Potri.005G058300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G058300.1 pacid=42803680 polypeptide=Potri.005G058300.1.p locus=Potri.005G058300 ID=Potri.005G058300.1.v4.1 annot-version=v4.1
ATGTCGAACCTGGAACCATCTGGGTCACAGATGCTGAGCCGGAGACCGTCTCGCAGCGCTGCCACCACTACATTCTCTCTTGAGGTTTTTGATAATGAGG
TCGTCCCTTCCTCACTCGGTTCCATCGCCCCTGTTCTCCGCATTGCAGCTGAAATCGAACATGAACGCCCTCGCGTTGCCTATCTCTGTAGGTTTTATGC
GTTTGAGAAAGCCCATAGACTGGATCCAAATTCAAGTGGTCGTGGTGTGAGGCAGTTCAAGACATCTTTGCTGCAGAGATTGGAGAGGGACAATAATTCC
AGTCTTGCCTCTCGGGTTAAAAAGACCGATGCGCGGGAAATCGAGAGCTTTTATCAGCAGTACTATGAACACTACGTTAGAGCACTTGACCAGGGAGAGC
AAGCAGACCGAGCTCAACTTGGAAAAGCCTATCAGACAGCTGGTGTTCTTTTTGAAGTGCTTTGTGCTGTTAACAAAACAGAGAAAGTAGAAGAAGTTGC
TCCTGAGATTATTGCAGCAGCCAGAGATGTCCAGGAAAAGAAAGAAATATATGCCCCTTTCAACATCCTTCCTCTGGATTCTGCCGGGGCTTCACAGTCC
ATAATGCAGCTTGAGGAGGTCAAAGCTGCTGTTGCTGCACTGTGGAACACTCGTGGTTTGAACTGGCCCACTGCATTTGATCCACAAAGGCAGAAAGCTG
GTGATTTAGACATTCTTGATTGGCTGAGGGCCATGTTTGGCTTTCAGAGAGACAATGTCAGGAACCAGAGGGAGCATTTGATTTTACTACTTGCCAACAA
ACATATAAGGCTTAATCCCAAGCCTGAACCTATTAGCAAGCTTGATGATCGAGCTGTTGATGAAGTTATGAACAAGCTTTTTAAGAATTATAAAACATGG
TGCAAGTTTTTAGGGCGGAAACATAGTTTAAGGCTCCCTCAAGGCCAGCCAGAAATCCAGCAGAGAAAGATACTGTACATGGGCCTTTTTCTTCTCATCT
GGGGCGAGGCAGCAAATGTTCGCTTCATGCCTGAATGTCTATGCTATATTTTTCATAATATGGCATATGAACTCCATGGCCTGTTGGCTGGAAACGTCAG
CATCGTTACTGGAGAAAATATTAAGCCTTCTTATGGTGGAGACGACGAGGCTTTCCTGAGGAAGGTTATAACACCAATTTACCATGTAATTGAAAAGGAA
GCCAATAAAAGCAAAAATGGTAAAGCTTCACACTCACAGTGGTGCAATTATGATGATCTAAATGAATACTTCTGGTCATCGGACTGCTTTTCTCTTGGTT
GGCCTATGCGTGATGATGGAAGTTTTTTTACATCTACAAGAGATGTGGGGAAAAAGGCTTCTTCAGAAAAACCCAGAAGCACAGGCAAAGCATATTTTGT
