External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G35750 284 / 1e-98
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
AT3G10210 148 / 1e-44
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
AT1G69340 66 / 2e-12
appr-1-p processing enzyme family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G014700
261 / 1e-89
AT4G35750 296 / 3e-103
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Potri.005G246800
248 / 2e-84
AT4G35750 282 / 9e-98
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Potri.006G043166
163 / 3e-50
AT3G10210 359 / 6e-127
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Potri.010G161500
61 / 8e-11
AT1G69340 926 / 0.0
appr-1-p processing enzyme family protein (.1)
Potri.008G093100
59 / 4e-10
AT1G69340 935 / 0.0
appr-1-p processing enzyme family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041841
290 / 6e-101
AT4G35750 323 / 7e-114
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Lus10028388
280 / 6e-97
AT4G35750 313 / 5e-110
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Lus10021230
231 / 1e-77
AT4G35750 265 / 7e-91
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Lus10005640
230 / 4e-77
AT4G35750 260 / 6e-89
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Lus10035431
166 / 4e-51
AT3G10210 353 / 2e-124
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Lus10031044
166 / 4e-51
AT3G10210 353 / 3e-124
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
Lus10030413
67 / 4e-13
AT1G69340 917 / 0.0
appr-1-p processing enzyme family protein (.1)
Lus10037119
62 / 4e-11
AT1G69340 966 / 0.0
appr-1-p processing enzyme family protein (.1)
Lus10036809
59 / 5e-10
AT1G69340 971 / 0.0
appr-1-p processing enzyme family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0512
CRAL_TRIO
PF13716
CRAL_TRIO_2
Divergent CRAL/TRIO domain
Representative CDS sequence
>Potri.005G107300.1 pacid=42804812 polypeptide=Potri.005G107300.1.p locus=Potri.005G107300 ID=Potri.005G107300.1.v4.1 annot-version=v4.1
ATGTGCACTCACATTTCTCTGTCTGACCAAGAACAACTTGTTGAGAAATTGGAGATTTTCAAATTTCAAGGCAGGGATAAAAATGGCCACAAAGTCCTTC
GAATTATCGGCAAATTCTTATCGGCTCGGTATTTGAGCGTGGATGCACTCAAGAATTATTTGGAGGAGAACATCTTTCCTAGGCTAAAGAAGAAGCCATT
TTCGGTCTTGTACCTTCACACACAAGTACAAAAGAGTGAGGATTTTCCAGGAATCTCAGCCCTCCGATCAATCTATGATGTTATACCAATCAACGCTAGG
GATAATCTCCAGGCAATTTACTTTGTCCACCCCAGCCTACAAGCCAAACTTTTCCTTGCCACGTTTGGCCGTCTTCACTTCGGCAGCAGGTTGTATGGGA
AGCTAAGATATATCAATAGGATTGATTATTTATGGGACCCTATAAGGAGAAATGAAGTAAAAATCCCTGAGTTCGTTTGTGATCATGATGAAGATTTGGA
GGGCCATCAGATGCTGGACTATGGGGTGGAGAGTGATCATCCTAGAGTTTGTGGTGCTCCCTTCATGGATTCTCCTGTGACCATGTACTCAACGAGGTGT
ATTTAG
AA sequence
>Potri.005G107300.1 pacid=42804812 polypeptide=Potri.005G107300.1.p locus=Potri.005G107300 ID=Potri.005G107300.1.v4.1 annot-version=v4.1
MCTHISLSDQEQLVEKLEIFKFQGRDKNGHKVLRIIGKFLSARYLSVDALKNYLEENIFPRLKKKPFSVLYLHTQVQKSEDFPGISALRSIYDVIPINAR
DNLQAIYFVHPSLQAKLFLATFGRLHFGSRLYGKLRYINRIDYLWDPIRRNEVKIPEFVCDHDEDLEGHQMLDYGVESDHPRVCGAPFMDSPVTMYSTRC
I
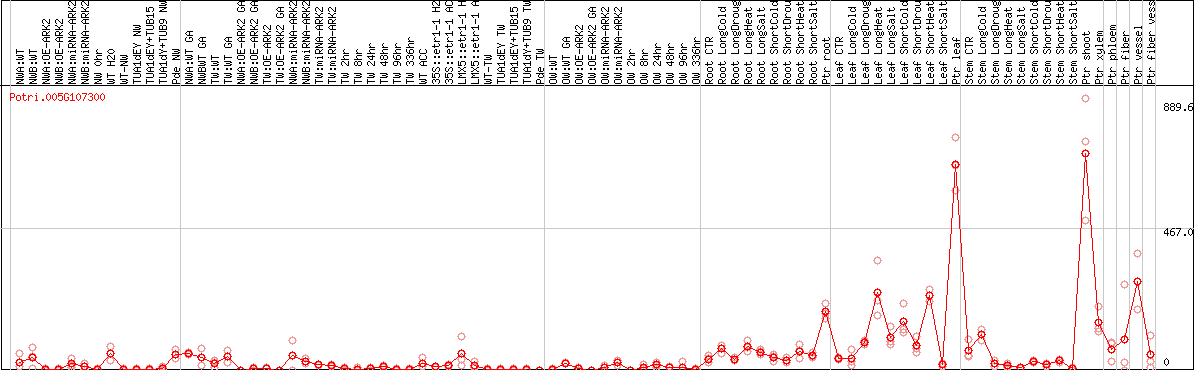
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.005G107300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.