External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G23140 380 / 3e-135
NCLPP7, NCLPP2, CLPP2
nuclear-encoded CLP protease P7 (.1)
AT1G02560 160 / 7e-48
NCLPP5, NCLPP1, CLPP5
NUCLEAR CLPP 5, NUCLEAR-ENCODED CLPP 1, nuclear encoded CLP protease 5 (.1)
AT5G45390 142 / 3e-41
NCLPP3, NCLPP4, CLPP4
NUCLEAR-ENCODED CLP PROTEASE P4, CLP protease P4 (.1)
AT1G66670 142 / 7e-41
NCLPP3, NCLPP4, CLPP3
CLP protease proteolytic subunit 3 (.1)
ATCG00670 137 / 4e-40
PCLPP, ATCG00670.1, CLPP1
CASEINOLYTIC PROTEASE P 1, plastid-encoded CLP P (.1)
AT1G12410 125 / 7e-35
EMB3146, CLP2, NCLPP2, CLPR2
NUCLEAR-ENCODED CLP PROTEASE P2, EMBRYO DEFECTIVE 3146, CLP protease proteolytic subunit 2 (.1)
AT4G17040 125 / 2e-34
HON5, CLPR4
happy on norflurazon 5, CLP protease R subunit 4 (.1)
AT1G11750 118 / 3e-32
NCLPP6, NCLPP1, CLPP6
NUCLEAR-ENCODED CLPP 1, CLP protease proteolytic subunit 6 (.1.2)
AT1G09130 111 / 5e-29
ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.1), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.2), ATP-dependent caseinolytic (Clp) protease/crotonase family protein (.3)
AT1G49970 82 / 6e-18
SVR2, NCLPP5, CLPR1
SUPPRESSOR OF VARIEGATION 2, NUCLEAR CLPP 5, CLP protease proteolytic subunit 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G071700
437 / 7e-157
AT5G23140 383 / 1e-135
nuclear-encoded CLP protease P7 (.1)
Potri.002G195200
160 / 4e-48
AT1G02560 474 / 2e-170
NUCLEAR CLPP 5, NUCLEAR-ENCODED CLPP 1, nuclear encoded CLP protease 5 (.1)
Potri.014G119700
160 / 7e-48
AT1G02560 493 / 6e-178
NUCLEAR CLPP 5, NUCLEAR-ENCODED CLPP 1, nuclear encoded CLP protease 5 (.1)
Potri.004G092100
149 / 3e-43
AT1G66670 441 / 2e-156
CLP protease proteolytic subunit 3 (.1)
Potri.003G103300
145 / 5e-42
AT5G45390 383 / 9e-135
NUCLEAR-ENCODED CLP PROTEASE P4, CLP protease P4 (.1)
Potri.009G114001
129 / 2e-36
AT1G11750 353 / 2e-123
NUCLEAR-ENCODED CLPP 1, CLP protease proteolytic subunit 6 (.1.2)
Potri.004G152900
129 / 2e-36
AT1G11750 397 / 4e-141
NUCLEAR-ENCODED CLPP 1, CLP protease proteolytic subunit 6 (.1.2)
Potri.001G115900
122 / 2e-33
AT1G12410 408 / 3e-145
NUCLEAR-ENCODED CLP PROTEASE P2, EMBRYO DEFECTIVE 3146, CLP protease proteolytic subunit 2 (.1)
Potri.003G083300
122 / 3e-33
AT4G17040 441 / 2e-157
happy on norflurazon 5, CLP protease R subunit 4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017397
369 / 6e-131
AT5G23140 394 / 1e-140
nuclear-encoded CLP protease P7 (.1)
