Potri.004G176000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G176000.5 pacid=42795185 polypeptide=Potri.004G176000.5.p locus=Potri.004G176000 ID=Potri.004G176000.5.v4.1 annot-version=v4.1
ATGTGTTTTCAGCTAGAAAATATGCAAAATGAAAGTAGTTCCAAAGTCAACGGGTTTGTCGATAGTGGTGGTGATAAAAACAGCTTGGATCAAAGTGGTT
CTATGGCAAATGGGTTGGTTTGTGAGTTGAAAGTTGATAGTGAAAACAATGTTAAAAAACTGGATGAGGTTGTTGGGAACTTGAATAATGTGCCATATAT
TTATGGACAAGATATTGTTAGACATAAAATGCACAATGTGGTTGGGGTAGTGAGTGAAGTTGCTGGTGAGTCAGATTCGGATGGAAGTATTATGGATGAT
GATGATGATGATGATGAGGATGAGGACGACGACGACGACGATGACGAAGATGAGTGTGATGTCGATGAAGATGCTGGCAATGAGGGAGCTGATGATGATG
GTAATGATAATACCAATGGCGATAGGAGTGAAGGGAGCGATAATTATAAGAGTGAAACTCTTCAGACTGATGAGGTTCGAGTAGTTTGGATGGGTGATGT
TAAACCAGTTCAACATGTCAATGATGTAACTGTCATTGATCGAGGATTCTTACACGGGGACTATGTTGCATCTGCTTCAGATCCAACAGGGCAAGTAGGT
GTTGTGGTGGATGTTAATATCTCTGTTGATTTGTTAGCTCCTGATGGATCCGTTAGAAAAGACATTTCATCCATAAATTTGGTGCGTGTTAGGGAATTTT
CTGTTGGTGACTTTGTGGTCTTTGGTCCCTGGTTAGGCAGAGTTGATGATGTTGTAGATGATGTGACTGTGTTATTTGATGATGGTTCTGTGTGCAAAGT
AAAGGGGGCTGAGCCACTGCATCTAAAACCAATTTCCAAGGGTATCTTTGAGGAAGATGAACATTTTCCTTACCATCCTGGCCAGCGTGTAAGGGCTACC
TCATCATCAGTTTTTAAAAACTCTCGGTGGCTATCCGGCTTGTGGGAAGCTAATAGATTGGAAGGTACTGTTACTAAAGTTACATCTGGATCAGTTTTCA
TCTATTGGATAGCTTCTGCTGGCTATGGACCTGATTCTTCAACTACCCCTGCTGAAGAGCAGGATCCAAAGAACTTAGAACTGCTGTCTTGTTTTGCACA
TGCAAGTTGGCAAGTGGGTGACTGGTGTCTTCTCCCATCATCTGTTGCTCAATCATCTTCTGTTACTTTAGACAAGGATTTATTGAAGTTGGGAATCCAT
GACCCAGCCAAATGTGAATTAGATTCTAGTCAACTGGGAAATGGATGTGATTCAGAAGGAGTTGCAAATGAGGAAGTAGATGATATGAATGGATCGATGG
CAATTGATCCAGTGACAACACCGGATGGAAATTTAGCCATTATTGCAAGTAATGAATCTAGCTCTTGTGGTAGTTCAACATCAGTTTCAAAGGTGCCTGC
CCATCGCAGAAAGATACGTAAAGTTTTAATCAGAAGAGAAAAGAAGAGTATAAAAAAAGAGGAAGATTTTGAAAGAGCCCTTTTGATTGTCAACACCAGG
