External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G02590 228 / 1e-73
bHLH
bHLH059, UNE12
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT1G03040 216 / 3e-68
bHLH
bHLH007
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT4G30980 179 / 6e-54
bHLH
LRL2, bHLH069
LJRHL1-like 2 (.1)
AT2G24260 179 / 2e-53
bHLH
LRL1, bHLH066
LJRHL1-like 1 (.1)
AT5G58010 172 / 2e-51
bHLH
LRL3, bHLH082
LJRHL1-like 3 (.1)
AT5G48560 96 / 8e-22
bHLH
bHLH078
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G50915 92 / 2e-21
bHLH
bHLH137
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT3G07340 92 / 1e-20
bHLH
bHLH062
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G34530 91 / 1e-20
bHLH
bHLH063, CIB1
cryptochrome-interacting basic-helix-loop-helix 1 (.1)
AT1G68920 92 / 2e-20
bHLH
bHLH049, ACE1
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G041000
467 / 2e-166
AT4G02590 207 / 2e-65
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.005G217800
252 / 2e-82
AT4G02590 307 / 2e-104
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.002G045400
250 / 1e-81
AT4G02590 295 / 8e-100
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.006G186600
177 / 6e-52
AT2G24260 198 / 2e-59
LJRHL1-like 1 (.1)
Potri.018G109500
177 / 8e-52
AT2G24260 194 / 5e-58
LJRHL1-like 1 (.1)
Potri.006G135600
158 / 1e-44
AT5G58010 189 / 1e-56
LJRHL1-like 3 (.1)
Potri.002G235400
98 / 2e-22
AT3G07340 257 / 2e-79
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G248500
98 / 3e-22
AT3G07340 349 / 5e-115
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G148900
97 / 3e-22
AT3G07340 254 / 2e-78
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014726
263 / 1e-88
AT4G02590 278 / 8e-96
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10006587
271 / 2e-88
AT4G02590 245 / 1e-79
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10015909
269 / 2e-88
AT4G02590 253 / 2e-82
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10002720
266 / 6e-88
AT4G02590 351 / 8e-122
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10024850
247 / 1e-80
AT4G02590 343 / 1e-118
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10018761
227 / 1e-72
AT4G02590 318 / 2e-108
