Potri.004G193700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G193700.1 pacid=42794802 polypeptide=Potri.004G193700.1.p locus=Potri.004G193700 ID=Potri.004G193700.1.v4.1 annot-version=v4.1
ATGCCTTTCCTATCGGACACGGCGAGCGCGATTAAGAGCCGGTTCGGGTTTCATGACCGGTCCGTCTCGGAATCTGTGCCGAGCACTCCAGATCTACTGA
AATCCGTCTCTCGAGATCATAACCTTGCTTCCGCCCAATCCTTGGTTGCCATGTCAGCAGCCCGGAGGATTGATGACTGGGAGGAGGACGACAACGGTGG
CGTTACAGGATCAGTGGTGTCGGTGCCGCGGCATGCTCAGAGCTTTGAGTTCTCCGAGGATCCTTCATTCTGGAAGGAACATAACGTGCAGGTTATTATA
AGATTAAGACCGTTAAGCAGTTCAGAAATATCTGTTCAAGGGCATGGCAAGTGCGTTAGGCAAGAAAGTTGTCAGACAATTACTTGGACCGGGCATCCAG
AGTCACGGTTTACTTTCGATCTTGTTGCGGATGAAACTGTTAGCCAGGAAAAAATGTTCAAAATGGCTGGATTGCCGATGGTGGACAATTGTATGGGAGG
CTACAACAGCTGCATGTTTGCATATGGCCAAACGGGAAGTGGAAAGACTCATACAATGCTTGGAGACATCGAGGGAGGGACTCGAAGACATAGTGTTAAT
TGTGGAATGACACCTAGGGTGTTTGAGTATTTGTTCTCAAGAATCCAGAAGGAAAAAGAGGTTCGTAAAGATGAAAAGATAAAATTTACCTGTAAATGCT
CTTTCTTGGAAATATACAACGAACAGATTCTTGATCTCCTGGACCCGTCATCTACCAACTTGCAGATAAGAGAAGATGTGAAGAAAGGGGTTTATGTGGA
GAATCTCAAGGAGATAGAAGTTGCAAGTGCTCGAGATGTGCTTCATCAACTTATTCAGGGTGCAGCAAATAGAAAAGTAGCAGCCACAAATATGAATCGT
GCAAGTAGTCGTTCTCATAGTGTATTTACATGCATCATTGAAAGTAAGTGGGAATCCCAAGGCGTAACTCATCATCGGTTTGCTCAGCTTAATCTTGTTG
ATTTAGCAGGATCTGAGAGGCAAAAGAGTTCTGGTGCTGAAGGTGAACGTCTCAAGGAAGCTACCAATATTAACAAGTCACTTTCGACATTGGGACTTGT
TATTATGAACCTTGTAAGTATATCAAATGGCAAGTCGCACCATGTTCCTTATCGGGATTCGAAGCTCACATTTTTGCTTCAGGATTCTCTGGGAGGAAAT
TCAAAAACCATCATAATTGCAAATATCAGTCCATCTCTTTGTTGCTCTTTGGAGACGTTAAGCACATTGAAATTTGCACAACGAGCAAAATTCATAAAAA
ATAATGCAATTGTTAACGAAGATGCATCAGGAGATGTTATTTTGATGCGCCTGCAAATTCAACAACTTAAGAAAGAAGTGTCCCGACTGCGTAGTTTAGT
CAATGGGGGAGCTGAAAATCTAGATAGTGATACTTCATCCTTAAGCTTTCTCGGATCTCCTGGAAGTTTTAAGTGGGAAGGGTTTCATGGATCATCTAGT
CCGCTTATGTCTGAAAATAGGATGTCTCAGAAGAAAGATTATGAAGTTGCCCTTGTCGGGGCTTTTAGAAGAGAAAAGGATAAGGACATAGCTTTGAAGG
CATTGACAGCTGAAAATCAAGCAGTTATGCAACTGACCAAACAGAGAGAGGATGAGATAAAAAGCTTAAAAATGATACTACGATTCCGAGAGGCAGGAGT
