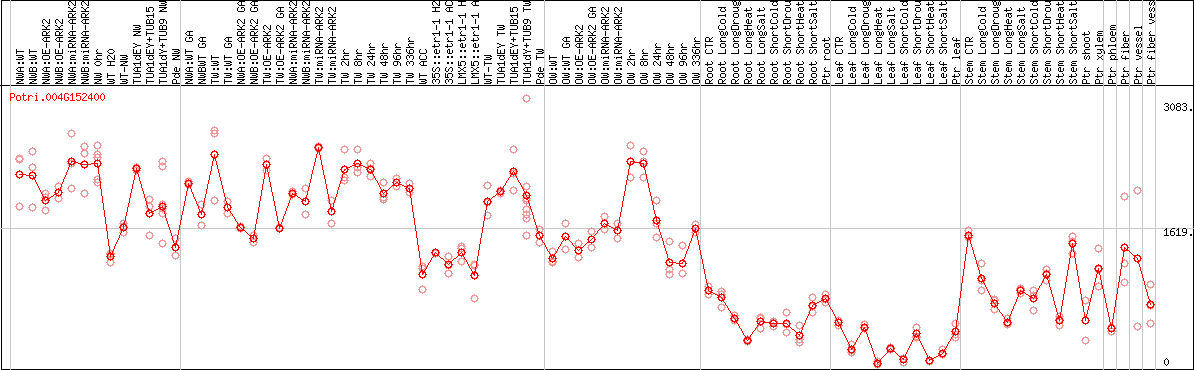
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G152400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
