External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G09250 109 / 1e-29
bHLH
bHLH149, AIF4
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G17100 97 / 9e-25
bHLH
bHLH147, AIF3
sequence-specific DNA binding transcription factors (.1.2)
AT3G05800 92 / 8e-23
bHLH
bHLH150, AIF1
AtBS1(activation-tagged BRI1 suppressor 1)-interacting factor 1 (.1)
AT3G06590 91 / 1e-22
bHLH
bHLH148, AIF2
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT2G47270 44 / 5e-06
bHLH
bHLH151, UPB1
UPBEAT1, sequence-specific DNA binding transcription factors;transcription regulators (.1)
AT1G09530 44 / 5e-05
bHLH
PIF3, POC1, PAP3
PHOTOCURRENT 1, PHYTOCHROME-ASSOCIATED PROTEIN 3, phytochrome interacting factor 3 (.1.2)
AT2G43060 42 / 9e-05
bHLH
AtIBH1, bHLH158
ILI1 binding bHLH 1 (.1)
AT3G59060 42 / 0.0001
bHLH
PIF5, PIL6
PHYTOCHROME-INTERACTING FACTOR 5, phytochrome interacting factor 3-like 6 (.1.2.3.4)
AT1G66470 42 / 0.0001
bHLH
AtRHD6, RHD6, bHLH083
ROOT HAIR DEFECTIVE6 (.1)
AT2G43010 42 / 0.0002
bHLH
SRL2, PIF4, AtPIF4
phytochrome interacting factor 4 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G007700
222 / 6e-74
AT1G09250 95 / 4e-24
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.008G103500
77 / 5e-17
AT3G17100 174 / 2e-54
sequence-specific DNA binding transcription factors (.1.2)
Potri.010G147400
74 / 4e-16
AT3G17100 174 / 3e-54
sequence-specific DNA binding transcription factors (.1.2)
Potri.005G201500
47 / 7e-07
AT3G58850 82 / 4e-21
phy rapidly regulated 2 (.1)
Potri.002G193300
46 / 2e-06
AT2G47270 79 / 5e-20
UPBEAT1, sequence-specific DNA binding transcription factors;transcription regulators (.1)
Potri.002G060100
43 / 2e-05
AT3G58850 80 / 4e-20
phy rapidly regulated 2 (.1)
Potri.005G060900
42 / 8e-05
AT4G00120 151 / 7e-46
INDEHISCENT, EMBRYO SAC DEVELOPMENT ARREST 33, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.013G001300
42 / 0.0001
AT1G09530 224 / 6e-65
PHOTOCURRENT 1, PHYTOCHROME-ASSOCIATED PROTEIN 3, phytochrome interacting factor 3 (.1.2)
Potri.004G088900
42 / 0.0001
AT5G37800 199 / 7e-62
ARABIDOPSIS THALIANA RHD SIX-LIKE 1, RHD SIX-LIKE 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017065
74 / 4e-16
AT3G06590 171 / 1e-53
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10037783
74 / 5e-16
AT3G06590 172 / 9e-54
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10031425
59 / 2e-11
AT1G09250 51 / 7e-09
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10026730
41 / 0.0001
AT2G43060 97 / 4e-26
ILI1 binding bHLH 1 (.1)
Lus10017634
41 / 0.0003
AT1G66470 171 / 2e-51
ROOT HAIR DEFECTIVE6 (.1)
Lus10010907
40 / 0.0006
AT3G06590 81 / 8e-19
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10033592
40 / 0.0006
AT1G66470 174 / 5e-53
ROOT HAIR DEFECTIVE6 (.1)
Lus10033488
40 / 0.0006
AT4G00120 143 / 4e-43
INDEHISCENT, EMBRYO SAC DEVELOPMENT ARREST 33, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Representative CDS sequence
>Potri.005G012400.1 pacid=42802787 polypeptide=Potri.005G012400.1.p locus=Potri.005G012400 ID=Potri.005G012400.1.v4.1 annot-version=v4.1
ATGGCAAGCAACCAATTAAATTGCGACATGTCATCCCAACAAGAACAAAACCAACGAAGAAAGAAACGGAGAAAATTGACTCATGAAACGACAGAGAGTC
ACATTCAAAATGACAGCAAAAACCAGCTAACAGTGAGGTGGCGAACTGAAGCAGAGCGGCGGATCTATTCGTCGAAGCTTCTTGAAGCTCTCCGTCGGTC
AAGCCGGACATCTTCTCATCCTGGGAAGAGGACTCGAGAAGTCCGAGAAACCGCTAACAGAGTCCTCGCGGTGGCTGCTAGAGGAAGAACTCAGTGGAGC
CGCGCGATTCTCGCGAAACGCGCGCGGTTGTTGAGAGTGAGGAAGGTGAAGAAACAGAAATTAAACAGTGATCGTCGGTTACAGGAGAGAGAGAAGAGGA
GGAAGTTACCGGCGGTGGAGAAGAAAGTGAAGGTTTTGAGCCGGCTAGTACCTGGTTGCCGGAAAGTTTCGTTTGTGAATCTTTTAGAGGAAGCGAGTGA
TTATATTGCGGCTTTGGAGATGCAAATTAAGGTTATGACCACTCTAAGTGAAATTCTCACCGCCGCTGGCGGCGAAGTAGGCGGTGGCGCGGGAGTTGAT
GGCGGCTTGTCAAGTTGA
AA sequence
>Potri.005G012400.1 pacid=42802787 polypeptide=Potri.005G012400.1.p locus=Potri.005G012400 ID=Potri.005G012400.1.v4.1 annot-version=v4.1
MASNQLNCDMSSQQEQNQRRKKRRKLTHETTESHIQNDSKNQLTVRWRTEAERRIYSSKLLEALRRSSRTSSHPGKRTREVRETANRVLAVAARGRTQWS
RAILAKRARLLRVRKVKKQKLNSDRRLQEREKRRKLPAVEKKVKVLSRLVPGCRKVSFVNLLEEASDYIAALEMQIKVMTTLSEILTAAGGEVGGGAGVD
GGLSS
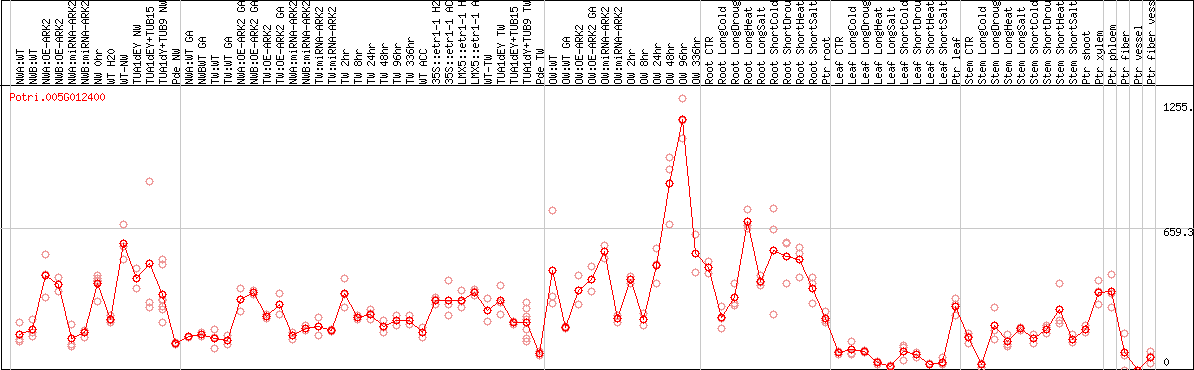
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G012400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.