ACAAGGGTCGATATAGCATGGCAGGATGGAGCTATAGAACGTGGGCTGAATTCTACAACATTGATTCCAATTGATAGTCCTGGTGATCATGAATTCGTTG
CTGAACAGTATGTTGTGGAGAAGGCCTCTGATGATGTTGATAACTCATTTGAATCTAGACGTGTTGGTGTTGTTAAAAGCCTTAATGCAAAAGAGCGAAC
AGCTTGTGTGAAGTGGTTAAAGCCAGTTACCAGGGCAGAAGATCCTCGTGAGTTTGACAAGGATGAAATTGTAAGTGTGTATGAGCTGGAGACTCATCCA
GATTATGACTATTCCTATGGGGATGTAGTCGTTCGATTATCCCCGGTTTCTGTGTCTGATCAAACCACCTCTGATTTGGAAACTATTGGGGAATCAAGGC
AGCAAAGTGGACAAAGTGAGATTATGAATACACAAAAGTGCTTAGGACATAAGAAAGGAGAGGATGCACCGAGTAATGATGTTTCTATGGACTCCTCAGA
TCTCTCTTGGGTTGGGAATATATCTGGTCTTAGGAATGGTGATCTCGAAGTCACATGGGCTGATGGGATGGTTTCAATGGTTGGACCTCAAGCAATTTTT
GTGGTTGGTCGAGATGACGATGATGATTCAGTTTCAGCTGGGAGTGAATTAAGTGAAGCTGCAGCTAGTTGGGAAACAGTTGATGATGATGTGAGGGACA
CTCATGATTATACTCAAGAGGCAGTTGTGCTGCAAGATGCAACTGGCATGAACTCAGAGGAGGAAGAGAGTGTGGAGAATTATAGTTCTGGAAGGAATGC
AGCACTTAATTTTCCACTCACTGCACTTGACTTTGTTGCAAGATTGGCCACTGGAATATTTTCACGAGGTCAGAAAAATATCGACCCAGATTTTTCAGGC
TACCAAGGTGAAAACAAACTGCATTCTCAGGGAACAAACTGTATTTCTGAAGAAAAAGATTCCAGTGATGAGTCCAGTGCTGAAAAATCTAATGTCAATA
ATAACTGTGGTATGCAAAATACCAATGAAAAAGACAAACATGTGTCAATGGAGGACCCAGGATCATCAAATGCAGAAGAAACTTCATGCAATTTAAGCAC
TGAAAAGTCAAATGCCATGACATGTTCTGAGGCTCGGATTCACCACTACTTTAAACATTTTGACACAGCTGAAGATCCTCTGGACCATCATTTTCTTGAT
TCAAATAGACTGATCAAAAATGGAAGGAAGTGGCTTAAGAAGGTTCAACAAGACTGGAACATCCTCCAGAACAACCTGCCAGATGAGATCTACGTACGGG
TTTATGAGGATCGAATGGATCTCTTAAGAGCAACAATAATTGGGGCATATGGAACACCTTACCAAGATGGACTCTTCTTTTTTGACTTTCACCTCCCACG
AGAGTATCCTGATGTCCCTCCTTCAGCATACTACCATTCTGGTGGTTGGCGAATAAATCCAAATTTGTACGAGGAGGGGAAGTTGTGCCTTAGCCTTTTA
AATACCTGGACTGGCAGAGGAAATGAAGTCTGGCACTCGACTTCTAGCATCCTTCAAGTCCTAGTTTCACTTCAAGGGTTGGTGCTAAATTCTAGACCAT
ATTTTAATGAAGCTGGATATGATAAGCAGATCGGGACAGCTGAAGGAGAGAAAAAATCGTTGTCATATAACGAGAATACTTTCTTGCTAAACTGCAAGAC
AATGATGTATCTTATGCGAAAACCCCCCAAGGATTTTGAATGCTTGGTAAAGGAGCATTTCAGGAGGCGTGGTTACTACATTCTTAAGGCCTGTGATGCG