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Lus10021846
167 / 2e-48
AT2G24260 275 / 3e-90
LJRHL1-like 1 (.1)
Lus10036286
140 / 4e-39
AT2G24260 214 / 2e-67
LJRHL1-like 1 (.1)
Lus10023562
92 / 3e-21
AT1G26260 178 / 9e-54
cryptochrome-interacting basic-helix-loop-helix 5 (.1.2.3)
Lus10040448
92 / 4e-21
AT4G34530 177 / 4e-53
cryptochrome-interacting basic-helix-loop-helix 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.005G053500.1 pacid=42804456 polypeptide=Potri.005G053500.1.p locus=Potri.005G053500 ID=Potri.005G053500.1.v4.1 annot-version=v4.1
ATGGCAGGAAATCCCCCACCTGAAGGGTTAGGAGATGATTTCTTTGAGCAGATCTTGGCAGGGCAGCCACCTGGTTATGGTGGTGAGGCTGTGGGGTCCA
CCAGCCTGCCCATGATGGGGTTGCAGTTAGGGTCTAGTGCTGCTAATGTTGGATTAACGAGGAGTAGTGCTAATAATAATATGGGAATGATGCCTTTAGG
GTTAAACTTGGAGCATCATGGGTTTTTAAGGCAACAACAAGATGATGGCAGTAGTTCTTTAGATACCAACAACAGCAATAATATCAATAATGCTTCTCCT
TTTTCTATCACTTCTGCTGGATTCTTGGGGAGAGATTCTGTACACATGACTAGCTTGTTCCCAACGTTTGGACAATTGCAGATTCATTCATCAATCCGGT
CTGCACCACCACCACTTGGACCACCTCAGATTCACCAGTTTAATAGCCAGCCAACATCAGGAGCAGTTTCAGCTGTACCACAACCACCTGGCATTCGACC
AAGGGTGCGAGCAAGACGGGGCCAAGCTACAGATCCGCACAGTATTGCTGAGAGGTTGCGAAGAGTAAGAATCACGGAGAGGGTGAAGGCGTTGCAGGAA
TTGGTTCCCACTTGCAACAAGACAGATAGGGCAGCCATGCTGGATGAAATTGTGGATTATGTGAAGTTTCTTAGACTCCAAGTCAAGGTTTTGAGTATGA
GCAGACTCGGTGCGGCGGGCGCTGTGGCTCAGCTTGTAGCTGATGTACCACTATCATCAGTTCAGGGAGAAGGCATTGAAGGAGGGGCCAATCAGCAAGC
TTGGGAGAATTGGTCAAACGATGGCACTGAACAGGAAGTAGCTAAGTTGATGGAAGAAGATGTTGGAGCTGCCATGCAGCTCCTTCAGTCCAAGGCACTC
TGCATCATGCCTGTATCACTTGCTTCAGCAATCTTCCGAGCACGCCCACCGAATGCTCCAACACTGGTCAAGCCCGAATCGAACCCCCCCTCATAA
AA sequence
>Potri.005G053500.1 pacid=42804456 polypeptide=Potri.005G053500.1.p locus=Potri.005G053500 ID=Potri.005G053500.1.v4.1 annot-version=v4.1
MAGNPPPEGLGDDFFEQILAGQPPGYGGEAVGSTSLPMMGLQLGSSAANVGLTRSSANNNMGMMPLGLNLEHHGFLRQQQDDGSSSLDTNNSNNINNASP
FSITSAGFLGRDSVHMTSLFPTFGQLQIHSSIRSAPPPLGPPQIHQFNSQPTSGAVSAVPQPPGIRPRVRARRGQATDPHSIAERLRRVRITERVKALQE
LVPTCNKTDRAAMLDEIVDYVKFLRLQVKVLSMSRLGAAGAVAQLVADVPLSSVQGEGIEGGANQQAWENWSNDGTEQEVAKLMEEDVGAAMQLLQSKAL
CIMPVSLASAIFRARPPNAPTLVKPESNPPS
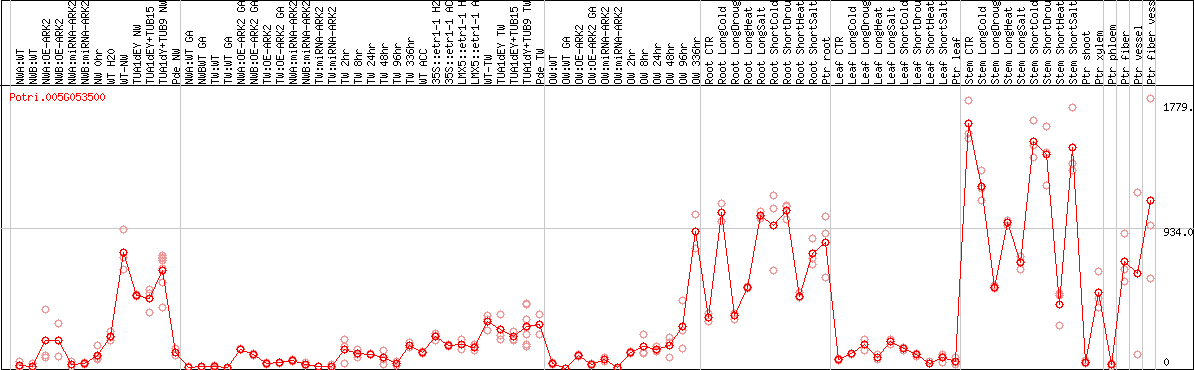
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G053500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.