AGAGACACGAACCTTTTGGCACATTTTTCGAAGTTTTGACAGATTGTGGACCTTTTATATTCTTGCTCTTCAGGCAATGATCATCATTGCTTGGAGTGGG
GTCTCGATTTTGAACATTGTTCAAAAGGATGTATTATATCAGCTATCCAGCATTTTCATCACAGCAGCCTGTCTTCGCTTGCTCCAAAGTATTCTGGACC
TTGTTCTGAACTTCCCAGGGTTCCACAAGTGGAAATTCACTGATGTCCTGAGAAATGTGCTCAAGATCATTGTTAGTCTTGCATGGGCAATCATTCTTCC
ACTATGTTATGTGCATTCCTTCAAAGTGGCCCCTGATAAAATCAAAGATTTGCTGTCATTCTTTAAAGAAGTGAAGGACATACCTGCACTATACCTTTTG
GCAGTGGCTGTATACATGCTTCCAAATATATTGGCAGCTGCTTTGTTTATCTTTCCCATGCTGCGAAGATGGATTGAAAACTCAGACTGGCTGATTATCA
GATTTCTCTTATGGTGGTCACAGCCAAGAATTTATGTTGGCAGGGGAATGCATGAGAGTCAATTTGTACTCATAAAGTATACTGTTTTCTGGCTGTTACT
TTTATGTTCAAAAATAGCATTCAGCTATTTTGTCCAGATAAAACCTCTGGTGAAGCCAACGAAAGCTATAATGAACATTCGGAATGTAGATTATGAGTGG
CACGAATTTTTTCCAAATGCTAAAAACAATTATGGAGCTGTTTTGTCACTCTGGTTACCAGTAATCTTGGTCTATTTTATGGATACTCAAATTTGGTATT
CTATTTTCTCAACCATATATGGTGGCTTTGCCGGGGCCTTTGATCGCCTAGGAGAGATTCGAACTCTGGGCATGCTGAGGTCACGGTTCCAGTCCTTACC
CGGTGCATTTAACACATATTTGGTTCCTTCAGATAAGAAACGGAAGAAGGGCTTCTCTTTTTCAAAGAGATTTTCTGAGGTTACTGCAAGCAAGAGAAGT
GAAGCTGCCAAATTTGCTCAGTTATGGAATGAAGTGATATGTAGTTTCCGTGAAGAAGATCTCATTAGTGACAGGGAGATGGATCTGTTGCTGGTACCAT
ATACATCAGATCCTAGCCTCAAATTAATTCAGTGGCCACCGATTATGCTTGCAAGCAAGATCCCGATAGCTTTGGATATGGCAGTGCAATTCCGATCAAG
GGATGCGGACCTCTGGAAACGCATATGTGCAGATGAATACATGAAATGTGCAGTCATTGAATGCTACGAATCCTTTAAACATGTCCTGAATATTTTGGTA
GTTGGAGAAATTGAGAAAAGGATACTTAGCATCATCTTTAAAGAAGTTGAGAGTAATATTTCAAAGAACACACTTCTTACAAATTTCAGAATGGGTCCCT
TACCAGCATTATGTAACAAATTTGTGGAGCTCGTGATATTATTGAAAGATGCTGATCCCTCCAAGCAAAATACCGTGGTGTTGATACTGCAGGACATGTT
AGAAGTATTTACTAATGATATGATGGTGAATGAGAATCGTGAATTGGTAGACCTTGGGCAAAGTGGCAAGGATTCTGGAAGACAAGTGTTTTCTGGTACT
GACACAAAACCTGCTATAATGTTCCCTCCTGTAGTTACAGCACAATGGGAAGAACAGATAAGACGCATTCATCTCCTTTTGACGGTGAACGAATTTGCCA
