CESA2.3 (Potri.005G027600) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | CESA2.3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G027600.2 pacid=42804254 polypeptide=Potri.005G027600.2.p locus=Potri.005G027600 ID=Potri.005G027600.2.v4.1 annot-version=v4.1
ATGGAAGTGAGTGCAGGCTTGGTGGCCGGCTCTCACAACAGGAATGAACTTGTTGTTATCCGTCGCGATGGAGAATTCGCTCCGAGATCGTTAGAGAGAG
TGAGTAGGCAGATATGCCACATATGTGGAGATGATGTAGGGCTAACGGTGGATGGAGAGCTATTCGTGGCCTGCAATGAGTGTGCCTTCCCCATCTGTAG
AACTTGTTATGAGTATGAGCGTAAAGAAGGAAATCAAGTCTGCCCTCAATGCAAGACCAGATTCAAGCGTCTCAAAGGATGTGCAAGAGTCCATGGGGAT
GATGAAGAAGATGGTACTGATGATTTAGAAAATGAGTTCAATTTCGATGGAAGGAATAGCAACAGACATGATATGCAGCACCATGGTGGACCTGAATCTA
TGCTTCACTATGATCCCGATTTGCCTCACGACCTTCATCATCCTTTGCCCCGAGTCCCTCTCCTTACTAATGGTCAAATGGTTGATGACATTCCTCCTGA
ACAACATGCTTTGGTCCCTTCATACATGGCTCCTGTCGGTGGTGACGGGAAAAGGATTCACCCCCTTCCCTTTTCAGATTCTTCTCTTCCTGCACAACCA
AGATCGTTGGATCCTTCCAAGGACTTGGCTGCTTATGGTTATGGAAGTATTGCTTGGAAGGAAAGGATGGAGAGCTGGAAGCAAAAACAGGACAAACTGC
AGATAATGAAGCGTGAAAATGGAGACTATGATGATGATGACCCCGATCTCCCATTAATGGATGAGGCAAGGCAACCACTATCGCGAAAGATGCCTATTCC
GTCCAGTCAAATCAATCCTTACCGAATGATCATCATTATTCGACTCGTAGTTCTTGGGTTCTTCTTCCATTATCGAGTCACACACCCTGTGAATGATGCG
TTTGCGTTGTGGCTCATATCTGTGATCTGTGAGATTTGGTTTGCTGTTTCATGGATACTTGATCAATTCCCAAAGTGGCTCCCTATTGACAGGGAAACTT
ATCTGGATAGATTATCTCTGAGGTATGAGAAGGAAGGCCAGCCTTCTCAGCTTTCTCCAGTTGACATATATGTAAGTACAGTTGATCCATTAAAAGAGCC
CCCTCTGGTGACTGCAAACACAGTTCTGTCCATTCTTGCAGTGGATTACCCTGTTGACAAGATTTCATGTTATGTTTCTGATGATGGTGCTGCTATGCTG
ACATTTGAGGCATTGTCAGAAACATCTGAATTTGCAAAGAAATGGGTCCCCTTCTGTAAGAAATTTAGCATTGAACCTCGGGCACCCGAGTTCTATTTTG
CTCAAAAGATAGATTATCTCAAAGATAAGGTTGATGCTTCATTTGTTAAGGAAAGAAGAGCAATGAAGAGAGAGTATGAAGAGTTTAAGGTTCGGGTCAA
