Potri.005G094200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G094200.3 pacid=42803262 polypeptide=Potri.005G094200.3.p locus=Potri.005G094200 ID=Potri.005G094200.3.v4.1 annot-version=v4.1
ATGTCTCGGCAAACGACATCATCTCATTTTCATAAATCGAAGACTCTCGATAACAAATATATGCTTGGAGATGAGATTGGAAAAGGAGCGTATGCTCGGG
TCTTTAAAGGGTTGGATTTGGAGAATGGAGATTTTGTTGCGATCAAACAAGTTTCATTGGAGAATATCGCCCAAGAGGATCTTAATATCATCATGCAAGA
GATTGATTTACTGAAGAATTTGAACCATAAAAACATTGTGAAATATCTTGGATCATTGAAGACAAAGTCTCATCTCCACATAATCCTTGAGTATGTGGAG
AATGGCTCGCTTGCAAATATTATCAAGCCAAATAAATTTGGGCCTTTTCCAGAGTCTTTGGTGGTTGTTTACATTGCCCAGGTTCTGGAAGGATTAGTTT
ATTTGCACGAACAGGGTGTTATCCATCGAGATATCAAGGGTGCAAACATATTGACGACCAAAGAGGGTCTTGTGAAACTTGCTGATTTTGGTGTTGCAAC
AAAATTAACTGAAGCTGATGTTAATACTCATTCGGTTGTTGGAACGCCATATTGGATGGCCCCAGAGGTTATTGAAATGTCAGGGGTTTGTGCTGCTTCT
GACATCTGGAGTGTTGGCTGCACTGTGATTGAACTCCTTACGTGTGTACCACCTTACTATGATTTGCAGCCAATGCCAGCTCTCTTTCGTATTGTTCAGG
ATGATCGTCCTCCGATACCTGATAGCTTATCTCCTGACATTACAGACTTTCTCCGCCAGTGCTTCAAGAAGGATGCAACGCAGCGGCCTGATGCAAAGAC
ACTGCTTTCTCATCCTTGGATATTAAATTCTAGACGTGCTTTGAATTCTTTCCGTCATAGTGGACCAATAAGAAGCATACAGGAAGATGTTTCAGCAGAA
GCAGAAATTTTAACTGGAGATAACCAGAGGACTGTTCAAATCAATTCTGTTGATAGAACAAAAGCATCTGTTGCCGACTTCAAAGCTGGCTCTAGGAAAG
AATCATTGCCAGATTCTGAGGATGTGAGTAAGTCTGATAAGAATACTTCTTCAGATGGTGATGTAGTTGAGGAAAGAATTGATAAATTGGAAGATGATCT
CCATTCAGATCAAGTCCCAACCCTAGCTATTCACGAAAACTCATCTCTAAAAACTAGTCCTGGCAGACTATCTACAAATAAAGTAGCTGCTGCTTCTCCT
TTGCTTCATGGATCGATGCCTTTGCATTACCAGGATGAGATTCTGACTATTGATGACTTGGAATCTCCCGATGCAAGAGGAAAAAACATTGAAAGAAGGA
ATGGAGGAAAAACAAGTCCTGCTCGTGTTGAGAATGGATCATTTGGTTTTGCAACAAGAAATCAAGATAATGGGCTTCGAAAGGCTGTGAAGACATCAAT
GACCTCAGGAGGAAACGAACTTAGTAAATTTAGTGACACTCCAAGAGATGCTTCACTGGATGATTTATTTCACCCGCTTGATAAAAATCCTGAGGATCGG
GCAGCTGAAGCTTCAACATCTACCTCAGCATCACATATGAACCAAGGGAATGCAATCATGGCTGATGCAGGGAAAAATGACCTGGCTGCAATTTTGAGAG
CGACTATTGCTCAAAAACAAATGGAGAGTGAAACGGGACAAACTAATGGTGGTGGTGATTTGTTTCGCCTAATGATGGGTGTTCTAAAGGATGGCGTGAT
TGATATTGATGGTTTGGATTTTGGAGACAAACTGCCGGCAGAAAATCTCTTTCCTCTACAAGCTGTTGAGTTTAGCAGGTTAGTTGGGTCATTAAGACCT
GAGGAATCAGAAGATGTGATTACGTCTTCATGCCAGAAACTTATTAGCATCTTTCACCAGCGCCCCGAGCAGAAAATTGTTTTTATTACCCAGCATGGTT
