External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G27740 177 / 3e-57
Family of unknown function (DUF662) (.1)
AT3G09980 161 / 6e-51
Family of unknown function (DUF662) (.1)
AT2G36410 159 / 1e-49
Family of unknown function (DUF662) (.1), Family of unknown function (DUF662) (.2), Family of unknown function (DUF662) (.3)
AT3G52920 154 / 5e-48
Family of unknown function (DUF662) (.1), Family of unknown function (DUF662) (.2)
AT3G52900 140 / 1e-42
Family of unknown function (DUF662) (.1)
AT5G03660 140 / 2e-42
Family of unknown function (DUF662) (.1), Family of unknown function (DUF662) (.2)
AT2G36355 128 / 4e-38
Family of unknown function (DUF662) (.1), Family of unknown function (DUF662) (.2), Family of unknown function (DUF662) (.3)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019529
193 / 2e-63
AT2G27740 201 / 9e-67
Family of unknown function (DUF662) (.1)
Lus10043372
189 / 6e-62
AT2G27740 207 / 6e-69
Family of unknown function (DUF662) (.1)
Lus10027724
159 / 8e-50
AT2G36410 237 / 1e-80
Family of unknown function (DUF662) (.1), Family of unknown function (DUF662) (.2), Family of unknown function (DUF662) (.3)
Lus10035563
159 / 9e-50
AT2G36410 238 / 8e-81
Family of unknown function (DUF662) (.1), Family of unknown function (DUF662) (.2), Family of unknown function (DUF662) (.3)
Lus10023876
154 / 2e-48
AT2G36410 239 / 9e-82
Family of unknown function (DUF662) (.1), Family of unknown function (DUF662) (.2), Family of unknown function (DUF662) (.3)
Lus10014379
132 / 8e-39
AT2G36410 214 / 4e-71
Family of unknown function (DUF662) (.1), Family of unknown function (DUF662) (.2), Family of unknown function (DUF662) (.3)
Lus10034160
118 / 4e-34
AT3G52900 154 / 2e-48
Family of unknown function (DUF662) (.1)
Lus10043423
117 / 3e-33
AT3G52900 150 / 4e-46
Family of unknown function (DUF662) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04949
Transcrip_act
Transcriptional activator
Representative CDS sequence
>Potri.004G186900.1 pacid=42795285 polypeptide=Potri.004G186900.1.p locus=Potri.004G186900 ID=Potri.004G186900.1.v4.1 annot-version=v4.1
ATGACGACACAAAAACCAATATTGGAGCAACAACAATCACAAATGCAGATCAGAATGATGAAGAACTCAGGTATTATCAGTAATATCAGTGAAAGCCCAC
TATCAAGGGATGATAAGGATGATGAGATGTCAAGATCAGCAGTGGCTATGTTTAGGGCGAAAGAAGAAGAGATTGAGAAGAAGAAGATGGAAGTCAGAGA
CAAGGTTCATGCTCACTTAGGACGTGCGGAAGAAGCAACTAAACGATTAGCAGAAATTCGTGAAGAGCTTGAAGCCCTGACAGATCCAATGAGAAAGGAA
GTTTCGATGGTACGTAAGAGGATTGACACAGTCAACAGGGAGCTGAAGCCTCTAGGACTGAGTTGCCAGAAGAAGGAAAGAGAATACAAAGAAGCCCTGG
AAGCCTTTAATGAAAAGAACAAAGAAAAAGCTCAGCTGGTTTCCAAATTAGTGGAGCTGGCGAGTGAAAGTGAGAAACTGAGGATGAAGAAACTGGAGGA
ACTGAGCAAAAATATTGAAACTTTAAGCTGA
AA sequence
>Potri.004G186900.1 pacid=42795285 polypeptide=Potri.004G186900.1.p locus=Potri.004G186900 ID=Potri.004G186900.1.v4.1 annot-version=v4.1
MTTQKPILEQQQSQMQIRMMKNSGIISNISESPLSRDDKDDEMSRSAVAMFRAKEEEIEKKKMEVRDKVHAHLGRAEEATKRLAEIREELEALTDPMRKE
VSMVRKRIDTVNRELKPLGLSCQKKEREYKEALEAFNEKNKEKAQLVSKLVELASESEKLRMKKLEELSKNIETLS
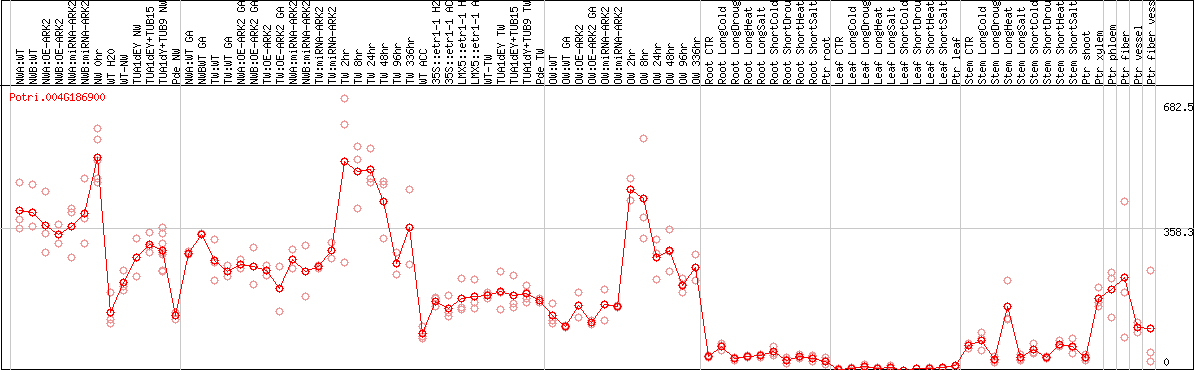
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G186900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.