CGCTTTGGTTGCCAAGGCACATAAGGTCCCGGAAGATGGATGGACAATGCAGGATGGTACTCCATGGCCTGGGAATAATGTTCGTGATCACCCTGGAATG
ATTCAGGTTTTCCTAGGCCAAAGTGGAGGGCATGACACAGATGGAAATGAATTACCACGTCTGGTATATGTTTCCAGAGAAAAGAGACCTGGATTTAATC
ACCACAAAAAGGCTGGGGCCATGAATGCTTTGGTCAGGGTCTCTGCCGTGCTATCAAATGCACGTTATCTTTTGAATTTGGATTGTGATCACTACATCAA
TAACAGTAAGGCTCTTAGAGAATCAATGTGTTTCATGATGGATCCATTGCTTGGAAAGAGAGTGTGCTATGTCCAGTTCCCACAGAGGTTTGATGGCATT
GACAGGAATGATCGATATGCAAACCGGAACACTGTATTCTTTGATATCAATATGAAAGGTTTGGATGGCATTCAAGGACCTATATATGTAGGGACTGGAT
GTGTTTTCAGAAGGCACGCGCTTTATGGTTATGATGCTCCAAAAACAAAGAAACCACCAACGAGGACATGCAACTGCTTGCCTAAGTGGTGCTGTGGATG
CTTTTGTTCAGGAAGGAAAAAGAAGAAGAAGACAAACAAACCTAAGTCCGAGTTAAAGAAGAGGAACTCTAGAACATTTGCACCTGTGGGTACTTTGGAG
GGTATTGAAGAGGGCATTGAAGGAATTGAAACCGAAAACATGGCTGTAACATCTGAGAAGAAATTGGAAAATAAGTTTGGACAATCCTCGGTGTTTGTAG
CATCTACCCTACTAGAGGATGGCGGCACACTGAAAAGTGCCAGCCCTGCATCCCTGCTCAAAGAAGCCATCCATGTCATTAGCTGTGGCTACGAAGATAA
AACAGAATGGGGCAAAGAGGTAGGGTGGATTTATGGTTCAGTTACGGAGGATATATTGACAGGCTTTAAAATGCATTGTCATGGATGGAGATCAATATAT
TGTATTCCTGCTAGGCCAGCATTTAAAGGTTCGGCGCCCATTAATCTTTCTGATCGTCTACACCAGGTCCTTCGATGGGCTCTTGGATCTGTTGAGATAT
TTTTGAGCAGGCACTGTCCTCTTTGGTATGGATATGGAGGGGGGTTGAAGTGGTTGGAGCGTCTGTCTTACATAAACGCTACTGTTTATCCTTTGACATC
TATTCCTCTACTGGCATATTGTACTCTTCCTGCAGTGTGCTTGCTTACCGGGAAATTTATCACTCCAGAGCTCAGCAATGCTGCCAGCTTGTGGTTTCTG
TCTCTTTTCATTTGTATTTTCGCAACAAGTATTCTGGAAATGAGATGGAGTGGAGTTGGTATCGATGAATGGTGGAGAAACGAACAATTTTGGGTGATTG
GAGGAGTATCAGCGCATCTATTTGCAGTATTCCAGGGGCTTTTAAAGGTTCTAGCTGGTGTTGATACGAACTTTACTGTGACGTCTAAAGGAGGAGATGA
CGATGAGTTCTCAGAGCTGTACGCATTTAAGTGGACCACGTTACTCATCCCACCAACAACATTGCTGATAATAAACCTGGTTGGAGTCGTGGCTGGTGTC
