Potri.004G217800 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Potri.004G217800.4 pacid=42795544 polypeptide=Potri.004G217800.4.p locus=Potri.004G217800 ID=Potri.004G217800.4.v4.1 annot-version=v4.1
ATGGATACCGGAGTAGAATCGTCGCCGGAGACGGGGGTGGTGGTTGGTGGAATTGCGCTTGATTTTCCTGTCAATGATACGGTATCGTTTTCGTCTCCGA
GGAGAATACCGAGAAAGCTTCAAAAAAGACTCTTAGAAGCCAAGACTCCAACAACCGGTAGTGTTGAAGAAATCGAAGCTAAACTTCGACACGCTCATCT
CCGTAGACAGGAATTCTATGAGAGGTTGTCAAGCAAGGCTAGACCAAAGCCAAGAAGCCCCTCACAGTGTTCATCTCATGAAGAAGACCTTGCGCAACGC
CTTGAAGCAAAGCTTCACGCCGCGGAGCAAAAAAGGCTGAGCATTCTGGCAAATGCTCAGATGCGCCTGGCTAGGTTGCATGAGTTGCGGCAGGCAGCTA
AAACTGGGGTGGAGAAGCGTTTTGAGAGAGAAAGGGAGAGGCTTGGAACAAAAGTGGAGTTACGGGTTCAGCAGGCTGAAGCAAATAGAATGCTTATGCT
AAAGGCTTACAGGCAAAGAAGGGCCACACTGAAGGAGAGAACATCCCAATCATTATTGCGAAGAAGGGCTCGGGAAAGTAAGTACAAGGAGCGTGTACGT
GCAGCAATTAACCAAAAGCGTGCAGCTGCTGAAATGAAGCGAATGGGATTGCTGGAAGCAGAAAAGAAGAGGGCATGTGCTAGGTTGTTGCAAGTTCAGC
GAGTAGCCAGGTCTGTCTCTCACCAACGTGAGATTGAGAGGAGGAGAATGAGGGAAAAGCTGGAAGATCGATTGCAAAGGGCAAAGAGACAAAGAGCAGA
ATTTTTGAGGCAAAGAGGACTTCAGCATAGTTCAGTTCGGGTAAACTGGAATAAGATGCACCAGCAGGCTGATCTCCTTTCTAGAAAATTAGCAAGGTGC
TGGAGGCAGTTCCTTAGGTCAAGGAGAACTACCATAGACCTGGCAAAAGACTATGATGCCTTGAAAATCAATGAGAACTGTGTGAAGTCAATGCCATTTG
AGCAGCTTGCTCGTCTGATTCAATTGACTGGTACTCTTCAGACTGTAGAAGGTCTACTTGATCGTCTTGAGAGCCGCCTTAGAGTCTCAATGGCTGTTGC
TGCCATGGATCATCCTTCCAGCTTGGACAACATTGATCACCTTCTCAAAAGAGTTGCCACCCCTAAGAAAAGGACCACTCCCAGGTCTTGTACAAGGAGT
AGAGAGGCCAAAAAAGTAGGTGCCTCTGGGGAGTCTGCCAGACGTGCCGCTAAAATGTCTAGGTATCCAGTGAGAATAGTTCTTTGTGCCTACATGATTT
TAGGTCATCCAGATGCTGTTTTCAGTGGTCAAGGGGAGCGTGAGATTGCTCTGGCCAAGTCTGCTGAATCTTTCATCCGAGAGTTTGAGTTGTTGATAAG
GATTATACTGGATGGGCCCATGCACAGTTCTGATGAGGAGTCTGAATCCATTTCACGGAAACGTTGCACGTTCAGGTCTCAGCTTGCAGCTTTTGATAAA
