Potri.004G212700 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G212700.5 pacid=42794862 polypeptide=Potri.004G212700.5.p locus=Potri.004G212700 ID=Potri.004G212700.5.v4.1 annot-version=v4.1
ATGATACTAAAGCTAGTTGCAAAGCTAAGGATGAGGAAACCAATGGTTCAAAAAGGGAAAAACATTGTTAGCAGATCATCATCTAGTTCAGGTAGTGCGC
CTGATACATCTGCTCTAAGAAGGTCAGCCAGAGAAACATCATCAAAGAAAAGCATGGCTCCAAGCCCTTCAAGTACTAGGAAGTCTGAACGTCTTGAGAA
TCGGACCCCTACAACTCCTGCAGTCATGAGGAAATCCGGGAGAGTTGAGAAGCAAAGCATGTTGAGTCCTTTGAGAAGGTCTAAGAGGGGTAAGAATCAG
TCATCATCCAGTTCTTTTGGTTCAAAGAAGTCTGGTAAAAGCTTGGGCTCATCAGTTATGAAAAAAAAGCACAGGAAAGAGAAGAGTGTGAAACTGTTGA
CTCTTGAACCTAATGAAATCGGCTACAGTGAGAAACATATTATCAAAGCTGTTCAGGTTGAAACAAAGATAACTGATGCTCGTGTTTACAGGTCATTGTT
CAAACAGCAACAAAAGAAAGCTAATCTGGAAGGTTTTTGTGAGGAGACGAAGAACAAGAAGGCTAAGTCATCTCAAGGATATTGCAGCAATCTCAGAGCT
AGTGCTTCTGAGAACGTTGATGGGGGTGGTGACTGCAGTCAGAGGGAGGTAGAGGAATTGATTGAAGAGTGCATTTTGAAAGATTCTGAGAAAAAAATGT
TAGGAAATTCCGTTTTGTCAGCTGGAGGTTCACCTGAGTTTGATAATAATCATCTGGACTTTGTCAACTATCTTTTAGAGTGCTGGCACAGGGGAGAGAA
TGTTGTTCTTATTGATGATCAGGAACAAATTGCAAAGGTGATTTATTTTATTTTGTCAATATCATCCAATGCCACTTGGCCATTTCTTATCATCACGACT
TCTGCTGCACTTCATTCATGGGAAGAAGGGCTATTTCGTTTGGCGCCATCTCTATATGCTGTGGTTTATCATGGAAACAAAGACATACGGAAAAGTATTA
GGACGTTGGAGTTTTACAGTGAAGGAGGTTGCATTATGTTTCAAATACTTATAACCTCACCAGAAGTTATTATCGAGGATCTAAATATGCTTGAATCAAT
GAAATGGGAAGCCATAATAGTTGATGAGTGCCAACGCTCCAGAATTTATTCACATTTTAAACAAATTAAGCTGCTAAGCACTGCTATGCGGCTTCTCCTT
GTAAATGGTCAGCTTAAGGATGGCATCACTGAGCACTTGCTGTCTCTGCTTGTTCAGCAGAGTGATCCGGATGGTAGTGAATGCTTGGTGATTGATTCCA
GCCATAAGACTGGCATTTTTAAGGAAAGGCTGTCACAATACATTGCAAATGGTTGCAAACCAGATTCCTCAAGGCTTAAAGAGTACTGGGTTCCTGTACA
GTTATCCATCATGCAGCTTGAGCAGTATTGTGCCATTCTACTTTTGAACTCCTTATTACTTTGCTCATCCTCAAAGAATGATCTTGCTGGTTCCCTACAT