TCAAATGCTATAAACAATGGCTACGAGTCATGGGGGCCTTTGTTTGGTAAGCTTTTCTTTGCATTCTGGGTGATAGTTCACCTGTACCCTTTCCTGAAAG
GTTTACTCGGCCGACAGAACAGAACTCCCACAATCATTATTGTCTGGTCAATTTTGCTTGCTTCAATCTTCTCTCTTTTGTGGGTTCGGATCGATCCATT
CTTGGCCAAGTCCAATGGCCCACTTCTGGAGGAATGTGGATTGGACTGTAATTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G027600.2 pacid=42804254 polypeptide=Potri.005G027600.2.p locus=Potri.005G027600 ID=Potri.005G027600.2.v4.1 annot-version=v4.1
MEVSAGLVAGSHNRNELVVIRRDGEFAPRSLERVSRQICHICGDDVGLTVDGELFVACNECAFPICRTCYEYERKEGNQVCPQCKTRFKRLKGCARVHGD
DEEDGTDDLENEFNFDGRNSNRHDMQHHGGPESMLHYDPDLPHDLHHPLPRVPLLTNGQMVDDIPPEQHALVPSYMAPVGGDGKRIHPLPFSDSSLPAQP
RSLDPSKDLAAYGYGSIAWKERMESWKQKQDKLQIMKRENGDYDDDDPDLPLMDEARQPLSRKMPIPSSQINPYRMIIIIRLVVLGFFFHYRVTHPVNDA
FALWLISVICEIWFAVSWILDQFPKWLPIDRETYLDRLSLRYEKEGQPSQLSPVDIYVSTVDPLKEPPLVTANTVLSILAVDYPVDKISCYVSDDGAAML
TFEALSETSEFAKKWVPFCKKFSIEPRAPEFYFAQKIDYLKDKVDASFVKERRAMKREYEEFKVRVNALVAKAHKVPEDGWTMQDGTPWPGNNVRDHPGM
IQVFLGQSGGHDTDGNELPRLVYVSREKRPGFNHHKKAGAMNALVRVSAVLSNARYLLNLDCDHYINNSKALRESMCFMMDPLLGKRVCYVQFPQRFDGI
DRNDRYANRNTVFFDINMKGLDGIQGPIYVGTGCVFRRHALYGYDAPKTKKPPTRTCNCLPKWCCGCFCSGRKKKKKTNKPKSELKKRNSRTFAPVGTLE
GIEEGIEGIETENMAVTSEKKLENKFGQSSVFVASTLLEDGGTLKSASPASLLKEAIHVISCGYEDKTEWGKEVGWIYGSVTEDILTGFKMHCHGWRSIY
CIPARPAFKGSAPINLSDRLHQVLRWALGSVEIFLSRHCPLWYGYGGGLKWLERLSYINATVYPLTSIPLLAYCTLPAVCLLTGKFITPELSNAASLWFL
SLFICIFATSILEMRWSGVGIDEWWRNEQFWVIGGVSAHLFAVFQGLLKVLAGVDTNFTVTSKGGDDDEFSELYAFKWTTLLIPPTTLLIINLVGVVAGV
SNAINNGYESWGPLFGKLFFAFWVIVHLYPFLKGLLGRQNRTPTIIIVWSILLASIFSLLWVRIDPFLAKSNGPLLEECGLDCN
|
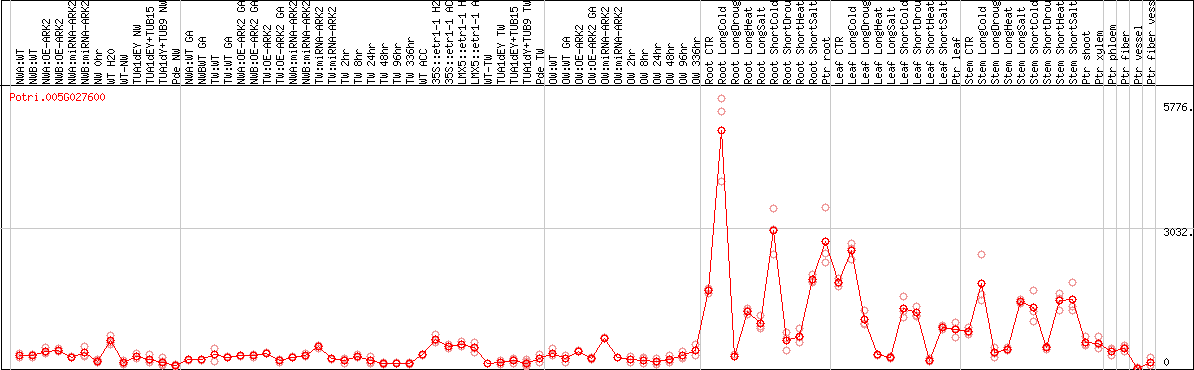
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G027600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