GAATGGTGCTCCTACTTGAATTGCTTTGTGGTGTGGAAGGTTAAGGATGCACAGTCATTGGAGGAGGACTTGGTGAGAGCGGCTAGCCAGCTTGAGCTCT
CTATGATCCAGAAATGCAAGCTCACCCCAGGAGGGAGTAATGATAATCTTACTCATGATATGAAAGCCATTCAGAATCAGGTTGCAGAAGATCAAAAACT
TTTAAGGGAAAAGGTACAACACCTGAGTGGAGATGCTGGTATTGAGCGAATGGAAATTGCACTATCTGAAACACGATCAAAGTATTTTCAAGCCAAGGAA
AATGGAAGTCCAGTAGGGTCACCAATAATGCATCTTCCATCTCCTAGCATGCCCATATATGCTCCTTCCGTTGCTAACACTGCTAATAGAAATAATGTGA
GTGATGGTATTGAGAGGCCAAGCCATGTAGATCGCTCCTTGTTCAGAGAAGATACCTCCTCAGCTAAAGAATTTGGTTCCTCTGATGGCCCATCAGGTTC
TGCTGTTGGAAAATTGTTAACAGAGAATGAAATGATCGTAAATGAATTTCTCCATGCGAATCGCCATGGTTTTGTTGATCGATTCAATATTTCTGACAAA
GATGAAAGTAGTATCAAGGCAAAGGTGAGAGAGACAATGGAGGCGGCTTTCTGGGATAGTGTCATGGAATCAATGAAACAGGATGAGCCCAAATATGATC
GGGTCGTTCAACTGGTGGGGGAGGTGAGGGATGGAATTCAGGAGCTGGCTCCAGAGAACTGGAAACAAGAGATTGTTGAAGCTATTGATCTAGATCTTCT
CTCTCAGGTTCTAAAGTCAGGTAACCTGGACATTGGTTACTGTGGAAAGATTCTAGAGTTTGCAATAGTTACGTTGCAGAAACTCTCCTCTCCAGCCCAG
GAGGATTTAATGAAGGCCCTGCATCAGAAATTGTTGAAAGAGTTGACTGAGACTTGTCAGACCCAAGATGAATCGAAGCATCCACATATTGCTGCAATGA
TCAAGGGTCTACGCTTTGTGCTGGAGCAGATTCAGGCTCTCAAGCAAGAGATTAGTAAAGTACGAATAAGGATGATGGAGCCTTTGTTAACAGGTCCTGC
TGGTTTGGATTACCTTAGAAAAGCCTTTGCCAACCATTATGGGTCTGATTCGGATGCCTGTATCTCTCTACCGCTGACAATGCAGTGGCTTTCATCTGTT
AAGAATAGTGAAGATCAAGAGTGGGAAGAGCACAAGAATTCTCTCTCGTCTTTGAAGAGCAATGACAGTTCATCACAGGTTTTTGTACCTCTCACTACCC
TTAGAACCGGTGGAAGCTTTCTGGTCAAAACAAATGGCAGTGCAATGGGTTCTACTTCTGTACCTTCAGAGACAGATAACCAACAACCAGAACCAGAATG
CACTGGTGAAAGAATTGATCTGTTGGTGAGGCTTGGATTGTTGAAGATTGTGAGTGGGGTTTCTGGTCTGACAAAAGAAACTTTGCCTGAAACTTTTATG
CTAAACCTGTCTCGGTTGAGGTCTGTGCAGGCTGAAATTCAGAAAATGATAGTAATTTCTACCAGCATTCTTGTTTACCAGCAAACCCTCCTGACGGAGC
GAGCGGTAAACAGCAATGCAGACATGGAGAGCATTCTATTAGAGCGCGGTAATAAACTGTCAGAAGTCCTAGACCGTGTTGACGATGTGGGTATTGAAGA
AATAGTCGAAGTAGTGAGCGGGTTCTCGCAAGATGATGAGGAGAAGCATAAGCCAAGAAAGTTAGTCATGGCTAGGATGTTAGCAAAGAGCTTGCAGGCC