TAAGAGGCTAGAGGCAGTTTCTGCTGGAAAGATTTCAGCTGAAACACACTTGTTGAAAGAAAAGGAGGAGCATCTGAAGCAAATTGAAGTTTTACGAACC
CAGGTTGATCGCAACCAAGAAGTCACAAGATTTGCCACAGAAAATTTGCGTCTAAAGGAAGAAATTAGAAGATTAAAGTCATTTTGTGAGGAAGGTGAAA
GGGTGATGATGAATGAACAAATCATGGTCTTACAGAATAAGTTACTAGAAGCACTGGATTGGAAACTTATGCATGAATCAGATTCCTTGGCAGTTCAGAA
AACTAGCTTAGACATGGAGACAGCAGCAGTACATGATGATCCACCAATTTTGAGTCAGGAACCAAGATCACCTTGTCAATCTTCAATTAAAGAGGAGAAC
GAATTCCTCCGCATACAGGTTATTCAAAACCAGGCCGAAATTGACTTGCTCCTTAAGAAGTTGGATTTCTGTTTTGAAGAGAAGGAAAGACTAGAAAGGC
ATGTTAATGATCTCATGATGAAACTTGAAGAAGAGAGATTTTTTAGAGCCACAAACGAGAAAACAGAGCAACTTGAGCTTCCTTTGTCAACTGATGCATC
AGTTATCAATGTGGAGCTCAAAACAATGGTTGATGCCATTGCCGCAGCAAGTCAACAAGAAGCTGAAGCTCATGAAAAAGCTATTACATTGTCTACGGAG
AATAATGAGCTGCAGTTGAAGCTTGAAACATTTATTGAGGCAAACAAGGAGCTACAGTCAAAACTAAAAGCTTTGATTGAGGAAAAAAATAGTCTCATTG
AAATGTATGAACGGGCAGCTTCAAAAAGTAACTACAAGAACGTGAATGATGATGAGAGTGAACAAAATGGCATGGAAGTTCATGATAATGATAGTCCTCC
TGAACTTGCCAATGCACGTGAGAGGGAGATGAAAACAGTTGAGAACCTAGAGCAGCAGCTTATGGAAACGCATGAAGAAAATGAAAAGCTGATGGGTTTG
TATGAGATAGCTATGCATGAGAGGGATGAACTGAGAAGATTGCTTTCTTCTGATGAACGGAACAGAGTTGGAGAGAAACTAATGGAAGTTGATGGAGAGA
AATGTCTTCAGTCTCCTGCATCTCCACTGAACGCGGAGCAAGCTTTCATTGAGTTTGATAAAGTTCTGAGAGAAATTGAAGTAACAGAAGAAGGGCTTCA
GATAAAACAGGAAGAGTTTCGATCCCTTGAGCTACTTTCCTCTGAAATGCAAGAGAAAAGAGCTCTTGTAGATAAGAAGTTATCAGCACTGAGATATTCC
GTGTCAAGCTTTGTTTCATCAATTGCTTACTTTGAACAGCGAGAACACAGAGCAAAGGCAAGGGTAAATGCTTCAACGTCTTACCTAGAGAAAAAGAAAG
AAGAGCTGGCTCGCCTTCAAGTTTGCAAGGAAGAATCTGAGGCCACCATGGGAAGGATTCAACAATCTGAAATTGAATTGAAGAACGTACTGGCAGTTTT
GAAATCCAAACTTATGGAGGAGAATCAGAGACAGGAGAGTGAAAAGGTCCTTTTTCCCATAGATAACATTGAGAATGTTGATACCTCACAGAGAAATTGG
CAATTAGGCGGTAAAGCTACAGAGTTATTGAAGTCCGAGGAAGAGAAGACCAAGATACAGAATGAAATGAAGCTGTCCCGCGAAAATCTTGGTCTTATAA
AGAAAGAATTTGATGTTTTGAGTAAAAAGTTGGGTAAGATAGAGGGTGAAATGCAAGTTGTACAAACGGATATACAGAAGGGATCCAAATCTGTGGAGGA
