Potri.005G101900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.005G101900.2 pacid=42802914 polypeptide=Potri.005G101900.2.p locus=Potri.005G101900 ID=Potri.005G101900.2.v4.1 annot-version=v4.1
ATGGGTATAATTAAGCGATTGCCGGTGTCAGCACGCAGCACAATGCGTTCAGGGATCTTGGTGTTTGATTTAACAAGAGTTGTAGAGGAGCTTGTTTTTA
ATAGCTTGGATGCTGGTGCTAAAAAGGTTTCGGTTTACGTGGCTGTTGGTACCTGCTATGTGAAAGTTTCTGATGATGGATGTGGGATTTCACGGGATGG
ATTGGTGTTACTGGGAGAAAGATACGTGACCTCAAAGGTTCAACGTTTGGCTGATATGGATGTTGCTAGTGGAAACTTTGGCTTCCGGGGAGAAGCGTTG
TCTTCTATCGCTGATGTCTCAGTGTTAGATGTATTGACAAAAGCTCGTGGAATGCCGAATGGATATCGCAAGGTCATGAAGGGATCTAAGTGTTTGTGTT
TGGGAATTGATGATGATATTAAAGATGTCGGCACTACAGTTGTTGTCCGTGATTTATTTTACAACCAACCAGTTCGGAGGAAATATATGCAATCCAGCCC
TAAGAAGATACTGCACTTGGTAAAAAAATGTGCACTTCGTGTTGCTCTTATGCATTCAGAGGTGTCCTTTAAAGTTGTTGATATTGAAAGTGACGAGGAG
CTTCTCTGCACAAATCCATCTTCTGCAATGTCACTGTTGATGAGTGGATTTGGAATTGAAGATTCCAGCTCTCTTCATGAACTGAATATCAGTGACAGTG
TATTAAAGCTTTCTGGCTATATATCTGGTCCTTGCAGTAGTTTTTTCATCAAGGCTTTCCAATATGTTTATATCAATTCACGGTTTGTTTGCAAAGGCCC
AATTCACAAATTGCTAAATCACCTGGCCTCTAGATTTGAGCATCCAGATCTGCAGAAGGCTAACAGTGTGTCTCAAAAAGGAAAGAAAAGCAGGCCTCAA
CCATGTCCAGCTTATATTTTAAATTTGAGTTGCCCTTTCTCTCTCTATGATTTAACCTTTGAGCCATCAAAGACACATGCTGAATTCAAGGATTGGAACC
CTATCCTTGCCTTCATTGAGAAGGTCATTCAACAACTCTGGAGGGAATGCACCGTTATTGGAGAATCCTCAACTCGGGCAACAGATACGTTCCAGAAAAA
TGACATTTGGCAGGAAGGTAATGATATCACATCAGTGAAACAAGATTTTTTTGATGCAGATTTTTCTGGATTTGCAATAAAGAAAGGCGGGGTTAAAATT
CACCAGTCTTCTCATCATCTCATTTCTTGCCCTTTGAAGATGCTAGACAAAGAGGTCGACCATCTTTTTCATGGGAAGCATGATAAAGTTCCTCAAGAAT
TCTACTCAAATGTTTCAGAATTCAAAGAGGAACAAGTTGATAAGGAATTTGTTCTTCAGGGTGACTACTCTTCCCAAACATGGAATGGATCCATCTCTGG
ATACATGCCAAGAGCAACCAAAACAGATGAGTGTCATCTCTTGACATCTGATAAGAATTTTTTATTGACAGATAATTGCTTCTTGGAAGATAGTTTCACT
ACCAGGGAAAGACTCAGTGATCACATGCAGAGCCATTTCTCAAGCTCAGAATGGCAAAACGAGTCCCCAAAAATTGATTCCGTGGCAAGAAATAAATCTC
