External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G72930 114 / 9e-33
TIR
toll/interleukin-1 receptor-like (.1.2)
AT1G72920 115 / 8e-32
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT1G72910 117 / 1e-31
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT4G11170 117 / 9e-31
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G72890 114 / 3e-30
Disease resistance protein (TIR-NBS class) (.1), Disease resistance protein (TIR-NBS class) (.2)
AT1G72940 112 / 5e-30
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G17615 112 / 6e-30
Disease resistance protein (TIR-NBS class) (.1)
AT5G36930 112 / 4e-29
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT1G72900 108 / 2e-28
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G72950 107 / 7e-28
Disease resistance protein (TIR-NBS class) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G004000
251 / 7e-87
AT1G72930 107 / 6e-30
toll/interleukin-1 receptor-like (.1.2)
Potri.005G004100
186 / 2e-60
AT1G27170 122 / 3e-32
transmembrane receptors;ATP binding (.1.2)
Potri.004G230000
168 / 2e-52
AT5G36930 169 / 4e-47
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.006G269950
167 / 2e-52
AT5G36930 147 / 3e-40
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.005G003900
159 / 3e-49
AT5G36930 146 / 8e-40
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.005G004233
157 / 3e-48
AT1G27170 153 / 8e-42
transmembrane receptors;ATP binding (.1.2)
Potri.011G012000
166 / 7e-48
AT5G36930 489 / 2e-153
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T126306
165 / 1e-47
AT3G14470 585 / 0.0
NB-ARC domain-containing disease resistance protein (.1)
Potri.012G135700
165 / 1e-47
AT5G36930 640 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011104
129 / 4e-35
AT5G17680 574 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10018972
123 / 6e-33
AT5G36930 192 / 3e-51
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10011741
123 / 7e-33
AT5G36930 540 / 4e-172
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10005171
122 / 2e-32
AT1G27170 576 / 0.0
transmembrane receptors;ATP binding (.1.2)
Lus10039850
122 / 2e-32
AT5G36930 573 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10014582
116 / 5e-32
AT1G27170 186 / 9e-56
transmembrane receptors;ATP binding (.1.2)
Lus10018616
119 / 2e-31
AT5G36930 578 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10015453
118 / 2e-31
AT1G27170 282 / 5e-84
transmembrane receptors;ATP binding (.1.2)
Lus10006928
118 / 2e-31
AT5G36930 165 / 5e-43
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10006929
112 / 7e-31
AT1G72890 146 / 1e-39
Disease resistance protein (TIR-NBS class) (.1), Disease resistance protein (TIR-NBS class) (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF13676
TIR_2
TIR domain
Representative CDS sequence
>Potri.005G004366.1 pacid=42802750 polypeptide=Potri.005G004366.1.p locus=Potri.005G004366 ID=Potri.005G004366.1.v4.1 annot-version=v4.1
ATGGATTTGTTCTCACTTATCAATCTGTTGATGCTTCTCCTCTCCAGAAAGGATAACGATGTTTTCTTGAGTTTTAGCCACCACGACATTGGCAAGAATT
TTGGTGATCATCTCTACAAAGACCTCAATTCTGCTGGGATTCGCACTCTTAGAGATGATGGTGGAATCTACGCAGGACAAAAGTCTGATATCAAGAAAGC
GATACGGGAATCCAGGATATCGGTTGTTGTGTTCTCTAAAGGCTATGCTTCTTCAACAAAGTGCCTGGACCAGCTTGTGCACATCATGGATTCCAGGAAT
AAAACTGGGCTCCTTGTGCTTCCCGTTTTTTACAACGTTGATCCGTCGGAGGTCAGTGAACAGAAAGGGTTGTTTGAAGAAGCATTTGCTAAACACGAAA
AAAGCTTCCATAAGAAAATGGCTAGAGTGGAAAGTTGGAGGGCCGCTCTTAAAGAAGCCGCAGACCTGGCAGGAATGGTGCTGCAGCAAGACAGGTATAC
AAGTAATTGA
AA sequence
>Potri.005G004366.1 pacid=42802750 polypeptide=Potri.005G004366.1.p locus=Potri.005G004366 ID=Potri.005G004366.1.v4.1 annot-version=v4.1
MDLFSLINLLMLLLSRKDNDVFLSFSHHDIGKNFGDHLYKDLNSAGIRTLRDDGGIYAGQKSDIKKAIRESRISVVVFSKGYASSTKCLDQLVHIMDSRN
KTGLLVLPVFYNVDPSEVSEQKGLFEEAFAKHEKSFHKKMARVESWRAALKEAADLAGMVLQQDRYTSN
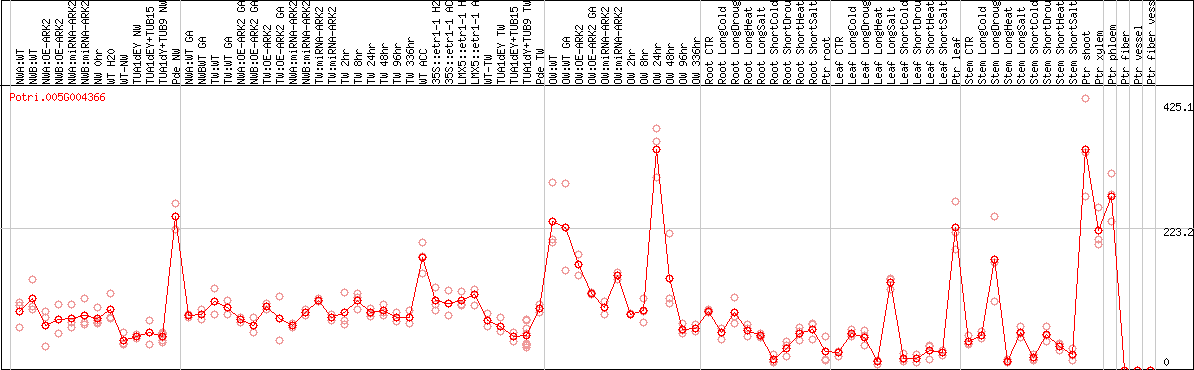
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G004366 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.