External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G72030 216 / 1e-69
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
AT2G39000 47 / 6e-06
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
AT4G28030 44 / 8e-05
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
AT1G24040 40 / 0.001
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G095800
50 / 5e-07
AT1G24040 287 / 4e-96
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.008G146000
49 / 1e-06
AT1G24040 317 / 9e-108
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.010G222900
46 / 1e-05
AT2G39000 370 / 5e-130
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Potri.006G176500
45 / 2e-05
AT4G28030 284 / 2e-96
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.008G039300
45 / 2e-05
AT2G39000 387 / 1e-135
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042534
258 / 1e-85
AT1G72030 179 / 3e-55
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Lus10021995
255 / 1e-84
AT1G72030 181 / 4e-56
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Lus10030837
49 / 2e-06
AT1G24040 289 / 2e-96
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Lus10019975
49 / 2e-06
AT4G28030 322 / 3e-111
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Lus10023517
49 / 2e-06
AT2G39000 388 / 4e-137
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10040401
48 / 4e-06
AT2G39000 386 / 3e-136
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10037996
46 / 1e-05
AT2G39000 335 / 2e-115
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10009229
45 / 3e-05
AT2G39000 320 / 5e-112
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0257
Acetyltrans
PF00583
Acetyltransf_1
Acetyltransferase (GNAT) family
Representative CDS sequence
>Potri.005G054600.1 pacid=42803146 polypeptide=Potri.005G054600.1.p locus=Potri.005G054600 ID=Potri.005G054600.1.v4.1 annot-version=v4.1
ATGGCTAATTTGCGCCTTTGCTATTTGCAGATTTCAAACAGTCTATGTACTGAACCTTGGTATGCCGGTAGAGGGTCCAGGTTTTATCATGGAAGTTGGA
AAACTGTAAGCAACAGAAAGAAAGTCAGCCAACCGATCGAAAGACCTGTGGGCTGGAGTGGTAGTTGTATTGTGCAATGCTGTAGTACATCTTCAACTAC
ATCAGCTGCGGAAGCAACTGAGCAAGTGTTTCTTGAAGATAGGAGTAGTATTGAAGAGAAGGAGGAGCAGTTTGAGTATTTGGCGAGTGAGTTTGGATGG
AAAGTAAGGAGGTTAGCTGAGAATCGAGATGAGATGAGGGAAGTAGCTCAAATCCAAGCTGAAGCATTTCACACACCAGTGGCTCTCTTTGATGACCTTT
TCTTCGAGTTCTTCAAGGCTGAAGTGCTCTCTGGGCTTCTTTACAAGCTTAAGAACTCACCTCCAGACAGATATGCATGTTTGGTCGCTGAGCCTGCAGC
TGAAAGGAAACTGGTGGGCATTGTGGATGTCACAGCTTTAAGAGATAAAGATGTCCTACAGCACCTTGAGGGAGCGGATGAATACCTTTATATTTCAGGG
ATTGCTGTTTCCAAAAGTTTCAGGCGGCGGAAAATAGGAAGTGTCTTGCTAAAAGCCTGTGATGTACTCTCCCATCTATGGGGTTTTGAATGTCTTGCCC
TCAGGGCATATGAAGATGATATGGGTGCTCGAAAGCTATACACAAATGCAGGTTACAAAGTGGTATCAAGCGATCCCCAATGGGTGACTTGGATAGGAAG
AAAACGCCGTGTTATTATGATCAAACGGTCCAATATGCTTGATTAG
AA sequence
>Potri.005G054600.1 pacid=42803146 polypeptide=Potri.005G054600.1.p locus=Potri.005G054600 ID=Potri.005G054600.1.v4.1 annot-version=v4.1
MANLRLCYLQISNSLCTEPWYAGRGSRFYHGSWKTVSNRKKVSQPIERPVGWSGSCIVQCCSTSSTTSAAEATEQVFLEDRSSIEEKEEQFEYLASEFGW
KVRRLAENRDEMREVAQIQAEAFHTPVALFDDLFFEFFKAEVLSGLLYKLKNSPPDRYACLVAEPAAERKLVGIVDVTALRDKDVLQHLEGADEYLYISG
IAVSKSFRRRKIGSVLLKACDVLSHLWGFECLALRAYEDDMGARKLYTNAGYKVVSSDPQWVTWIGRKRRVIMIKRSNMLD
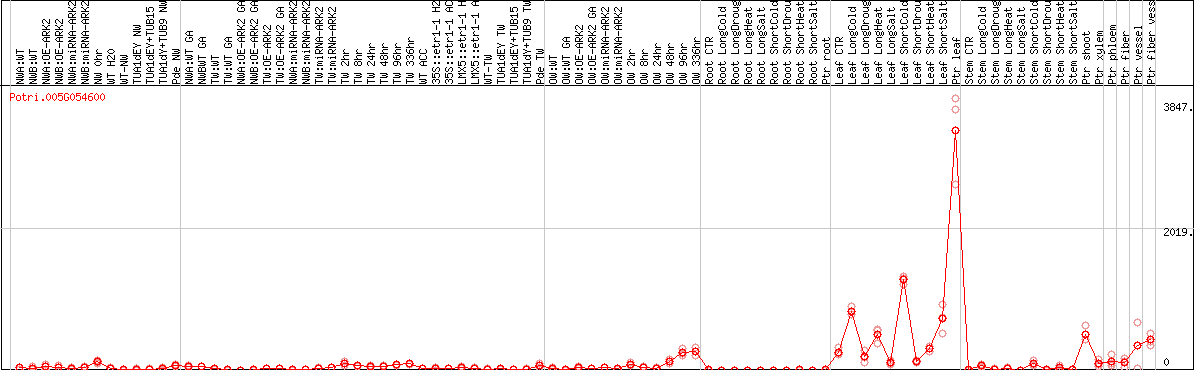
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G054600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.