Potri.004G228600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.004G228600.6 pacid=42795112 polypeptide=Potri.004G228600.6.p locus=Potri.004G228600 ID=Potri.004G228600.6.v4.1 annot-version=v4.1
ATGACTTCTAAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAAGAGGAAGAGAAAACGGCGATGAGGAGGAAGGCGAAGAGGAAGGAGAAGAAGAAGT
TGAAGAGAAAACAAGCGAGGAGAGAGATAGCGGAGAAGGAGAGGGAAGAAGAGGAGAGGAGACTCAACGATCCTGAGGAACAGAGGCGATTGAAGGAGAT
GGAGGAAGAGGAGAAGGAGAGAATGGAAAGGGAGAGGAAAGCGTTTGAGGAGAGAGAGAGGCAGTGGATGGAAGTTATGGAGAGGAAGAGGAAAGAAGAA
GAAGAAGAAGAAAGAAGGAAGGTTCTAGAAGAAAAGGAAGAGGAAGTTAATACACTACTTCAGGATGGAAAGGAGAATGGGTGTGATGAGGATGATGATT
GGGAATACATAGAAGAAGGGCCTGCGGAAATTATTTGGCAAGAAAATGAGATAATTGTTAGGAAGAAAAAGGTCAGGGTTCCAAAGAAGATTGCAGATCG
ACAGGGTACAAAAGAGGATGTTGATAGGCCCACATCAAATCCTCTTCCTCCACAATCTGAAGCTTTTACTGAGTATCAGAATGCGTCCGTTGTTTCAGCA
CAACAGATGCTTGAGAATGTCGCTCAACAAGTACCAAATTTTGGAACTGAACAGGATAAAGCTCATTGTCCATTCCATTTAAAAACCGGAGCTTGTCGGT
TTGGTCAACGCTGCAGCAGAGTTCATTTTTATCCTGATAAAGCATGCACATTGCTTATCAAGAACATGTATAATGGTCCTGGCCTTGCTTGGGAGCAAGA
TGAGGGGCTTGAGTATACAGATGAGGAGGTTGAATGCTCTTATGAAGAATTTTATGAGGATGTGCATACAGAGTTCTTGAAATATGGAGAAATAGTTAAT
TTTAAGGTCTGTAAAAATGGTTCATTCCACTTGCGGGGAAATGTATATGTACACTATAAGTCATTGGATTCAGCCATACTTGCTTATCATTCTATCAATG
GTCGCTACTTTGCCGGCAAACAGGTAAAATGTGAATTTATCAATTTGACCAGATGGAAGGTTGCCATATGTGGAGAGTTTATGAAGTCAAGGCTGAAGAC
ATGTTCTCATGGATCGGCTTGCAATTTTATTCATTGTTTCCGCAATCCTGGTGGGGACTATGAATGGGCTGATTGGGATAGACCACCACCAAGATTTTGG
GTGAAAAAAATGGCCTCTTTATTTGGCTACTCTGCTGAATCTTCGTATGAGAAGCAGACAGTGGAAGAAAAGCTGAGGCAAACAAGGATGTCTAGCAATA
TGTTAACAGCAGATAGAGACAGGCATGATCGAATGCGTAGATCTAGATCTAGAGACATGGATTGTGCAATCCATAGCTCTCATAAAAGTGGTGACATTGA
AGATGATTTTCCAGAAGTCACTCACTGGAGAAGCCATTCAAAAGATGATGGATCTAATTCTGATGGGCGTCAGTCAGATAGAGATACAGAAAGAGACAGG
GACAGGGACAGAGACAGAGACAGAGAAAGATATAGGCATCATTATCATGCAAGGAAAAGGTCAAAACACCATAGCAAAGTGTCAGAACGCTCAACTGATT
ACGGGGAGAACAATAGCAGGTCCCATGAGATTGATCAAAATGAGTTGTCAGATGATGGCAGAAACAGGCATGAGCATTTTGGTTCCACCAAAAGCTCTGG