TTGGGAGTGCATTCTCCTTTGATCATTATGGATTCAGAAACGAATTACCGTTTAGCAAGAGTAATATAAAACCCATTTTACAGAGTTGCTCCTCACAGAA
AAGCCTGTCACTTGATAGGGATTTTTTTGCTGACAAGGAAGCATTTGAATTTCTTAATGATGGCTTCAAGAACAAAAGAAGACGGCTTTGGACAGCTGAA
AATGTGGGTATCCCAAAAGGGGATACGATATTTGATATCTTTCCCTGTGCTTTACTGCAAGACAATGCATCGTGTACTCAGCAGTTGCCTGCAGATACTG
ATGGAGCTGAGATGTCTGCAGCTTTTGATCTTCTGCCAGGTGCTTATGTAAATTCTTCTTCACCAAATGGAAAAATTTTAGCTAAAGGGAAAGGTCTTGC
ATCTAATTCTATCCTACAATTAGAAATGTATGCCTCAGGAAATCATTCATCAATGTCTGATTGGTGTTCTGTAACTTCAAGTGCTTTTTTTCAAGCAAAA
GTTTGGGATGCTGAACACTTTCCAGATGATAATGCAAGTGAGGGGAGCAAGGGATGGGGTAAAAAGGAAAATTGTTGGCATCTTCCTGATAGCTGGGAGA
TTATGTCAAAGCCTTCCAGCCAAGATAACTTCTTCTCAAGTTGTACAAGCAGTGTGTTAGATTTTAAAAATTCAGCTGATTCTAGTAAAGATATCTGCAA
ATTGCCTCAGTGGCAGGATCAAAATAATGAATTTTCTTTACAACATTCAGATATATCTGTTGGTGAGACAGACTGGCTCCTTTTGGACCCAGGTTCTAAA
GATCCTAAAAGGAATGATGAGTGTGAAAGGCAAGAGAATCAACTAAGATATAAAGCTTGTGTACGAGATCGTGTTGCCAAAGAACGATATAGAAGAAGCA
ATTCAACCCCACCATTTTACAGACTGAAGAGGAGGTTTATTTCCTTAAACAACCACTCAATGAGGAAAGAAGAAGAGCCATACACTCAGTTGTTCCATGA
TTGGCTCACTTCTCCAGAAGCTAATGACTTTGAGCATCTACCTCTCCAGCCTAGTCATGTGGAGGAGGATTTAACACAAAGAACCAAATCCAATGGAAAG
AACATGCCAGATACCATGCCGAACAAGGAGACTCCAGAAGGAAATCCAGAGCACTTCCAACACCCTAAGGCTTATGATTCTTCACCTGAAGCCTTTATGC
CAAAGGACACCCAGGAGTCAATGGATTATAGGATCAAATGGCGGAATGGCTGCCAACAAATTGCAAATCACAACACGTCATCCAATGTTGGCAGTCAGCG
AAATATACTTGACATCTCTTCGGGGTTCTTGCATCTTGCTGGAAATTTATTAGTTCCAGAATCTATTCATAAGAAGTGCCTTCAGGATGCCAGAGTCCTT
CATCAGGTAGACAAGAAATTCATCCCAATTGTAGCTGGTGGGACACTTGCTGTTATAGACCAGCATGCAGCAGATGAAAGGATTCGACTGGAAGAACTTC
GTCAAAAGGTATTATCTGGAGAAGAGAAGACAGTTACCTATCTTGATGCTGAACAAGAGCTGATACTTCCTGAAATAGGGTATCAGCTACTTCACAATTA
TGCTGAACAAGTAAGAGAATGGGGATGGATCTGCAGTATTCAAGGTTCAGGGACCTTCAAGAAGAATTTGAATATTCTCCACCAGCAGCCAACTGTGATT
ACACTTCTTGCAGTACCATGCATTTTAGGGGTCAATTTGTCTGATGGAGATCTCTTGGAATTTCTGCAGCAGCTTTCTGATACAGATGGCTCTTCAACTT
TGCCGCCATCTGTTCTTCGTGTCCTCAATTATAAAGCATGCAGAGGCGCAATCATGTTCGGAGATTCGTTGCTACCATCAGAATGTTCACTTATTGTCGA
