Potri.005G010133 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G010133.5 pacid=42804102 polypeptide=Potri.005G010133.5.p locus=Potri.005G010133 ID=Potri.005G010133.5.v4.1 annot-version=v4.1
ATGGGAAATGATACCAAGGTTAGTCGTAAAGGTAAGGATGAGGAAAGTAATGATATAAAAGGGAGAAACATTGGTAACAGATCATCAAGCAGTTTGGGTG
CTGCTAATGATACTTTTGGTCTAAGAAAGTCAGCTAGAGAAACATCATTCAAGAAAAACTTGACTCCGAGCCCTTCAAGTAGTAGAAAGTCTGAACGTAT
TGAGAAGCAGACCCCCCCTGCAATTCCTTCTGTTACGAGGAAATCTGAGAGATTGGTTGAGAAACAAAGTTTACCGACTCCTTCTAGAAGGCCTGAGAAG
GGTAAGAATCAGTCGTCATCTAGTTCTTCTGGTTCAACGAAGTGTGGTAAAAGCTCGGGCTCTTTAATTATGAAGGAAAAGCACAAGAAAGAGAAGAGTG
TGAAACAACTTGAAACTGAAGAAGTTGGAAACAATGACAAACCTGTTATCAAAACTGTCCTGGTTGGAATTAAGAGAATGGATGCTCGTGCTTACAGGGA
ACTGTTCAAACAGAAACAAAAGAAAGCTAAACTGGAAGGTAGCGAAGATCTGATAGTAAATAGGACACCTGGGGTTGATACTGAGGTGAAGAGTGGTTGC
AAAGTGATGTCTTCTAAACAAAAAAGAAGTATTGACGACTTGAATTTTGATGCTACGGAAATGGTTTCAAATGAAGATGGCGATGCAGCTCCATATGAGT
GCGGCAGAACTGATTCTGTGGATTCATGTTCCGAAAGGCAAAGGTTAAAGAAGCGCAATAAAGTGTCCAACAATAACACTGATTCGCCTTCTCTGAAAGC
TGCTTTAATAGGAACTTCTGGAGCTCCAGTGCATAAGATGTCTCAAGTCATGCCAAGTTCAGTTGGCCTTCTTAACACAACTGATGCGAATCATGTTTCC
AACTTCTCACAGTTGACTAGTAAATTATCACAAGTACTGAAAGCCGACATGGTTGGGTATAATGGAGGAAGGAACTTGCATGATGATTCTGAGAAGAGTC
TCCATCTTTTTTTGAAGCCTGAGATAGCAAAACTTTGTGAAATTTTGCAACTCCCAGAAAATGTCAAAGTCATGGTGGAACAGTTTCGGGAATATGTACT
GAATAATCACCATGTCAGTAGAGAACCACCTAGCTTATTGCAGGGATTTCTGATATCCTTGTGTTGGACTGCAGCTTCCATGCTAAAGCACAAACTTGAT
CACAAAGAATCACTTGCACTTGCAAAAGAGCATTTGAATTTCAGCTGCAAGAAAGACGAGGCAGATTTCGTTTACTCAAAGTTGCGATGTCTGAGGAAAT
TATTTCTTTATCGCACTGGGACTTGTAAGGTAGCTGGCTCTCCAAAAGCTTCAGGTTTTTCATTAGAAGATTTCGGTCAGAATCAGTCAAATGGAAGATC
TTCACTGTCAACACCATCTAACAAGCAAAAAGTGAGAATGGAGGTTGAAAACCTCAGATCTGGTCAGGAGTTCTCCATTAATCAGGTCCTTTCTCATCTG
GAATTGGCACAAAAAGATTACTCAAAAAGTATCAAAGACATTGAAAAGAAATGTGATAAACAGATGAGGAAGCTATTGCAGAGGCAACAAGAAGAAAGGG
AAGAGTTTGAAAAGAAGTATGAGCAAGACAAGGCAGAGCTCGAACATAAGCAAAGAACAGAGGCAGCTGTTATTCGTTTACACAGTAATAGTTCAGTGGA
TAAACTTAAGATGCTGGATAATGTGTATGCAAAAGAGTTCGAAAAACTTAAACGGCAGATGGATATGCGTCTTAATAATCTTTTGAAGTTGCAACTGGCT
ACAAGGAACAAGTTGCAAGAGAGGAAGGCTCAATGGATTGAAGGAGTGAAGTCCTGGGCACATGCTGAATTAATAAGTAAACCACCTGCAAATGAGTCTG