GATATTCTTATTTCTGCTCGAAAGTGCTGTGATCATCCCTACATCATGGATCCATCGCTACAAATTTCATTAACAAAAGACAGTAAAGAAGCTGACATTT
TAGATATTGGAATAAAAGCTAGTGGCAAGCTACAACTCCTGGATGCAATGCTTTTCAACATAAAAGAGAGGGGATTAAGAGTGCTAGTTCTTTTTCAGTC
TAGTGGAGGTTCTGGAAAGGATAATGTTGGAGATATTTTAGATGACTTTATACGTCAAAGATTTGGCAAGGGTTGTTATGAACGTGTTGATGGGCATGTA
CTCCCTTCAAGGAAGCAAGCTGCTTTGAAAAACTTCAACAATCTTCAGGAAGGGAGGTTTGTGTTCTTATTAGAAACCCGTGCTTGTAGTCCCAGCATAA
AACTTTCATCTGTGGATACTGTTATCATATTTGCCAGCGATTGGAAGCCAAATACTGATATAAGGAACCTGCAAAAAATAACGCTTTATTCAGAGTCTGA
ACAGATAAATATATTTCGTCTATACTCTTCTTGTACTGTGGAAGAAAAAGTTTTAATTGTTGCAAGGCAAGATAAGACACTTGATAGAAATTTACAGAGG
ATAAACCAGGGTGCTAGTCACATGCTTCTGATGTGGGGGGTTTCATATCTGTTTGACAAATTGTCAGAGTTTAATTGTGGCAATGACCCTGCCTCCAGTG
GAACATTGTTGTCTGAGCAGTCACATATGAAAGATGTTATACAGGAGTTCTTAACCATAGTAACTCAGAAGGGAAAAGACAAGAATCTAATTAACTCTAT
AATTTTGAATGTTAAACAAAATCAAGGAAGTTATACTACTAATCTTCCATTGCATGGTGAGCCAAAAATTCAATTGTTGGATGAAGAGCTGCCTCATGTA
TTTTGGGAAAGGTTACTTAAGGGAAAACAGCCACAGTGGAAATATAGTTCTGGTTTGTTTCAGAGGAACAGAAAAAGGGTTCAATATTTTGATGACACAC
AGAAGAACCCTGAAGTCGAGGCTGATGAAGTTGTAAAGAAGCGCAAGAAAGTGGCCATTGATAACTCCAATTCACCATCTCTAAAAGCTGCTCCAATAGG
AACTTCTGGAGCTCCAGTGTGTAGCATGTCTCAATTCATGCCAAGTTCAACTGGTTGTCTTACTACAACTGATGCAAATCATGTTTCCAACTTCACCCAT
TTGAATAATAAATTATCACTTCTACCAAAAGCTAATACGGTTGATTATAATGAAAGAATGAATTTACATTATTCTCGGAAGAGTCTCCATCTCGTTTTGA
AGCCAGAGATAGAAAAACTTAGTGAAATTTTACAACTCCCAGAGGATGTCAAGGTCATGGTTGACCAATTTCTGGAATATGTACTGAATAATCACCATGT
CAGTAGAGAGCCAGCTAGCATATTGCAAGCATTTTTGATATCCTTGTGTTGGACTGCAGCTTCCATGATTAAGTACAAACTTGATCGCAAAGAATCACTT
GCACTGGCAAAACAGCATTTGAACTTTTGCTGCACGAAGGATGAGGCAGATTTTGTTTATTCAAAGTTGCGATATCTGAAGAAAGTATTTCTTTATCATA
CAGGAAATTTTAAGTTGGCTGGCTCTCCAAAAGCTGCAGAATTTTCAACTAAAGATCTCAGTACAAATCAGTCAAATGGAAGACCTTCACTGTCAACACC
