External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39660 219 / 1e-69
AGT2
alanine:glyoxylate aminotransferase 2 (.1)
AT3G08860 204 / 1e-63
PYD4
PYRIMIDINE 4 (.1)
AT2G38400 179 / 8e-54
AGT3
alanine:glyoxylate aminotransferase 3 (.1.2)
AT1G80600 82 / 3e-18
WIN1
HOPW1-1-interacting 1 (.1)
AT5G46180 74 / 3e-15
DELTA-OAT
ornithine-delta-aminotransferase (.1)
AT3G22200 66 / 1e-12
HER1, GABA-T, POP2
POLLEN-PISTIL INCOMPATIBILITY 2, HEXENAL RESPONSE1, GAMMA-AMINOBUTYRATE TRANSAMINASE, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1), Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.2)
AT3G48730 54 / 3e-08
GSA2
glutamate-1-semialdehyde 2,1-aminomutase 2 (.1)
AT5G63570 50 / 4e-07
GSA1
"glutamate-1-semialdehyde-2,1-aminomutase", glutamate-1-semialdehyde-2,1-aminomutase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G085600
244 / 4e-79
AT4G39660 814 / 0.0
alanine:glyoxylate aminotransferase 2 (.1)
Potri.006G106800
194 / 1e-59
AT3G08860 771 / 0.0
PYRIMIDINE 4 (.1)
Potri.016G132200
193 / 2e-59
AT2G38400 769 / 0.0
alanine:glyoxylate aminotransferase 3 (.1.2)
Potri.007G080000
100 / 1e-26
AT4G39660 278 / 5e-93
alanine:glyoxylate aminotransferase 2 (.1)
Potri.017G053500
79 / 6e-17
AT1G80600 677 / 0.0
HOPW1-1-interacting 1 (.1)
Potri.011G082800
77 / 4e-16
AT5G46180 734 / 0.0
ornithine-delta-aminotransferase (.1)
Potri.005G095800
75 / 2e-15
AT1G80600 636 / 0.0
HOPW1-1-interacting 1 (.1)
Potri.016G018500
73 / 5e-15
AT3G22200 826 / 0.0
POLLEN-PISTIL INCOMPATIBILITY 2, HEXENAL RESPONSE1, GAMMA-AMINOBUTYRATE TRANSAMINASE, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1), Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.2)
Potri.006G020900
71 / 3e-14
AT3G22200 834 / 0.0
POLLEN-PISTIL INCOMPATIBILITY 2, HEXENAL RESPONSE1, GAMMA-AMINOBUTYRATE TRANSAMINASE, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1), Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004480
230 / 1e-73
AT4G39660 801 / 0.0
alanine:glyoxylate aminotransferase 2 (.1)
Lus10029922
228 / 9e-73
AT4G39660 801 / 0.0
alanine:glyoxylate aminotransferase 2 (.1)
Lus10025262
179 / 1e-53
AT3G08860 722 / 0.0
PYRIMIDINE 4 (.1)
Lus10009086
176 / 5e-53
AT3G08860 625 / 0.0
PYRIMIDINE 4 (.1)