AATGGAACTTGCACTTCAGGCTGTAACCCACGAGAAGGAAACTCTCTTAGAAATCAGAGAGGCTGGAATGCATGAAATTCAGAGTATGATTCTTGAATAT
CAGCAAAGCGTCTTCGATGCAGATCTGAAGGGGGCGGAGAAGGAGATGTTGGAAGAAGAATTACAGCTGGAGCTGAGAAGAAATGAAGAGTTGAAAATAC
AGCGGGCAGAAGCTTCTGAGAAGATGACCAAGCTGTTGGAAAACACAAGCTCCCGTCCATGCTTTGCTAGAAAAATGGAAGAGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G193700.1 pacid=42794802 polypeptide=Potri.004G193700.1.p locus=Potri.004G193700 ID=Potri.004G193700.1.v4.1 annot-version=v4.1
MPFLSDTASAIKSRFGFHDRSVSESVPSTPDLLKSVSRDHNLASAQSLVAMSAARRIDDWEEDDNGGVTGSVVSVPRHAQSFEFSEDPSFWKEHNVQVII
RLRPLSSSEISVQGHGKCVRQESCQTITWTGHPESRFTFDLVADETVSQEKMFKMAGLPMVDNCMGGYNSCMFAYGQTGSGKTHTMLGDIEGGTRRHSVN
CGMTPRVFEYLFSRIQKEKEVRKDEKIKFTCKCSFLEIYNEQILDLLDPSSTNLQIREDVKKGVYVENLKEIEVASARDVLHQLIQGAANRKVAATNMNR
ASSRSHSVFTCIIESKWESQGVTHHRFAQLNLVDLAGSERQKSSGAEGERLKEATNINKSLSTLGLVIMNLVSISNGKSHHVPYRDSKLTFLLQDSLGGN
SKTIIIANISPSLCCSLETLSTLKFAQRAKFIKNNAIVNEDASGDVILMRLQIQQLKKEVSRLRSLVNGGAENLDSDTSSLSFLGSPGSFKWEGFHGSSS
PLMSENRMSQKKDYEVALVGAFRREKDKDIALKALTAENQAVMQLTKQREDEIKSLKMILRFREAGVKRLEAVSAGKISAETHLLKEKEEHLKQIEVLRT
QVDRNQEVTRFATENLRLKEEIRRLKSFCEEGERVMMNEQIMVLQNKLLEALDWKLMHESDSLAVQKTSLDMETAAVHDDPPILSQEPRSPCQSSIKEEN
EFLRIQVIQNQAEIDLLLKKLDFCFEEKERLERHVNDLMMKLEEERFFRATNEKTEQLELPLSTDASVINVELKTMVDAIAAASQQEAEAHEKAITLSTE
NNELQLKLETFIEANKELQSKLKALIEEKNSLIEMYERAASKSNYKNVNDDESEQNGMEVHDNDSPPELANAREREMKTVENLEQQLMETHEENEKLMGL
YEIAMHERDELRRLLSSDERNRVGEKLMEVDGEKCLQSPASPLNAEQAFIEFDKVLREIEVTEEGLQIKQEEFRSLELLSSEMQEKRALVDKKLSALRYS
VSSFVSSIAYFEQREHRAKARVNASTSYLEKKKEELARLQVCKEESEATMGRIQQSEIELKNVLAVLKSKLMEENQRQESEKVLFPIDNIENVDTSQRNW
QLGGKATELLKSEEEKTKIQNEMKLSRENLGLIKKEFDVLSKKLGKIEGEMQVVQTDIQKGSKSVEEMELALQAVTHEKETLLEIREAGMHEIQSMILEY
QQSVFDADLKGAEKEMLEEELQLELRRNEELKIQRAEASEKMTKLLENTSSRPCFARKMEE
|
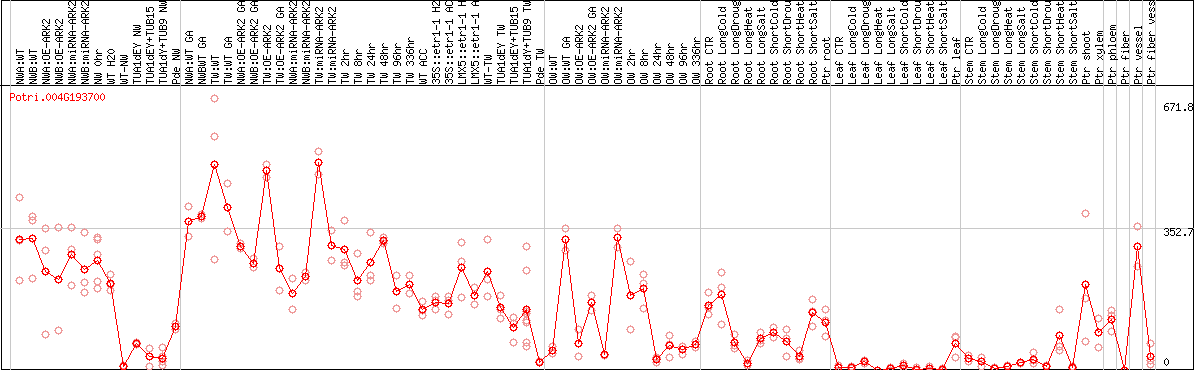
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G193700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