ATCTAACATGCAAAAGGTTAGAATAGAGGTTGAAAACCTGAGACCTAGTCAAGAATTCTTCATTGATCAGGCCCTTTCCCATCTAGGATTGACACAGAAA
GATTACTCAGAAAACATTGGAGAGAAATGTGATGAACAGATGAACAAGCTATTGCAGAGGCAACGAGAAGAAAGGGAAGAGTTCAAAAAAAAGTACGAGG
AAGAAAAGGCAGAACTTGAGCTTATGCAGAGAACAGAGGCAGCTGTTATTCATTTACACAGTAATAGTTCAATGAGAACAGATAAACTCAAGGTACTGGA
TAATGTATTTGCAAAAGAGTTCAGAGAGCTTAAACGGAAGATGGAGATACGCCTTAATAATGTTTTGGAATTTCAACTGGCTACAAGGAACAAGTTGCAA
GAGAGGAAGGCTCATTGGATTGGAGTGAAATTGTCTGGATTACTAAATAAACCACTTGCAGATGAGTCTGGATATGATCAACAAAACGCTGCAACTTTGA
ATTCTTGTTCCAAAGAACAGACTTCTGAACGAGCCCAAAGCATGCCAGATGGAGAGGTTCTATTGGAAGCTCTTGAAACTGTGAGTTTGAATGAGGATGT
ATTTTCTGGGGTATTGTCTGCATCTGAACCAATGTTTGATGGGGCTAGTTCCAGCATGCTAGACAGAGAGGTTCCTTTGGAAATGCCCCAGACTGCAAGT
GTGAGGAATATTTCAGAGAATATTGTCTATTTGAATGCATCTTCAGGTGAAGGACAAATTCCTGTTACACAGGTTGCAGTGAGAGTCCTTGAGGCTATCA
GCTCTAGTGATGGCCCAGAGAATACCATTCATAAATCATCATCTGAAAGTCGTAATAGAGATGCATTAATGGTGCCTGATAGTGAATTTCCCTTGGGAGT
GACTGAGATTGTCAGTTCCACTGGTGGCCTGGAGAATGCTGCTTCTGCAAATCCTTCACCATCTGAAGGATGTACTGTCAGAACAACTTCATGCATGGAT
GGCAGAGAAGTTCTGTTGGAAGTGCCTGAAACCGCTTCTTTGGAAGCTGAGCATGGTAACAGAGTTATGGAGAAAGATGGAATATCGGCTATGGTATCTG
ATAATGCCACTGAAGAAGATCAGCAGAATGGATTGGTGTCTATGCTTAATCAAGATTCTCAATCTGATAATATTATTGCAGTCAACCAGCAAAATGGAGA
GGTTCTTTTAGGAGTGCCCCAAACCAATGAAGTTGGCCTGCAGGATGAGGAGGTTCCATCAGGAGTTCATGGAACTCCTGTAGAAGGGAGTGCTAGTAAT
GGTGGAGAGAACACTGGTGTTTATGTTACAGCCTTTTCTATCGGTACTGGAGTTGATCAGCTGGCTGGAGTGCTTCCATCAGGAGGGTTTGAAACCGCTA
CTAGTGCAGAATTGGAGGGTAGCCGCACTCAGCGAGAGATTGACAGCATACATGCTGTAGCCTCAGATACTAGTCAATCAGCGGAATCCTCTAGACTGCA
AGATGGAGTTGCACAAGTTTGTGATAACCAGATTGCTTTTCAACAAGTTGATGCCTCAGCATCCCAACCTCTTGTTGTAGCTTCTGGCCAATCTCCTAAT
GATGCATCTGTCACTGAGCATCTGCTGGAATTGTTACTATCAACTGGCTCTCCTACTCCTAGTGGTTCACAGCCTGCAACTTCATTTGCACAACTTTCAC