Lus10008025
81 / 1e-17
AT1G80600 614 / 0.0
HOPW1-1-interacting 1 (.1)
Lus10023299
77 / 3e-16
AT5G46180 743 / 0.0
ornithine-delta-aminotransferase (.1)
Lus10038510
77 / 3e-16
AT5G46180 741 / 0.0
ornithine-delta-aminotransferase (.1)
Lus10031196
73 / 9e-15
AT1G80600 606 / 0.0
HOPW1-1-interacting 1 (.1)
Lus10010465
70 / 9e-14
AT3G22200 756 / 0.0
POLLEN-PISTIL INCOMPATIBILITY 2, HEXENAL RESPONSE1, GAMMA-AMINOBUTYRATE TRANSAMINASE, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1), Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.2)
Lus10003810
70 / 1e-13
AT3G22200 758 / 0.0
POLLEN-PISTIL INCOMPATIBILITY 2, HEXENAL RESPONSE1, GAMMA-AMINOBUTYRATE TRANSAMINASE, Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1), Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0061
PLP_aminotran
PF00202
Aminotran_3
Aminotransferase class-III
Representative CDS sequence
>Potri.005G082100.3 pacid=42803009 polypeptide=Potri.005G082100.3.p locus=Potri.005G082100 ID=Potri.005G082100.3.v4.1 annot-version=v4.1
ATGCAATATTTGTTTGATGAAAATGGAAGGAGGTATTTGAATGCTTTTTCTGGGATAGTGTCTGTCTCTTGCGGCCACTGCCATCCTCAAATTTTAAATG
CAATTACGGAGCAAAGCAAGCTGCTCCAACATGCTACTACCATTTACTTGAACCACACTATTGCTGATTTTGCTGAGGCCTTAGCTGCTAAGATGCCTGG
AAATCTCAAGGGCGAAATTCATCATGTTATTAATCCAAATCCACACAACAATTATGGTACTTCTGGAAAAGTTGCTGGTTTCATATCCGAAACTATTCAG
GGAGTTGGAGGAGCAGTTGAATTAGCCCCTGGATACTTGACAATGGTTTATGACATTGTGCGTAAGGCAGGGGGAGTTTGCATAGCTGACGAAGTGCAAT
CGGGGTTTGGCCGCACAGGAAGCTGCTATTGGGGTTTTGAGACTCAAGGTGTCATCCCTGACATTGTTACCATGGCAAAGGGTATTGACAATGGTTTGCC
ATTGGGAGCTGTAGTAACAACACCAGAAATAGCACAAGTCATGGCACAGAAAATTCAATTCAATACTTTTGGTGGAAACCCTGTGTTTTCTGCCAGTGGA
CATGAAGTACTGAGAGTCATTGACCAAGAGAGGCGTCAAGAATAG
AA sequence
>Potri.005G082100.3 pacid=42803009 polypeptide=Potri.005G082100.3.p locus=Potri.005G082100 ID=Potri.005G082100.3.v4.1 annot-version=v4.1
MQYLFDENGRRYLNAFSGIVSVSCGHCHPQILNAITEQSKLLQHATTIYLNHTIADFAEALAAKMPGNLKGEIHHVINPNPHNNYGTSGKVAGFISETIQ
GVGGAVELAPGYLTMVYDIVRKAGGVCIADEVQSGFGRTGSCYWGFETQGVIPDIVTMAKGIDNGLPLGAVVTTPEIAQVMAQKIQFNTFGGNPVFSASG
HEVLRVIDQERRQE
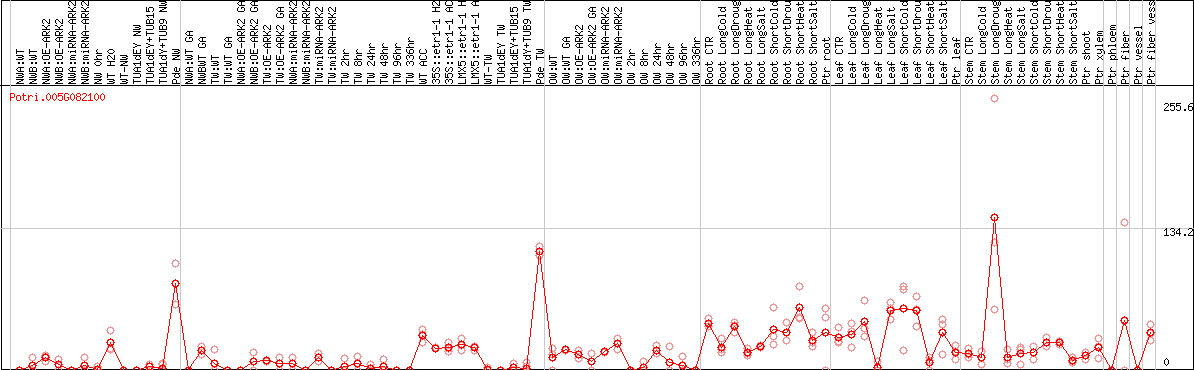
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G082100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.