External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G11850 201 / 3e-61
PLDGAMMA1, MEE54
maternal effect embryo arrest 54, phospholipase D gamma 1 (.1)
AT4G11840 197 / 1e-59
PLDGAMMA3
phospholipase D gamma 3 (.1)
AT4G11830 196 / 2e-59
PLDGAMMA2
phospholipase D gamma 2 (.1.2)
AT2G42010 188 / 3e-56
PLDBETA1
phospholipase D beta 1 (.1)
AT4G35790 184 / 6e-55
ATPLDDELTA
ARABIDOPSIS THALIANA PHOSPHOLIPASE D DELTA, phospholipase D delta (.1.2.3)
AT4G00240 183 / 2e-54
PLDBETA2
phospholipase D beta 2 (.1)
AT3G15730 144 / 5e-41
PLDALPHA1
phospholipase D alpha 1 (.1)
AT1G52570 138 / 8e-39
PLDALPHA2
phospholipase D alpha 2 (.1)
AT5G25370 131 / 4e-36
PLDALPHA3
phospholipase D alpha 3 (.1)
AT1G55180 92 / 1e-22
PLDALPHA4, PLDEPSILON
phospholipase D alpha 4 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10001293
224 / 1e-69
AT4G35790 1071 / 0.0
ARABIDOPSIS THALIANA PHOSPHOLIPASE D DELTA, phospholipase D delta (.1.2.3)
Lus10012699
224 / 2e-69
AT4G35790 1075 / 0.0
ARABIDOPSIS THALIANA PHOSPHOLIPASE D DELTA, phospholipase D delta (.1.2.3)
Lus10004156
197 / 1e-59
AT4G35790 1079 / 0.0
ARABIDOPSIS THALIANA PHOSPHOLIPASE D DELTA, phospholipase D delta (.1.2.3)
Lus10005627
194 / 5e-59
AT4G35790 969 / 0.0
ARABIDOPSIS THALIANA PHOSPHOLIPASE D DELTA, phospholipase D delta (.1.2.3)
Lus10006819
189 / 1e-56
AT2G42010 1465 / 0.0
phospholipase D beta 1 (.1)
Lus10005782
184 / 1e-54
AT2G42010 1456 / 0.0
phospholipase D beta 1 (.1)
Lus10028401
172 / 3e-51
AT4G35790 1050 / 0.0
ARABIDOPSIS THALIANA PHOSPHOLIPASE D DELTA, phospholipase D delta (.1.2.3)
Lus10014146
174 / 4e-51
AT2G42010 1254 / 0.0
phospholipase D beta 1 (.1)
Lus10042282
171 / 1e-50
AT2G42010 940 / 0.0
phospholipase D beta 1 (.1)
Lus10041855
170 / 6e-50
AT4G35790 1280 / 0.0
ARABIDOPSIS THALIANA PHOSPHOLIPASE D DELTA, phospholipase D delta (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF12357
PLD_C
Phospholipase D C terminal
Representative CDS sequence
>Potri.004G218900.1 pacid=42795815 polypeptide=Potri.004G218900.1.p locus=Potri.004G218900 ID=Potri.004G218900.1.v4.1 annot-version=v4.1
ATGGTCATGGGATCAGCAAACGTTAACAAGATATCCTTGGATGGTTCAAGGGATACAGAAATTGCTATGGGGGCTTACCAACCAACATACACATGGGCAA
GAAAAAGTTCTCGTCCACATGGACAGGTATATGGTTACCGGATGTCTCTTTGGGCAGAGCATCTAGGCAACCTCGAGGAAGCTTTCGGCGAGCCTCAACA
CCTTGAATGCATGAAACGAGTGAGAAAGATTTCCAGACACAACTGGAAAGCATATGTTTCCGAGGAGGGTAAAGAAATGAGAGGACACTTGCTACAATAT
CCAATTCAAGTGAGCAGAAGTGGAAAAGTGAGCGCATTGCCAGGTCATGAAACATTTCCTGATGTTGGTGGCAAAGTTCTTGGATCACCCACCACCCTTC
CTGATGTTCTAACAACGTAA
AA sequence
>Potri.004G218900.1 pacid=42795815 polypeptide=Potri.004G218900.1.p locus=Potri.004G218900 ID=Potri.004G218900.1.v4.1 annot-version=v4.1
MVMGSANVNKISLDGSRDTEIAMGAYQPTYTWARKSSRPHGQVYGYRMSLWAEHLGNLEEAFGEPQHLECMKRVRKISRHNWKAYVSEEGKEMRGHLLQY
PIQVSRSGKVSALPGHETFPDVGGKVLGSPTTLPDVLTT
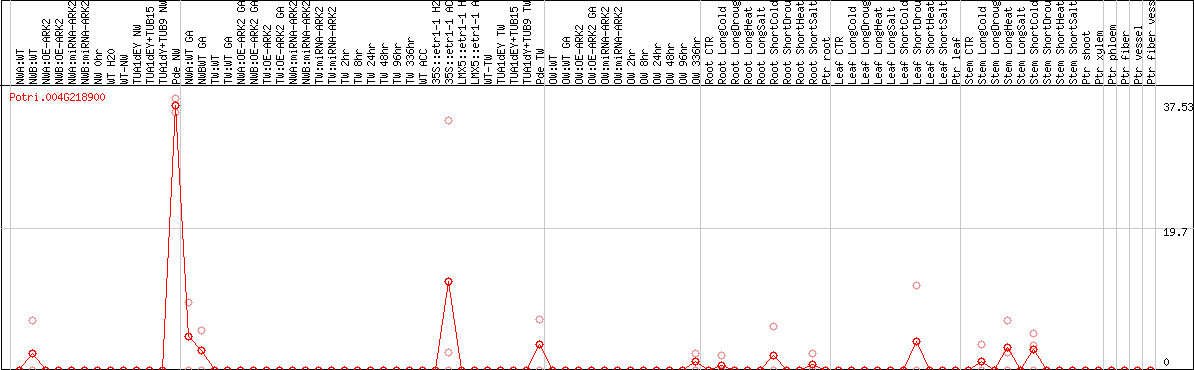
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G218900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.