CTATTGACTCGATAGCTGTTGGTGGATCAGGGATGCATATTTCAAACATGAGGGCTGCACCTGTTACCCCTGGAATTAGTAATCGTCCTGGAACTGCACT
TGCTGTGCGAATGCCTGTATCTATGTCTCAAGATCCACTTCAAAATGAGTTGGATAGATTAAGCAAAGAAACTGAAGAAATCATCAAGATCCATGAAGAT
ACAAAGCTGCAGTTGAAATCTGATTGCGAGAAGGAGATAGTGGAGGTTGTAGCTCAAATTCATAAAAAACATGATATTAAACTTCAGGAGATAGAATCTG
ATTTCCAGTGTAAGAAGAAAGAGATGAATGACAATCAAAACAAAGTTCTTATGAACAAGATATTGGCTGAGGCTTTCAAGACCAAGTGCATGGATAGTAG
GGCATCTAGCACCCTTGGCAAGCAGCAAGAGATCACCTCTAGTGCTGTGCAGCAGCTACTTCGGCAATCTCAACCCACTGCTCAGAGGCCTCCTATTGTT
GCTTCATCTGGTGTTTCTGCAGATGGTCATCAAACAAGTCCCTCACTTTCCCCTCCCTCCCCACCTCTAGAGGTTGTTCGTTGCTCATCACTTTTATCAG
GCACCCCTACCAGACCACCACATATTGGTTCCATCTCTCCTATCACTAACAATCTCCAACTTGGTAGTGGGATTCGAGCTCCTGCCCCACACCTGCAACC
TTTTAGACCTTCAGCATCCATATCAACAACTGGTCTTTCTTCCTTTCTACATGGGATGCAAAGTCAACAAGTACCTTCAACTTCTCCCACGCTTTCTGAG
ATTCCTTCACGAGCTCCAGCATCTGTGCAACAATCTGGTCCTCAGACGACGACAAATTGTTGTGAAAGCATGGGGGTCTCGCCTTCTAGCACATATCTCT
CTGGACTTGATTCGTTAATGGATGGAGGTTATCAAACAAGTACAAATGCAACTCAACCCTGCAGTTTTCCTCCAGTAACAAATTTGATTTCAAACCCAAA
CCAGTTGACCCAGCCAGAGCTTAGCATGCTACATAGTGTGAATAGTGTGCTGACAAACCCAGCAAGTGAGGAATTGCAGTTGTCTCCAAGCTTTTGTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G212700.5 pacid=42794862 polypeptide=Potri.004G212700.5.p locus=Potri.004G212700 ID=Potri.004G212700.5.v4.1 annot-version=v4.1
MILKLVAKLRMRKPMVQKGKNIVSRSSSSSGSAPDTSALRRSARETSSKKSMAPSPSSTRKSERLENRTPTTPAVMRKSGRVEKQSMLSPLRRSKRGKNQ
SSSSSFGSKKSGKSLGSSVMKKKHRKEKSVKLLTLEPNEIGYSEKHIIKAVQVETKITDARVYRSLFKQQQKKANLEGFCEETKNKKAKSSQGYCSNLRA
SASENVDGGGDCSQREVEELIEECILKDSEKKMLGNSVLSAGGSPEFDNNHLDFVNYLLECWHRGENVVLIDDQEQIAKVIYFILSISSNATWPFLIITT
SAALHSWEEGLFRLAPSLYAVVYHGNKDIRKSIRTLEFYSEGGCIMFQILITSPEVIIEDLNMLESMKWEAIIVDECQRSRIYSHFKQIKLLSTAMRLLL
