External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G12700 147 / 7e-41
RPF1
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
AT1G12620 139 / 2e-38
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G12300 136 / 2e-37
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G62930 135 / 4e-37
RPF3
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63080 135 / 5e-37
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G63320 124 / 1e-35
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G62670 128 / 2e-34
RPF2
rna processing factor 2 (.1)
AT3G22470 128 / 3e-34
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G62910 124 / 4e-33
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G63130 122 / 2e-32
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G050240
357 / 9e-122
AT1G12700 465 / 1e-155
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G050180
339 / 1e-114
AT1G12700 488 / 2e-164
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G046200
325 / 5e-109
AT1G12700 501 / 5e-170
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G050400
324 / 8e-109
AT1G12700 504 / 3e-171
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G050500
322 / 6e-108
AT1G12700 523 / 2e-178
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G045000
312 / 3e-104
AT1G12700 502 / 4e-170
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G050300
305 / 2e-101
AT1G12700 466 / 5e-156
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G046100
298 / 2e-98
AT1G12700 512 / 1e-173
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G038400
293 / 8e-97
AT1G12700 478 / 7e-161
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014244
157 / 5e-45
AT1G12700 397 / 7e-130
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10003433
156 / 1e-44
AT1G12700 396 / 2e-129
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10014242
152 / 2e-44
AT1G62930 256 / 5e-78
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014245
154 / 5e-44
AT1G12700 427 / 5e-141
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10021074
150 / 6e-44
AT1G62930 273 / 5e-86
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10022861
150 / 2e-42
AT1G12700 370 / 7e-121
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10003446
142 / 2e-42
AT1G62930 137 / 2e-37
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014247
149 / 5e-42
AT1G62930 340 / 3e-109
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10008593
144 / 3e-40
AT1G12700 400 / 7e-131
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10029737
144 / 5e-40
AT1G63130 424 / 6e-140
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF13041
PPR_2
PPR repeat family
Representative CDS sequence
>Potri.005G050060.1 pacid=42802348 polypeptide=Potri.005G050060.1.p locus=Potri.005G050060 ID=Potri.005G050060.1.v4.1 annot-version=v4.1
ATGCAAGGTCTGTGCCAATTAGGAAGACCTAAGGAAGCTCTCAATCTTTTCAAGGAGATGTGTTCTTATGGCCCGCATCCAAATTTGGTGACTTATGTGA
TTTTACTAGATGGCTTCTGCAAGCATGGGCATTTGGATGAGGCATTAAAACTGCTCAAGTCAATGAAAGAGAAGAAACTAGAACCTAATATTGTCCATTA
TACTATCCTTATTGAAGGCATGTTTATCGCTGGGAAGCTTGAAGTTGCGAAGGAACTATTTTCTAAGCTTTTTGGGGATGGAACACGGCCTGATATACGG
ACATACACTGTCATGATCAAGGGACTGCTGAAAGAAGGGCTGTCAGATGAAGCATACGATTTATTTAGAAAAATGGAAGATGATGGCTTCTTGCCAAACA
GTTGCTCATATAATGTTATGATTCAAGGATTTCTTCAAAATCAGGACTCATCAACTGCTATACGACTTATTGATGAAATGGTTGGTAAAAGATTCTCGGT
GAATTTATCTACATTTCAGATGTTATTGGATTTAGAATCTCAAGATGAAATCATAAGCCAATTTATGCGTGGAAGCTCTCAAGGTAGAAAAATGAAGTGA
AA sequence
>Potri.005G050060.1 pacid=42802348 polypeptide=Potri.005G050060.1.p locus=Potri.005G050060 ID=Potri.005G050060.1.v4.1 annot-version=v4.1
MQGLCQLGRPKEALNLFKEMCSYGPHPNLVTYVILLDGFCKHGHLDEALKLLKSMKEKKLEPNIVHYTILIEGMFIAGKLEVAKELFSKLFGDGTRPDIR
TYTVMIKGLLKEGLSDEAYDLFRKMEDDGFLPNSCSYNVMIQGFLQNQDSSTAIRLIDEMVGKRFSVNLSTFQMLLDLESQDEIISQFMRGSSQGRKMK
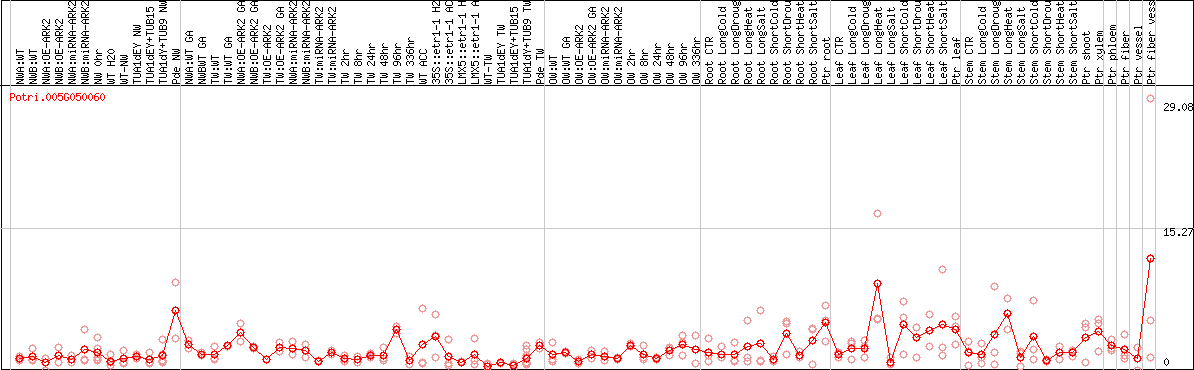
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G050060 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.