TGCTTCCTCTAATGGAATTACTTGAAGTTCCTAAACCTCGTGTTATATGCTCTATCCTTCAACTTATAAATCAGATCGTGAAAGATAATACTGATTTCCA
GGAAAATGCTTGTCTTGTTGGCTTGATCCCTGTGGTTACGAGTTTTGCAGGACCTGATCGCCCTCGTGAAGTTCGCATGGAAGCAGCTTATTTCTTGCAG
CAGCTATGCCAGTCAAGCTCCTTGACATTGCAAATGTTCATTGCTTGCCGAGGAATACCTATCCTGGTGGGTTTTCTAGAGGCTGATTACGCGAAGCACA
GGGACATGGTTCACCTAGCTATCGATGGCATGTGGCAGGTCTTTAAGCTTCAGAGATCAACCCCAAGAAATGATTTCTGTCGCATAGCTGCAAAAAATGG
AATACTGCTCAGACTTATCAACACGCTTTATAGCTTGAATGAGGCAACTAGGCTAGCTTCAATATCTGTGGGAACTGGATTTCCACTAGATGGTTTATCT
CAACGACCACGATCTGGTCCCTTGGATTCCAATCATCCTATTTTCATTCAGAGCGAGACAGCTCTTTCTGCATCTGATCAGCCTGATGTCTTTAAAGTTA
GACATGGAATGATTGATCATTCCTTGCCTTTTGGGACACAAGAACCATCACGTGCTTCAACTTCACATTCCCAAAGGTTGGATGCCATTCAACCAGATGC
CAGATTTTTTGGGACAGACACAGATGGATCTCAGGCTAGCAATGAAACAATCGAAGCTATAGCTGCTTCCAAGTTGTCAGATCCAGCAGCTTTAGGAAAA
GCTCCAAACATGGCCACCAAAGAACCTTCAGGTGCTGTTTCTAAAGAGCGAGATAACTTGGACCGGTGGAAAAGTGATCCATCTCGCCCAGAAATTGACC
TTAGGCAGCAGCGAGTCACTGGTTCCACACAAAGGACATCTACTGACAGGCCTCCAAAGTTGATAGAAAGTGCATCAAATGGTCTTACTTCTATGATATC
TGCCCAGCCAGAGCAAGTTCGTCCTCTTCTAAGCTTGTTGGAGAAGGAACCCCCCTCTAGGCATTTCTCTGGTCAGTTGGAGTATGCACGTCACCTCACA
GGGTTAGAGAGACATGAAAGCATACTGCCTTTATTACATGCATCTGAGAAGAAAACAAATGGTGGATTAGAGTTTCTGATGGCAGAGTTTGCAGAGGTAT
CTGGGCGTGGAAGAGAGAATGGAAATCTAGACTCCATTCCTAGAATTTCACACAAGACAGTGAGTAAGAAGGTTGGATCTCTTGCACCTAATGAAGGCGC
TGCTTCCACATCTGGTATTGCATCTCAAACAGCATCTGGTGTACTGTCTGGTTCGGGTGTTCTAAATGCTAGACCTGGGAGTGCAACATCATCTGGGTTA
CTTTCCCAGATGGTATCAACTATGAATGCAGAGGTTGCAAGGGAATACCTGGAGAAAGTGGCAGACCTTCTGCTGGAGTTTTCTCAAGCTGATACAACTG
TAAAATCCTATATGTGTAGCCAAAGCTTGCTTAGTCGTCTTTTTCAGATGTTCAACAGAATTGAGCCCCCTATTCTTTTGAAGATACTTGAGTGTATCAA
TAACCTGTCAACTGACCCGAACTGCTTAGAGAATCTTCAGCGCGCAGATGCAATTAAGTACCTGATCCCAAATCTTGAACTCAAAGACGGGCCTCTTGTA
GATCAAATTCATAGTGAGGTCCTCAATGCACTTTTTAACCTCTGCAAGATAAATAAGAGGAGGCAGGAACAAGCTGCTGAAAATGGAATCATTCCACACT
TGATGAATTTTATTATGTCAGATTCTCCTTTAAAACCTCATGCATTACCTCTACTGTGTGACATGGCTCATGCATCACGTAATTCAAGAGAGCAATTGAG
GGCTCATGGTGGATTGGATGTATACTTGAGCCTGCTGGATGATACGGTTTGGTCAGTGACAGCATTGGACTCTATTGCTGTTTGCTTGGCTCATGACAAT
GACAACCGCAAAGTGGAACAAGCATTGCTAAAAAAGGACGCAGTTCAGAAATTAGTGAAGTTTTTCCAATGCTGCCCGGAACAACAATTTGTGCACATAT