Lus10010196
369 / 1e-130
AT5G23140 395 / 4e-141
nuclear-encoded CLP protease P7 (.1)
Lus10040981
313 / 4e-108
AT5G23140 350 / 1e-122
nuclear-encoded CLP protease P7 (.1)
Lus10013434
311 / 4e-107
AT5G23140 347 / 3e-121
nuclear-encoded CLP protease P7 (.1)
Lus10040982
300 / 8e-103
AT5G23140 338 / 5e-118
nuclear-encoded CLP protease P7 (.1)
Lus10013435
179 / 1e-54
AT5G23140 209 / 5e-66
nuclear-encoded CLP protease P7 (.1)
Lus10002422
148 / 2e-43
AT1G02560 361 / 4e-126
NUCLEAR CLPP 5, NUCLEAR-ENCODED CLPP 1, nuclear encoded CLP protease 5 (.1)
Lus10001450
150 / 3e-43
AT1G02560 365 / 1e-126
NUCLEAR CLPP 5, NUCLEAR-ENCODED CLPP 1, nuclear encoded CLP protease 5 (.1)
Lus10000063
126 / 6e-37
AT5G23140 128 / 2e-38
nuclear-encoded CLP protease P7 (.1)
Lus10007589
129 / 5e-36
AT5G45390 293 / 2e-99
NUCLEAR-ENCODED CLP PROTEASE P4, CLP protease P4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0127
ClpP_crotonase
PF00574
CLP_protease
Clp protease
Representative CDS sequence
>Potri.005G092600.1 pacid=42803007 polypeptide=Potri.005G092600.1.p locus=Potri.005G092600 ID=Potri.005G092600.1.v4.1 annot-version=v4.1
ATGAAAGCCCTCTTCTCTACCACCAAGCTCATCACTTCCACAGCTAAACCAAAACCCTCTTTTCTCTTGACCCCCAAAACCTCTACCACTAGAGCTTATA
GTCTAATCCCAATGGTTATCGAACACTCTTCAAGAGGAGAAAGAGCTTATGATATATTTTCTAGGCTTTTAAAAGAGAGAATTGTCTGTATTAATGGCCC
AATTAATGATGATACTTCTAATGTTGTTGTTGCTCAGCTTTTGTTTCTCGAGTCTGAGAATCCTTCTAAGCCTATCCATATGTATCTGAATTCTCCTGGT
GGACATGTTACCGCAGGTCTTGCTATTTACGATACAATGCAGTACATAAGGTCTCCAGTTAATACCATTTGTTTGGGTCAAGCTGCATCTATGGCTTCTC
TCCTTTTGGCTTCTGGGGCTAAGGGTGAGAGGAAGGCACTTCCAAATGCAACAATTATGATTCATCAGCCATCAGGTGGGTACAGCGGACAGGCAAAGGA
TTTGACAATTCACACCAAGCAGATAGTTCGGGTCTGGGATGCACTGAATCAATTGTACTGCAAGCATACAGGGAAGCCAATTGATGTAATTCAGAAGAAT
ATGGATAGAGATTATTTTATGACCCCAGAAGAGGCAAAGGAATTCGGAATTATTGATGAGGTGATTGATCAAAGACCCATGACTCTGGTGACTGATGCTG
TTGGCGATGAAAGTAAACAAAAGGGTTCGAGCTAG
AA sequence
>Potri.005G092600.1 pacid=42803007 polypeptide=Potri.005G092600.1.p locus=Potri.005G092600 ID=Potri.005G092600.1.v4.1 annot-version=v4.1
MKALFSTTKLITSTAKPKPSFLLTPKTSTTRAYSLIPMVIEHSSRGERAYDIFSRLLKERIVCINGPINDDTSNVVVAQLLFLESENPSKPIHMYLNSPG
GHVTAGLAIYDTMQYIRSPVNTICLGQAASMASLLLASGAKGERKALPNATIMIHQPSGGYSGQAKDLTIHTKQIVRVWDALNQLYCKHTGKPIDVIQKN
MDRDYFMTPEEAKEFGIIDEVIDQRPMTLVTDAVGDESKQKGSS
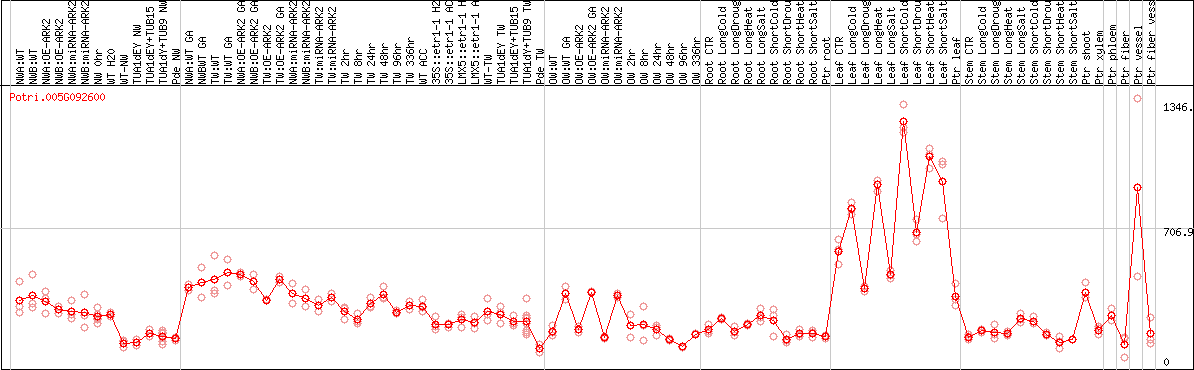
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G092600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.