AGAGCTAAAGCAGACCACACTTTGTTTTCAATGCGCCCACGGGCGACCAACAACTATTCCTGTTGTCAACCTGGAGGCGTTGCAAAAGCAGGTAGCAAAA
CTTGGAGTGCTAAATGATGGTTCAAATGACTTGTGGCATGGATTACGCCGACAGGAACTCAGCCTTGAACGTGCAGCACAGCGTTTAAGTGCAGCTAGAG
GCTAG
|
|||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.005G101900.2 pacid=42802914 polypeptide=Potri.005G101900.2.p locus=Potri.005G101900 ID=Potri.005G101900.2.v4.1 annot-version=v4.1
MGIIKRLPVSARSTMRSGILVFDLTRVVEELVFNSLDAGAKKVSVYVAVGTCYVKVSDDGCGISRDGLVLLGERYVTSKVQRLADMDVASGNFGFRGEAL
SSIADVSVLDVLTKARGMPNGYRKVMKGSKCLCLGIDDDIKDVGTTVVVRDLFYNQPVRRKYMQSSPKKILHLVKKCALRVALMHSEVSFKVVDIESDEE
LLCTNPSSAMSLLMSGFGIEDSSSLHELNISDSVLKLSGYISGPCSSFFIKAFQYVYINSRFVCKGPIHKLLNHLASRFEHPDLQKANSVSQKGKKSRPQ
PCPAYILNLSCPFSLYDLTFEPSKTHAEFKDWNPILAFIEKVIQQLWRECTVIGESSTRATDTFQKNDIWQEGNDITSVKQDFFDADFSGFAIKKGGVKI
HQSSHHLISCPLKMLDKEVDHLFHGKHDKVPQEFYSNVSEFKEEQVDKEFVLQGDYSSQTWNGSISGYMPRATKTDECHLLTSDKNFLLTDNCFLEDSFT
TRERLSDHMQSHFSSSEWQNESPKIDSVARNKSLGSAFSFDHYGFRNELPFSKSNIKPILQSCSSQKSLSLDRDFFADKEAFEFLNDGFKNKRRRLWTAE
NVGIPKGDTIFDIFPCALLQDNASCTQQLPADTDGAEMSAAFDLLPGAYVNSSSPNGKILAKGKGLASNSILQLEMYASGNHSSMSDWCSVTSSAFFQAK
VWDAEHFPDDNASEGSKGWGKKENCWHLPDSWEIMSKPSSQDNFFSSCTSSVLDFKNSADSSKDICKLPQWQDQNNEFSLQHSDISVGETDWLLLDPGSK
DPKRNDECERQENQLRYKACVRDRVAKERYRRSNSTPPFYRLKRRFISLNNHSMRKEEEPYTQLFHDWLTSPEANDFEHLPLQPSHVEEDLTQRTKSNGK
NMPDTMPNKETPEGNPEHFQHPKAYDSSPEAFMPKDTQESMDYRIKWRNGCQQIANHNTSSNVGSQRNILDISSGFLHLAGNLLVPESIHKKCLQDARVL
HQVDKKFIPIVAGGTLAVIDQHAADERIRLEELRQKVLSGEEKTVTYLDAEQELILPEIGYQLLHNYAEQVREWGWICSIQGSGTFKKNLNILHQQPTVI
TLLAVPCILGVNLSDGDLLEFLQQLSDTDGSSTLPPSVLRVLNYKACRGAIMFGDSLLPSECSLIVEELKQTTLCFQCAHGRPTTIPVVNLEALQKQVAK
LGVLNDGSNDLWHGLRRQELSLERAAQRLSAARG
|
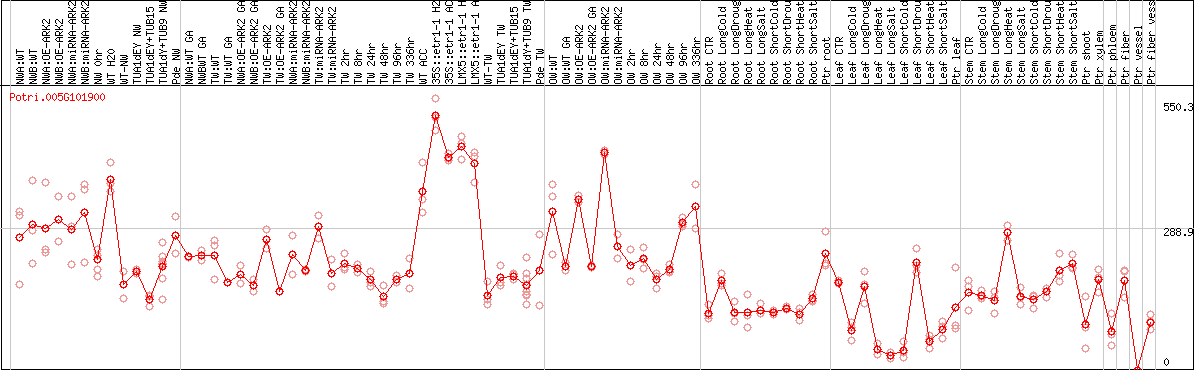
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G101900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