AAGGAGGACATTTATAGATGATGATAAAGACAGCAAGGATGAGGCTCATCGAATGTATGGATACAGATTGGATAGGGACATAAATAGAGAAGAAAGCCGT
CATCGCACTGGGAAAAGCTCAAGACACCAGAGCAGAGTAAGGTCCCCTGATGATCATGGGGATGGTGAGAGTAGAACGCTTGATAGTGGTTCCAGCAGAG
AGGGGTCTCCTAGGGACAGAGACAGACACCATCATCACATAAGCAAAAGCTTAGGAAACCAGAAAAATGTCTTGGAATTCCCCGATAATCATTGGGATAA
TGGACCCAGAACTTATAGTACCAATTCCAGTAATGATCTGTTAGACAGGGACAGGGTAAGGCACTTTGGTCGTGGAAGGAAACGTTCAGAGCACAGTGAA
CCGGAGTTTTCAGATGATGAAGGTGCAATTGTGAAGAGGTTGAAAGACAAGTCTCTTGTTGGCAAATCTGACATAAGAAAATCTCACCTACAAAGAAAAC
CAACAGGTGGTCACTCAACCAGAAGAAATTATTCAAGCAAAGAGACTGTTGATGTGAATTCTCCTGTTGAGAGTCATAAACTAGACGGAAACACTGTTTA
TGGCCAAGTGAAACAATGTAGTGGAAGATGGGAATCAAACAGTCCTGGTTCAAGATACAACTTTTCTGAAGAAATTGACGAGCAGGATCGTTGGGAACCT
GAAAGCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.004G228600.6 pacid=42795112 polypeptide=Potri.004G228600.6.p locus=Potri.004G228600 ID=Potri.004G228600.6.v4.1 annot-version=v4.1
MTSKEEEEEEEEEEEEEEKTAMRRKAKRKEKKKLKRKQARREIAEKEREEEERRLNDPEEQRRLKEMEEEEKERMERERKAFEERERQWMEVMERKRKEE
EEEERRKVLEEKEEEVNTLLQDGKENGCDEDDDWEYIEEGPAEIIWQENEIIVRKKKVRVPKKIADRQGTKEDVDRPTSNPLPPQSEAFTEYQNASVVSA
QQMLENVAQQVPNFGTEQDKAHCPFHLKTGACRFGQRCSRVHFYPDKACTLLIKNMYNGPGLAWEQDEGLEYTDEEVECSYEEFYEDVHTEFLKYGEIVN
FKVCKNGSFHLRGNVYVHYKSLDSAILAYHSINGRYFAGKQVKCEFINLTRWKVAICGEFMKSRLKTCSHGSACNFIHCFRNPGGDYEWADWDRPPPRFW
VKKMASLFGYSAESSYEKQTVEEKLRQTRMSSNMLTADRDRHDRMRRSRSRDMDCAIHSSHKSGDIEDDFPEVTHWRSHSKDDGSNSDGRQSDRDTERDR
DRDRDRDRERYRHHYHARKRSKHHSKVSERSTDYGENNSRSHEIDQNELSDDGRNRHEHFGSTKSSGRRTFIDDDKDSKDEAHRMYGYRLDRDINREESR
HRTGKSSRHQSRVRSPDDHGDGESRTLDSGSSREGSPRDRDRHHHHISKSLGNQKNVLEFPDNHWDNGPRTYSTNSSNDLLDRDRVRHFGRGRKRSEHSE
PEFSDDEGAIVKRLKDKSLVGKSDIRKSHLQRKPTGGHSTRRNYSSKETVDVNSPVESHKLDGNTVYGQVKQCSGRWESNSPGSRYNFSEEIDEQDRWEP
ES
|
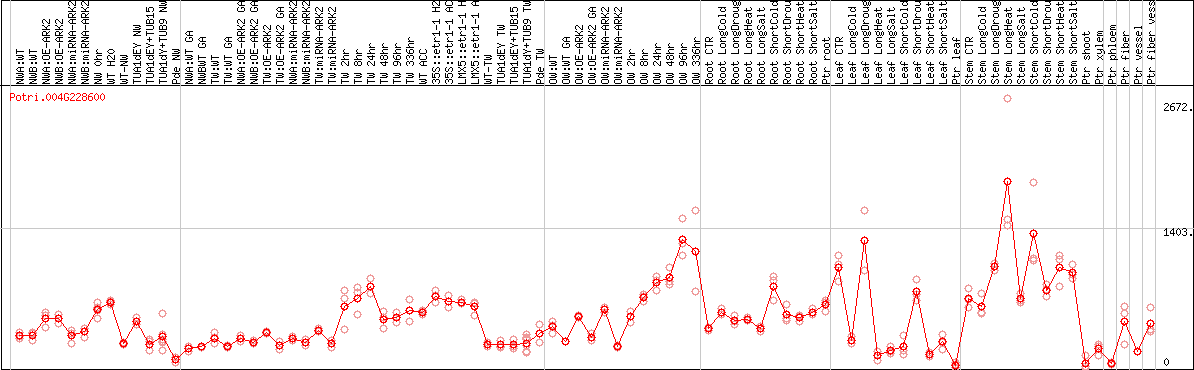
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.004G228600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