GGGGATCCTGTTTTTGAAATAGTTTCCCGTGCTGTTTATTTGGCTCTAAGAGGAATTGTGCTTGGGGGCAGTGGACCCCGGGGAAGAAAATTATCACAAA
CGGCACTCCGGTCAATTGGAGCTGTCATGCTGGCAGAAAGAGTGGTGGCAGCTGCTGAAGTATTAGTGGTAGCAGCTACTGTATCAATCGGCGTTCACAG
GCCATGGTATATAACTCTGACCGATAACATGTGA
|
|||||||||||||||
|
AA sequence
|
>Potri.004G217800.4 pacid=42795544 polypeptide=Potri.004G217800.4.p locus=Potri.004G217800 ID=Potri.004G217800.4.v4.1 annot-version=v4.1
MDTGVESSPETGVVVGGIALDFPVNDTVSFSSPRRIPRKLQKRLLEAKTPTTGSVEEIEAKLRHAHLRRQEFYERLSSKARPKPRSPSQCSSHEEDLAQR
LEAKLHAAEQKRLSILANAQMRLARLHELRQAAKTGVEKRFERERERLGTKVELRVQQAEANRMLMLKAYRQRRATLKERTSQSLLRRRARESKYKERVR
AAINQKRAAAEMKRMGLLEAEKKRACARLLQVQRVARSVSHQREIERRRMREKLEDRLQRAKRQRAEFLRQRGLQHSSVRVNWNKMHQQADLLSRKLARC
WRQFLRSRRTTIDLAKDYDALKINENCVKSMPFEQLARLIQLTGTLQTVEGLLDRLESRLRVSMAVAAMDHPSSLDNIDHLLKRVATPKKRTTPRSCTRS
REAKKVGASGESARRAAKMSRYPVRIVLCAYMILGHPDAVFSGQGEREIALAKSAESFIREFELLIRIILDGPMHSSDEESESISRKRCTFRSQLAAFDK
EWCSYLNCFVVWKVKDAQSLEEDLVRAASQLELSMIQKCKLTPGGSNDNLTHDMKAIQNQVAEDQKLLREKVQHLSGDAGIERMEIALSETRSKYFQAKE
NGSPVGSPIMHLPSPSMPIYAPSVANTANRNNVSDGIERPSHVDRSLFREDTSSAKEFGSSDGPSGSAVGKLLTENEMIVNEFLHANRHGFVDRFNISDK
DESSIKAKVRETMEAAFWDSVMESMKQDEPKYDRVVQLVGEVRDGIQELAPENWKQEIVEAIDLDLLSQVLKSGNLDIGYCGKILEFAIVTLQKLSSPAQ
EDLMKALHQKLLKELTETCQTQDESKHPHIAAMIKGLRFVLEQIQALKQEISKVRIRMMEPLLTGPAGLDYLRKAFANHYGSDSDACISLPLTMQWLSSV
KNSEDQEWEEHKNSLSSLKSNDSSSQVFVPLTTLRTGGSFLVKTNGSAMGSTSVPSETDNQQPEPECTGERIDLLVRLGLLKIVSGVSGLTKETLPETFM
LNLSRLRSVQAEIQKMIVISTSILVYQQTLLTERAVNSNADMESILLERGNKLSEVLDRVDDVGIEEIVEVVSGFSQDDEEKHKPRKLVMARMLAKSLQA
GDPVFEIVSRAVYLALRGIVLGGSGPRGRKLSQTALRSIGAVMLAERVVAAAEVLVVAATVSIGVHRPWYITLTDNM
|
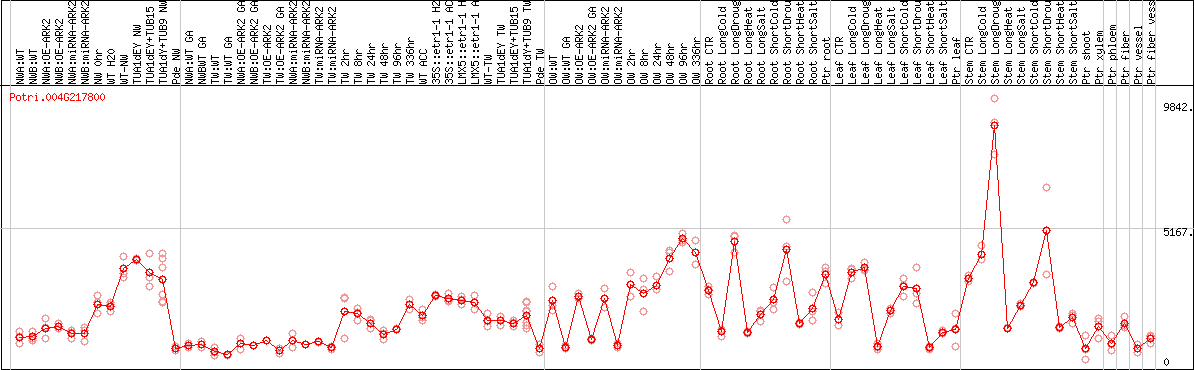
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G217800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