GATATGATCAAGAAAACACTGTCACTTTGAATTCTTGTTCCAGAGAACAGACTCCCAAACGAGTCCAAAGCATGCCAGATGGAGATGTTCCACTGGAAGT
TACTGAAACTGTGAGTTCAAATGAGGATGTACTTCCTGGGGTAATGGCTGCTTCCAAACCAATGTCCGATGGAGCTGCATCCAGCATGCTTGACCAAGAG
GTTCCCTTGGAAGTGCCTCAGACTGCAAGTGCGAGGGATGTTTCAGAGGATGTTGTGTCTGTAAATTCATCTCCATGTGAAGAACAAATTCCTGATTTGA
AGATTACATTGGGAATACCTGAGGCTAACAGCTGTAATGATGGTCCTGAGAACAGCATTCATAAATCATCATCTGAAGACGGTTCTGGTAGAGTTGCATT
AATGGTGCCTGATAGAGAATTTCCATTGGGAGTGACTGAGATTGTCAGTTCCACCGGTGGCATGGAGAATTCTGCTTTAAGTCCTTCACCTTCTGAAGGA
CAAACTTCTGCCAGAACCACGTCTTGCATTGATGGAAGAGAAGTTTTATTGGAAGTGCCTGAAACAGCCCCTCCGGAAGCTGAGGAGGCCGTTAACACAG
CTTTGGATAAAGATGGAGTAGCCAGTATGGAATTGGGTAATGCCATTGAAGTTGATAAGCAGAATGGTGCAGTCTGTATTCTTAATCAAGAGTCTCACCG
TGATGTAGCAGCAGTCAACCTGCAAAATGGAGAGTCTCTGTTAGAAGTGTCTGAAAACAATAGAGTCAACCAGTCGGATGAAGTGGTTCCATCAGGGGTT
TGCGAAACTCCAGTGGTAGGGAGTGGTACCACTGGTCAAGAGAAAAGTAGAGTTTGTGTTACCACCTTGGCTTGTGGTACTGGAGTTGATCAGCAGGCTG
GAGTGCTTCCATCAGGAGGTTTTGAAACTGCTACTGTTGCAGAAGTGGGGAGTGGCCCCACTTCGAGAGAGATTGACAGAATGCCTGCTGTAGCCTCAGA
TAGTAGTCAACCAACTGAACCCTTTAGATTGCAAGATAGAGCTGCACAATTTTGTGATAACTGGATTGCTTTTCAACAAAGTGATGCCTCAGCATCCCAA
CCTGTTGTAGTGTCCAATCAATCTCCAAATGATGCACCTGTCAGAGAGCATACGCTGCATTTGTTACCATCCATCGACTCCCCTACTAGTTCACAGCTTA
CAACTTCATTTGCGCAACATGTGCCTATTGATTTGATAGCTGTTGGTGGACCACAGACTCATATATCAAACATGAGGACAGAACCTGTTACTTCTAGAAT
TAGCAATCATTCTGCAACTGCACCTGCTGTTCGAATGCCTGTATCCACGTCTCAAGATCCACTGCAAAATGAATTGGATAGAATACGCACAGAAACAGAC
CAAATTATCAAGATCCACGAAGATACAAAGCTGCGGCTGAAATCTGATTGTGAGAAGGAGATACAGGAAGTTGTCGCTCAAATTCGCAGAACGTATGATT
TTAAACTTAAGGATTTAGAATATGAATTCCTTCGTAAGAAGAAAGAGATGGATGACAATCAAAGCAAAGTTCTCATGAATAAGATATTGGCTGAGGCATT
CAGGACCAAGTGCAAGGATAATAGGGCATCTAGGCAGCAAGAGATGACCTCTGGTGTTATGCAGCAGCTGCTTCAGCCATCCCAACCTTCTACACAGAGA
CCTTCTATTGTTACTGGCCCGTATTCAACTGGTCTTCCTGCAGTTAGTCTGCAGACAACTCCCACATCTTCCCTTCCTGCCCCACCTGTACAGGCTGTTC
ATTGCTCTGCACTTTTTTCAGCCACCCCAACCAGACCACCACATATCAGCTCCATCTCTCCTACCACAAGCAATCTCCAAGTTGGTACTGAGATTCGAGC
ACCTGCTCCACACCTGCAACATTTTAGACCTTCAGCATCCATGTCGGCAACTAGTCTTTCTGCCTTCCCAAGTGGGATCCTTCAACATATACCTACAACT
TCTCCCACACTTTCTGAGTTTCCTTCTCTAGCTCCAGCAACTGTGCAACAACCTGGTCCTCGGATTATGACAAATTTGCTCAAAAGCATGGGAGTTTTTC