ATGATGTCCCAACAAACCTTGAAGCACGTAGGAGGATTTCATTCTTTACAAATTCGTTGTTTATGGACATGCCACGTCCTCCTCGAGTCCGTAAAATGCT
CTCATTCAGTGTCTTAACCCCCTACTATAGTGAAGAGACAGTTTATTCCAAATCTGATCTTGAAATGGAGAATGAAGATGGTGTATCAATCATCTACTAC
TTGCAAAAGATTTACCCAGATGAATGGAATAACTTCATGGAAAGAATAAATTGCAAAAAAGAGAGTGAGGTTTGGGAAAATGAAGAAAATATTTTGCAAC
TTCGTCATTGGGGCTCCTTGAGAGGTCAAACTCTCTGTCGCACAGTTAGAGGGATGATGTACTATAGACGAGCTTTAAGACTTCAAGCTTTCCTTGATAT
GGCTAAAGAAAGTGAGATACTAGAAGGTTACAAAGCCATCACAGATCCAACTGAGGAAGATAAGAAAAGTCAAAGATCTGTGAGTGCTCAAATAGAGGCA
GTAGCGGACATGAAATTCACTTATGTTGCTACCTGTCAGAACTATGGAAATCAGAAGAGGAGTGGAGATCGTCGTGCAACAGACATTCTCAATTTAATGG
TCAATAATCCCTCTCTCCGTGTTGCATATATTGACGAAGTTGAAGAAAGAGAAAGGGAAGGTGGCAAGGTACAGAAGGTTTATTATTCTGTATTGGTAAA
AGCTGTTGATAATCTTGATCAGGAAATATATCGCATAAGATTGCCAGGTACAGCTAAATTAGGAGAAGGAAAGCCTGAGAATCAGAATCATGCCATTATA
TTTACTCGAGGGGAAGCTCTGCAAGCCATTGACATGAATCAGGATAATTACTTAGAAGAAGCACTTAAAATGCGTAACCTTCTTGAAGAATTTAATGAGG
ATCATGGAGTGCTTCCACCTACAATTTTGGGTGTTCGGGAGCATATCTTTACTGGAAGTGTCTCTTCTTTGGCTTGGTTCATGTCAAACCAAGAAACGAG
TTTTGTCACCATTGGTCAAAGGGTTCTAGCAAGACCATTGAAGGTCCGGTTCCATTATGGGCATCCAGATGTGTTTGATAGAATCTTTCACGTAACACGT
GGTGGCATCAGCAAGGCTTCTCATGGCATCAATTTGAGTGAGGACATATTTGCTGGTTTCAACTCAACACTGAGACGAGGGAATGTCACTCATCATGAGT
ATATTCAAGTTGGGAAAGGTCGAGATGTCGGGCTCAACCAAATCTCTCTGTTTGAAGCAAAGGTAGCATGTGGTAATGGGGAGCAGACACTCAGCAGGGA
TATCTATCGACTAGGCCATCGTTTTGATTTCTTCCGCATGCTGTCCTGTTATTATACAACGATCGGATTTTACGTCAGTTCAATGATAGTTGTCCTTACG
GTATATGCTTTCTTGTATTGTAAACTTTACCTGTCATTAAGTGGATTGGAGGAATCAATAATTAAGTATGCACGGGCTAGGGGAAATGATCCCCTAAAAG
CAGCCATGGCCTCACAGTCTCTTGTTCAGATTGGTTTTCTGATGGCGTTGCCCATGGTCATGGAGATGGGGCTAGAGAGAGGGTTCAGAACAGCATTAGG