TGGAGCCATTCTTGAAAATTATAACGAAATCATCTCGGATAAATACGACGTTAGCTGTTAATGGCTTGACACCTCTGCTCATTGGGAAACTGGATCATCA
GGATGCCATTGCTCGTCTTAATCTGCTTAAACTAATAAAGTCTGTGTATGAGCACCACCCCCGTCCAAAGCAGTTGATCGTGGAGAATGATTTGCCCCAA
AAACTCCAGAACTTGATCGAGGAACGTAGGGATGGGCAAAGCTCCGGAGGCCAAGTGCTAGTTAAACAAATGGCTACTTCGCTACTTAAAGCACTTCATA
TAAATACCGTTCTGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G094200.3 pacid=42803262 polypeptide=Potri.005G094200.3.p locus=Potri.005G094200 ID=Potri.005G094200.3.v4.1 annot-version=v4.1
MSRQTTSSHFHKSKTLDNKYMLGDEIGKGAYARVFKGLDLENGDFVAIKQVSLENIAQEDLNIIMQEIDLLKNLNHKNIVKYLGSLKTKSHLHIILEYVE
NGSLANIIKPNKFGPFPESLVVVYIAQVLEGLVYLHEQGVIHRDIKGANILTTKEGLVKLADFGVATKLTEADVNTHSVVGTPYWMAPEVIEMSGVCAAS
DIWSVGCTVIELLTCVPPYYDLQPMPALFRIVQDDRPPIPDSLSPDITDFLRQCFKKDATQRPDAKTLLSHPWILNSRRALNSFRHSGPIRSIQEDVSAE
AEILTGDNQRTVQINSVDRTKASVADFKAGSRKESLPDSEDVSKSDKNTSSDGDVVEERIDKLEDDLHSDQVPTLAIHENSSLKTSPGRLSTNKVAAASP
LLHGSMPLHYQDEILTIDDLESPDARGKNIERRNGGKTSPARVENGSFGFATRNQDNGLRKAVKTSMTSGGNELSKFSDTPRDASLDDLFHPLDKNPEDR
AAEASTSTSASHMNQGNAIMADAGKNDLAAILRATIAQKQMESETGQTNGGGDLFRLMMGVLKDGVIDIDGLDFGDKLPAENLFPLQAVEFSRLVGSLRP
EESEDVITSSCQKLISIFHQRPEQKIVFITQHGLLPLMELLEVPKPRVICSILQLINQIVKDNTDFQENACLVGLIPVVTSFAGPDRPREVRMEAAYFLQ
QLCQSSSLTLQMFIACRGIPILVGFLEADYAKHRDMVHLAIDGMWQVFKLQRSTPRNDFCRIAAKNGILLRLINTLYSLNEATRLASISVGTGFPLDGLS
QRPRSGPLDSNHPIFIQSETALSASDQPDVFKVRHGMIDHSLPFGTQEPSRASTSHSQRLDAIQPDARFFGTDTDGSQASNETIEAIAASKLSDPAALGK
APNMATKEPSGAVSKERDNLDRWKSDPSRPEIDLRQQRVTGSTQRTSTDRPPKLIESASNGLTSMISAQPEQVRPLLSLLEKEPPSRHFSGQLEYARHLT
GLERHESILPLLHASEKKTNGGLEFLMAEFAEVSGRGRENGNLDSIPRISHKTVSKKVGSLAPNEGAASTSGIASQTASGVLSGSGVLNARPGSATSSGL
LSQMVSTMNAEVAREYLEKVADLLLEFSQADTTVKSYMCSQSLLSRLFQMFNRIEPPILLKILECINNLSTDPNCLENLQRADAIKYLIPNLELKDGPLV
DQIHSEVLNALFNLCKINKRRQEQAAENGIIPHLMNFIMSDSPLKPHALPLLCDMAHASRNSREQLRAHGGLDVYLSLLDDTVWSVTALDSIAVCLAHDN
DNRKVEQALLKKDAVQKLVKFFQCCPEQQFVHILEPFLKIITKSSRINTTLAVNGLTPLLIGKLDHQDAIARLNLLKLIKSVYEHHPRPKQLIVENDLPQ
KLQNLIEERRDGQSSGGQVLVKQMATSLLKALHINTVL
|
DESeq2's median of ratios [POPLAR]
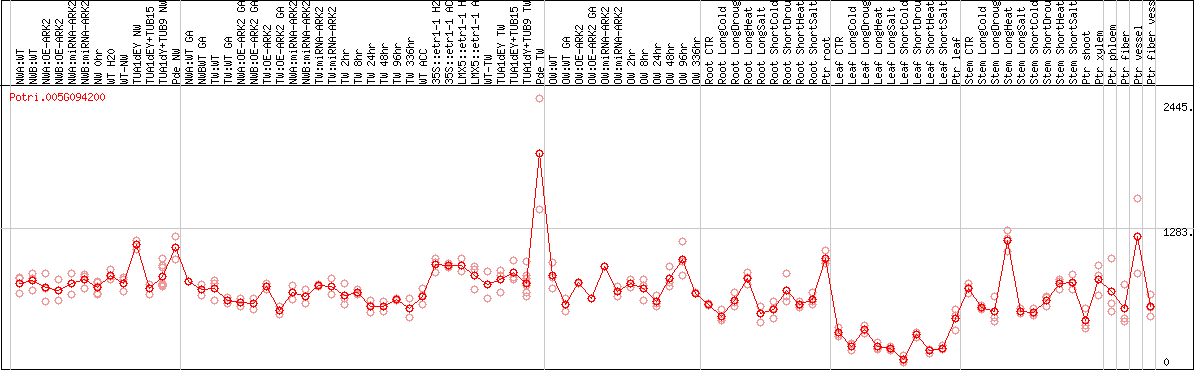
Coexpressed genes
Potri.005G094200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