CTTCTAGAACATCCTTCCCCAGACCAGAATCGTTAATGGATGTAGATAATCAAACGAGCACAGAGATTCCTTCCGTAGCTCCTGCAACTGTGCAACAATC
TGGTCCTCAGATTATGACAAATTTGCTTGAAAGCATGGGAATTTTTCCTTCTAGAACATCCCTCTCCAGACCAGAATCGTTATTGGATGTAGATAATCAA
ACAAGTACAGATGCAACTCAGCCCTGCAGTTTTCCCCCGCTAACAGATTTGAATTATAACATAAACCCATTGGTTCAAGAAAGTGAGGTTGTTTGTTTAT
CAGATGATGATTGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G010133.5 pacid=42804102 polypeptide=Potri.005G010133.5.p locus=Potri.005G010133 ID=Potri.005G010133.5.v4.1 annot-version=v4.1
MGNDTKVSRKGKDEESNDIKGRNIGNRSSSSLGAANDTFGLRKSARETSFKKNLTPSPSSSRKSERIEKQTPPAIPSVTRKSERLVEKQSLPTPSRRPEK
GKNQSSSSSSGSTKCGKSSGSLIMKEKHKKEKSVKQLETEEVGNNDKPVIKTVLVGIKRMDARAYRELFKQKQKKAKLEGSEDLIVNRTPGVDTEVKSGC
KVMSSKQKRSIDDLNFDATEMVSNEDGDAAPYECGRTDSVDSCSERQRLKKRNKVSNNNTDSPSLKAALIGTSGAPVHKMSQVMPSSVGLLNTTDANHVS
NFSQLTSKLSQVLKADMVGYNGGRNLHDDSEKSLHLFLKPEIAKLCEILQLPENVKVMVEQFREYVLNNHHVSREPPSLLQGFLISLCWTAASMLKHKLD
HKESLALAKEHLNFSCKKDEADFVYSKLRCLRKLFLYRTGTCKVAGSPKASGFSLEDFGQNQSNGRSSLSTPSNKQKVRMEVENLRSGQEFSINQVLSHL
ELAQKDYSKSIKDIEKKCDKQMRKLLQRQQEEREEFEKKYEQDKAELEHKQRTEAAVIRLHSNSSVDKLKMLDNVYAKEFEKLKRQMDMRLNNLLKLQLA
TRNKLQERKAQWIEGVKSWAHAELISKPPANESGYDQENTVTLNSCSREQTPKRVQSMPDGDVPLEVTETVSSNEDVLPGVMAASKPMSDGAASSMLDQE
VPLEVPQTASARDVSEDVVSVNSSPCEEQIPDLKITLGIPEANSCNDGPENSIHKSSSEDGSGRVALMVPDREFPLGVTEIVSSTGGMENSALSPSPSEG
QTSARTTSCIDGREVLLEVPETAPPEAEEAVNTALDKDGVASMELGNAIEVDKQNGAVCILNQESHRDVAAVNLQNGESLLEVSENNRVNQSDEVVPSGV
CETPVVGSGTTGQEKSRVCVTTLACGTGVDQQAGVLPSGGFETATVAEVGSGPTSREIDRMPAVASDSSQPTEPFRLQDRAAQFCDNWIAFQQSDASASQ
PVVVSNQSPNDAPVREHTLHLLPSIDSPTSSQLTTSFAQHVPIDLIAVGGPQTHISNMRTEPVTSRISNHSATAPAVRMPVSTSQDPLQNELDRIRTETD
QIIKIHEDTKLRLKSDCEKEIQEVVAQIRRTYDFKLKDLEYEFLRKKKEMDDNQSKVLMNKILAEAFRTKCKDNRASRQQEMTSGVMQQLLQPSQPSTQR
PSIVTGPYSTGLPAVSLQTTPTSSLPAPPVQAVHCSALFSATPTRPPHISSISPTTSNLQVGTEIRAPAPHLQHFRPSASMSATSLSAFPSGILQHIPTT
SPTLSEFPSLAPATVQQPGPRIMTNLLKSMGVFPSRTSFPRPESLMDVDNQTSTEIPSVAPATVQQSGPQIMTNLLESMGIFPSRTSLSRPESLLDVDNQ
TSTDATQPCSFPPLTDLNYNINPLVQESEVVCLSDDD
|
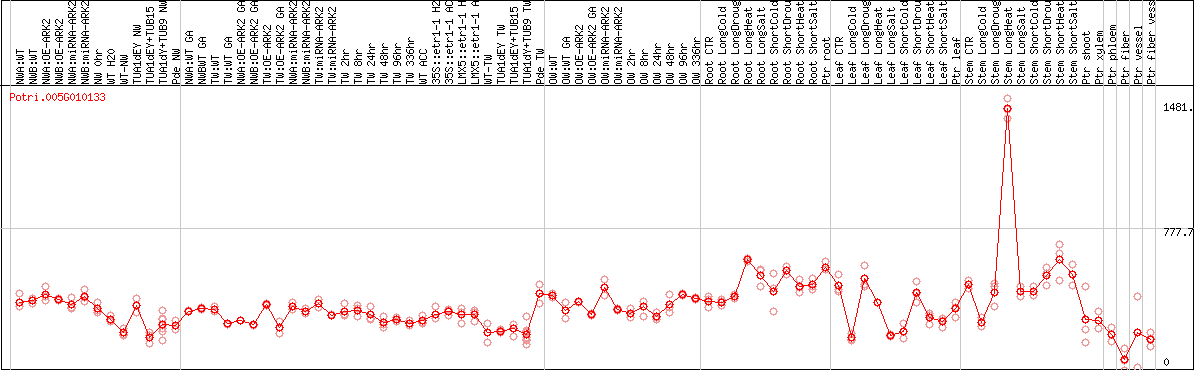
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G010133 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