TGACATAATAATCATGCAGCTTCAGCTTGCTTCTGTCTTCTTCACCTTTTCCCTTGGGACTAAGGTCCATTATTTTGGGCGCACTATCCTGCATGGTGGA
GCAAAATACAGGGCTACTGGGCGTGGTTTTGTGGTTCGGCATCAGAAATTTGCTGAGAACTATAGGATGTACTCGAGGAGTCACTTTGTGAAAGGACTGG
AGCTTTTAATATTGCTTATATGTTATAAGATATATGGAAAAGCAGCAAGTGGAGTAGGCTTTGCACTTGTCACGGCCTCAATGTGGTTCTTAGTGACTTC
TTTTTTGTTTGCTCCTTTCCTTTTCAATCCATCCGGATTTGAGTGGCAGAAGATTGTGGATGATTGGGATGATTGGTCAAAGTGGATAAGCAGCCAGGGT
GGCATTGGTGTCCCAGCAAACAAGAGCTGGGAGTCTTGGTGGGATGAGGAACAGGAGCACCTGCAACACACTGGATTTTTAGGCCGGTTCTGGGAGATAT
TTCTCTCTTTGCGTTTCTTCATCTATCAATATGGAATTGTATACCAGCTAAAAGCAGTCAAGGAAAGCACGCCTGGAAGAAGCCGGAGTGCTATTGTATA
TGGTCTGTCGTGGCTGGTCATTGTTGCTATGATGATCATCTTAAAGATTGTTTCCATGGGAAGAAAGAAGTTTAGTGCAGATTTTCAGCTCATGTTTAGG
CTTCTTAAGCTGTTTCTCTTTATTGGATCTGTGATCACTCTCGTTATCCTGTTCACCACTCTTCACCTCACAGTTGGAGACATCTTTCAGAGTCTGCTAG
CTTTTCTGCCGACTGGTTTGGCAATTCTGCAGATAGCACAAGCTTGCAGGCCAGTAGTGAAGGGCCTTAAAATGTGGGGATCTGTGAAAGCTCTAGCCCG
TGGGTACGAGTACATGATGGCTTTGGTCATCTTTGCACCAGTGGCTGTACTTGCCTGGTTCCCATTCGTTTCAGAATTCCAGACCAGGCTGCTGTTTAAC
CAAGCTTTCAGCCGAGGGCTTCAGATTCAACGTATCTTGGCTGGCGGAAAGAAAAACAAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G058300.1 pacid=42803680 polypeptide=Potri.005G058300.1.p locus=Potri.005G058300 ID=Potri.005G058300.1.v4.1 annot-version=v4.1
MSNLEPSGSQMLSRRPSRSAATTTFSLEVFDNEVVPSSLGSIAPVLRIAAEIEHERPRVAYLCRFYAFEKAHRLDPNSSGRGVRQFKTSLLQRLERDNNS
SLASRVKKTDAREIESFYQQYYEHYVRALDQGEQADRAQLGKAYQTAGVLFEVLCAVNKTEKVEEVAPEIIAAARDVQEKKEIYAPFNILPLDSAGASQS
IMQLEEVKAAVAALWNTRGLNWPTAFDPQRQKAGDLDILDWLRAMFGFQRDNVRNQREHLILLLANKHIRLNPKPEPISKLDDRAVDEVMNKLFKNYKTW
CKFLGRKHSLRLPQGQPEIQQRKILYMGLFLLIWGEAANVRFMPECLCYIFHNMAYELHGLLAGNVSIVTGENIKPSYGGDDEAFLRKVITPIYHVIEKE