VNGQLKDGITEHLLSLLVQQSDPDGSECLVIDSSHKTGIFKERLSQYIANGCKPDSSRLKEYWVPVQLSIMQLEQYCAILLLNSLLLCSSSKNDLAGSLH
DILISARKCCDHPYIMDPSLQISLTKDSKEADILDIGIKASGKLQLLDAMLFNIKERGLRVLVLFQSSGGSGKDNVGDILDDFIRQRFGKGCYERVDGHV
LPSRKQAALKNFNNLQEGRFVFLLETRACSPSIKLSSVDTVIIFASDWKPNTDIRNLQKITLYSESEQINIFRLYSSCTVEEKVLIVARQDKTLDRNLQR
INQGASHMLLMWGVSYLFDKLSEFNCGNDPASSGTLLSEQSHMKDVIQEFLTIVTQKGKDKNLINSIILNVKQNQGSYTTNLPLHGEPKIQLLDEELPHV
FWERLLKGKQPQWKYSSGLFQRNRKRVQYFDDTQKNPEVEADEVVKKRKKVAIDNSNSPSLKAAPIGTSGAPVCSMSQFMPSSTGCLTTTDANHVSNFTH
LNNKLSLLPKANTVDYNERMNLHYSRKSLHLVLKPEIEKLSEILQLPEDVKVMVDQFLEYVLNNHHVSREPASILQAFLISLCWTAASMIKYKLDRKESL
ALAKQHLNFCCTKDEADFVYSKLRYLKKVFLYHTGNFKLAGSPKAAEFSTKDLSTNQSNGRPSLSTPSNMQKVRIEVENLRPSQEFFIDQALSHLGLTQK
DYSENIGEKCDEQMNKLLQRQREEREEFKKKYEEEKAELELMQRTEAAVIHLHSNSSMRTDKLKVLDNVFAKEFRELKRKMEIRLNNVLEFQLATRNKLQ
ERKAHWIGVKLSGLLNKPLADESGYDQQNAATLNSCSKEQTSERAQSMPDGEVLLEALETVSLNEDVFSGVLSASEPMFDGASSSMLDREVPLEMPQTAS
VRNISENIVYLNASSGEGQIPVTQVAVRVLEAISSSDGPENTIHKSSSESRNRDALMVPDSEFPLGVTEIVSSTGGLENAASANPSPSEGCTVRTTSCMD
GREVLLEVPETASLEAEHGNRVMEKDGISAMVSDNATEEDQQNGLVSMLNQDSQSDNIIAVNQQNGEVLLGVPQTNEVGLQDEEVPSGVHGTPVEGSASN
GGENTGVYVTAFSIGTGVDQLAGVLPSGGFETATSAELEGSRTQREIDSIHAVASDTSQSAESSRLQDGVAQVCDNQIAFQQVDASASQPLVVASGQSPN
DASVTEHLLELLLSTGSPTPSGSQPATSFAQLSPIDSIAVGGSGMHISNMRAAPVTPGISNRPGTALAVRMPVSMSQDPLQNELDRLSKETEEIIKIHED
TKLQLKSDCEKEIVEVVAQIHKKHDIKLQEIESDFQCKKKEMNDNQNKVLMNKILAEAFKTKCMDSRASSTLGKQQEITSSAVQQLLRQSQPTAQRPPIV
ASSGVSADGHQTSPSLSPPSPPLEVVRCSSLLSGTPTRPPHIGSISPITNNLQLGSGIRAPAPHLQPFRPSASISTTGLSSFLHGMQSQQVPSTSPTLSE
IPSRAPASVQQSGPQTTTNCCESMGVSPSSTYLSGLDSLMDGGYQTSTNATQPCSFPPVTNLISNPNQLTQPELSMLHSVNSVLTNPASEELQLSPSFC
|
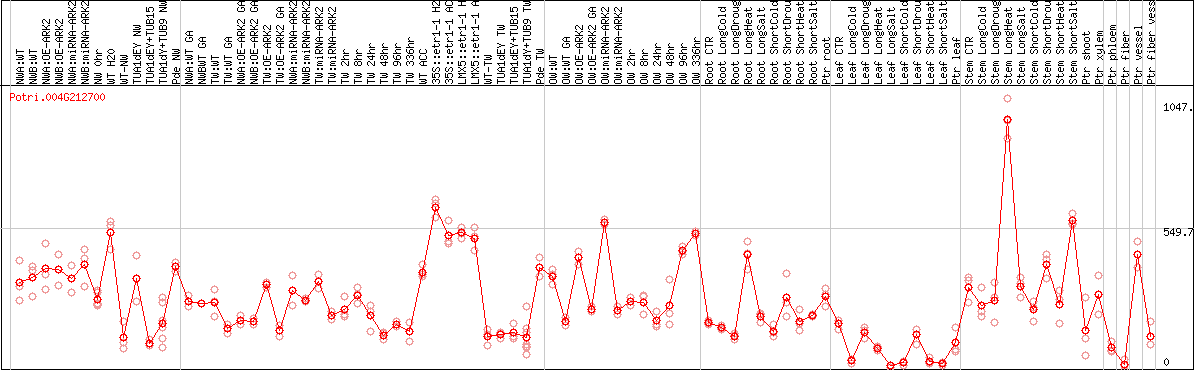
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G212700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