TATATGCAAGGCAATTTAATTGGATCTCTCTCTCGAGATGGCTCTATAAGCAGTAAGGAAAGTTCCAATCTAACTTCGGTTGGTTTCAAGTTGATGTTAG
CGAAGATTGTGCCAAAGCTTTACTTGGCATTGAATGAGGTTGGCGCTGATTGTCATGAATTTAAGCATCTGCTACAGTCATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G176000.5 pacid=42795185 polypeptide=Potri.004G176000.5.p locus=Potri.004G176000 ID=Potri.004G176000.5.v4.1 annot-version=v4.1
MCFQLENMQNESSSKVNGFVDSGGDKNSLDQSGSMANGLVCELKVDSENNVKKLDEVVGNLNNVPYIYGQDIVRHKMHNVVGVVSEVAGESDSDGSIMDD
DDDDDEDEDDDDDDDEDECDVDEDAGNEGADDDGNDNTNGDRSEGSDNYKSETLQTDEVRVVWMGDVKPVQHVNDVTVIDRGFLHGDYVASASDPTGQVG
VVVDVNISVDLLAPDGSVRKDISSINLVRVREFSVGDFVVFGPWLGRVDDVVDDVTVLFDDGSVCKVKGAEPLHLKPISKGIFEEDEHFPYHPGQRVRAT
SSSVFKNSRWLSGLWEANRLEGTVTKVTSGSVFIYWIASAGYGPDSSTTPAEEQDPKNLELLSCFAHASWQVGDWCLLPSSVAQSSSVTLDKDLLKLGIH
DPAKCELDSSQLGNGCDSEGVANEEVDDMNGSMAIDPVTTPDGNLAIIASNESSSCGSSTSVSKVPAHRRKIRKVLIRREKKSIKKEEDFERALLIVNTR
TRVDIAWQDGAIERGLNSTTLIPIDSPGDHEFVAEQYVVEKASDDVDNSFESRRVGVVKSLNAKERTACVKWLKPVTRAEDPREFDKDEIVSVYELETHP
DYDYSYGDVVVRLSPVSVSDQTTSDLETIGESRQQSGQSEIMNTQKCLGHKKGEDAPSNDVSMDSSDLSWVGNISGLRNGDLEVTWADGMVSMVGPQAIF
VVGRDDDDDSVSAGSELSEAAASWETVDDDVRDTHDYTQEAVVLQDATGMNSEEEESVENYSSGRNAALNFPLTALDFVARLATGIFSRGQKNIDPDFSG
YQGENKLHSQGTNCISEEKDSSDESSAEKSNVNNNCGMQNTNEKDKHVSMEDPGSSNAEETSCNLSTEKSNAMTCSEARIHHYFKHFDTAEDPLDHHFLD
SNRLIKNGRKWLKKVQQDWNILQNNLPDEIYVRVYEDRMDLLRATIIGAYGTPYQDGLFFFDFHLPREYPDVPPSAYYHSGGWRINPNLYEEGKLCLSLL
NTWTGRGNEVWHSTSSILQVLVSLQGLVLNSRPYFNEAGYDKQIGTAEGEKKSLSYNENTFLLNCKTMMYLMRKPPKDFECLVKEHFRRRGYYILKACDA
YMQGNLIGSLSRDGSISSKESSNLTSVGFKLMLAKIVPKLYLALNEVGADCHEFKHLLQS
|
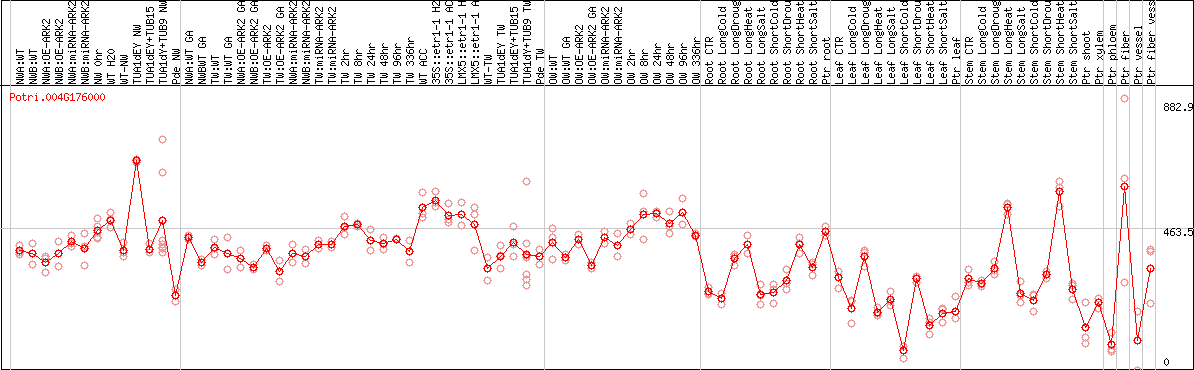
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G176000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