ANKSKNGKASHSQWCNYDDLNEYFWSSDCFSLGWPMRDDGSFFTSTRDVGKKASSEKPRSTGKAYFVETRTFWHIFRSFDRLWTFYILALQAMIIIAWSG
VSILNIVQKDVLYQLSSIFITAACLRLLQSILDLVLNFPGFHKWKFTDVLRNVLKIIVSLAWAIILPLCYVHSFKVAPDKIKDLLSFFKEVKDIPALYLL
AVAVYMLPNILAAALFIFPMLRRWIENSDWLIIRFLLWWSQPRIYVGRGMHESQFVLIKYTVFWLLLLCSKIAFSYFVQIKPLVKPTKAIMNIRNVDYEW
HEFFPNAKNNYGAVLSLWLPVILVYFMDTQIWYSIFSTIYGGFAGAFDRLGEIRTLGMLRSRFQSLPGAFNTYLVPSDKKRKKGFSFSKRFSEVTASKRS
EAAKFAQLWNEVICSFREEDLISDREMDLLLVPYTSDPSLKLIQWPPIMLASKIPIALDMAVQFRSRDADLWKRICADEYMKCAVIECYESFKHVLNILV
VGEIEKRILSIIFKEVESNISKNTLLTNFRMGPLPALCNKFVELVILLKDADPSKQNTVVLILQDMLEVFTNDMMVNENRELVDLGQSGKDSGRQVFSGT
DTKPAIMFPPVVTAQWEEQIRRIHLLLTVNEFANDVPTNLEARRRISFFTNSLFMDMPRPPRVRKMLSFSVLTPYYSEETVYSKSDLEMENEDGVSIIYY
LQKIYPDEWNNFMERINCKKESEVWENEENILQLRHWGSLRGQTLCRTVRGMMYYRRALRLQAFLDMAKESEILEGYKAITDPTEEDKKSQRSVSAQIEA
VADMKFTYVATCQNYGNQKRSGDRRATDILNLMVNNPSLRVAYIDEVEEREREGGKVQKVYYSVLVKAVDNLDQEIYRIRLPGTAKLGEGKPENQNHAII
FTRGEALQAIDMNQDNYLEEALKMRNLLEEFNEDHGVLPPTILGVREHIFTGSVSSLAWFMSNQETSFVTIGQRVLARPLKVRFHYGHPDVFDRIFHVTR
GGISKASHGINLSEDIFAGFNSTLRRGNVTHHEYIQVGKGRDVGLNQISLFEAKVACGNGEQTLSRDIYRLGHRFDFFRMLSCYYTTIGFYVSSMIVVLT
VYAFLYCKLYLSLSGLEESIIKYARARGNDPLKAAMASQSLVQIGFLMALPMVMEMGLERGFRTALGDIIIMQLQLASVFFTFSLGTKVHYFGRTILHGG
AKYRATGRGFVVRHQKFAENYRMYSRSHFVKGLELLILLICYKIYGKAASGVGFALVTASMWFLVTSFLFAPFLFNPSGFEWQKIVDDWDDWSKWISSQG
GIGVPANKSWESWWDEEQEHLQHTGFLGRFWEIFLSLRFFIYQYGIVYQLKAVKESTPGRSRSAIVYGLSWLVIVAMMIILKIVSMGRKKFSADFQLMFR
LLKLFLFIGSVITLVILFTTLHLTVGDIFQSLLAFLPTGLAILQIAQACRPVVKGLKMWGSVKALARGYEYMMALVIFAPVAVLAWFPFVSEFQTRLLFN
QAFSRGLQIQRILAGGKKNK
|
DESeq2's median of ratios [POPLAR]
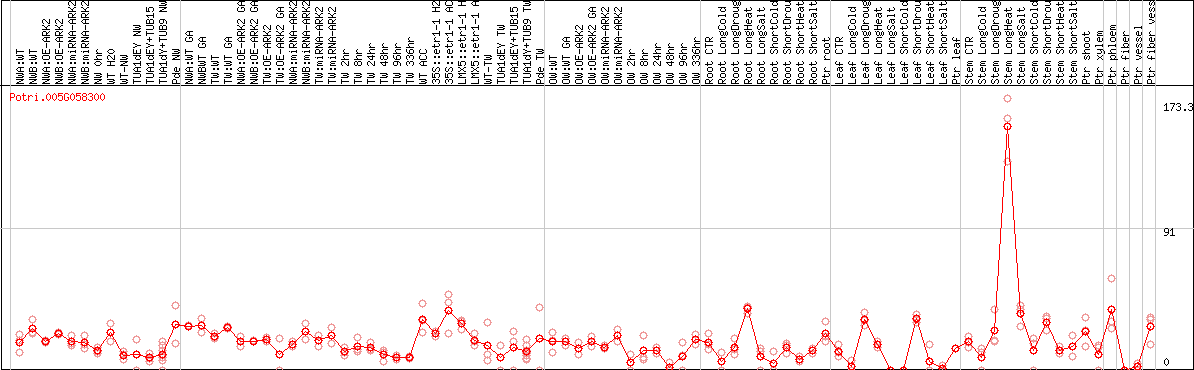
Coexpressed genes
Potri.005G058300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
